|
|
Post by Admin on Nov 6, 2019 18:46:29 GMT
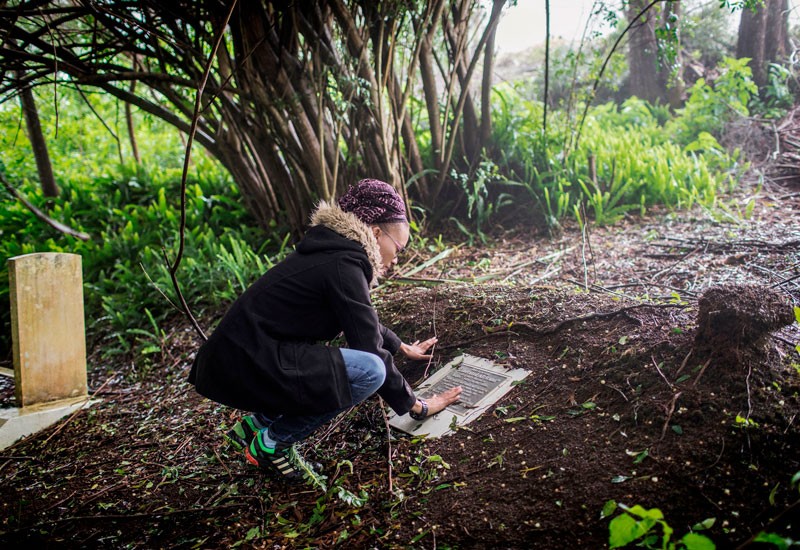 Annina Van Neel Hayes looks at a plaque commemorating the burial place of the remains of some liberated African slaves. Genomes from enslaved Africans who were freed and died on a remote Atlantic island in the mid-nineteenth century are offering clues about their origins in Africa. The findings come from the largest study of genome data obtained from remains of enslaved people and offer insights into the transatlantic slave trade, in which an estimated 12 million Africans were kidnapped and enslaved in North and South America and the Caribbean. 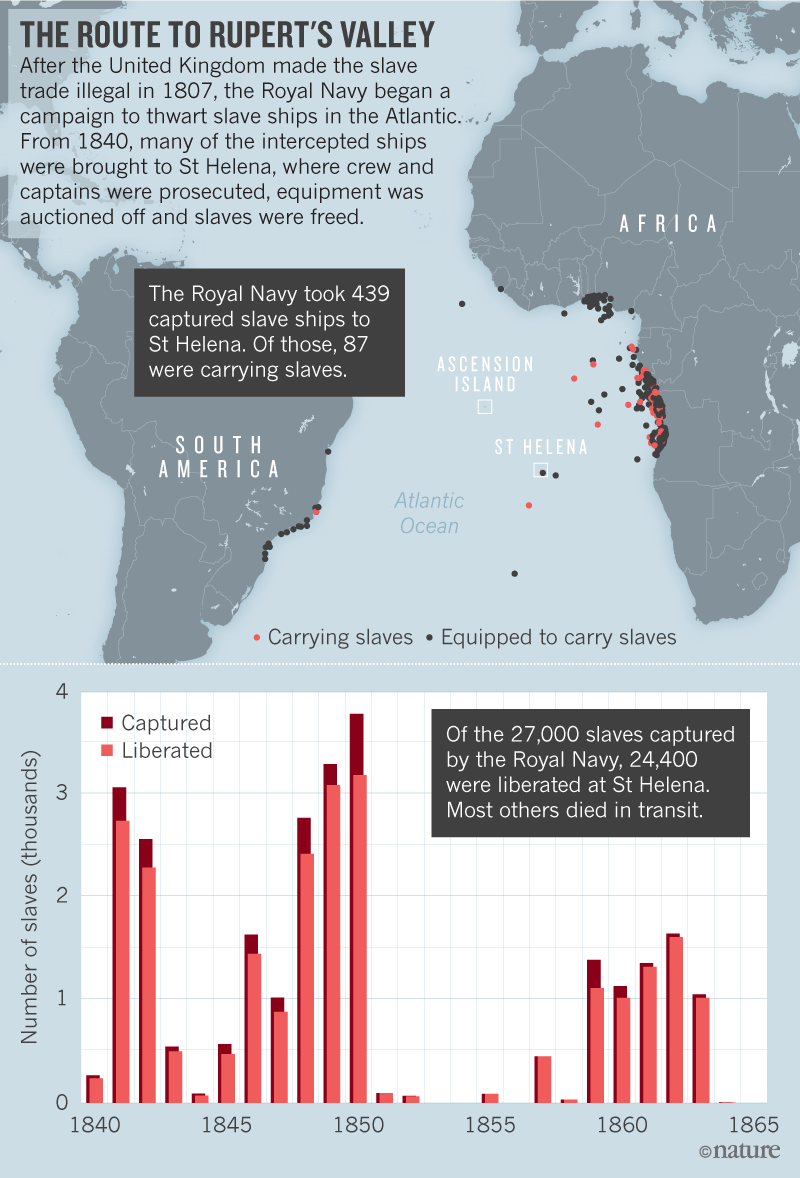 No island paradise St Helena, which lies in the Atlantic Ocean nearly 2,000 kilometres west of Angola, occupies a unique chapter in the history of the transatlantic trade in people. After Britain outlawed the slave trade in 1807, its navy intercepted slave ships and sent an estimated 24,000 people to St Helena (see ‘The route to Rupert’s Valley’). They had been aboard ships heading largely to Brazil and Cuba between 1840 and the late 1860s. Many of the people freed arrived in poor health and were housed in squalid conditions in an isolated coastal valley, and as many as 10,000 died on the island. In 2006, construction work for St Helena’s first airport uncovered mass burials. Archaeologists unearthed the remains of 325 people — more than half under 18 and many younger than 12.  Seventeen were male — backing up records indicating that, in its final decades, the transatlantic slave trade captured far more men than women. Analysis of the genome data found that none of the people were closely related, nor did they belong to a single African population. Comparisons with genome data from thousands of modern Africans from dozens of populations suggest that the people from St Helena are most closely related to people living today in central Gabon and northern Angola. But the researchers caution that gaps in present-day genome data from potential homelands, such as the Democratic Republic of the Congo, make it difficult to say for certain where the people buried in St Helena were taken from. “Although it’s very hard to exactly pinpoint their origins, I think what we see in our results is that they are not coming from a single population,” says Sandoval-Velasco. This insight suggests that the liberated Africans taken to St Helena lived in a challenging multicultural setting where they might not have understood the language and customs of others left on the island. “We hope that by illustrating the history and the condition of a few, we are at the same time illustrating the condition of the many, but it shouldn’t stop there,” Sandoval-Velasco says. Individual stories Ancient-genome analysis is a powerful tool for shining a light on people exploited in one of history’s darkest chapters, says Rosa Fregel, a population geneticist at the University of La Laguna in the Canary Islands, who was not involved in the St Helena study. “Usually it’s just about numbers — how many people from each country. Here, we are talking about particular people and their origin,” says Fregel, who is applying ancient genomics to illuminate the histories of people captured in the Indian Ocean slave trade. “Ancient DNA has the potential to tell their story.” The data lay a solid foundation for studies that could pinpoint the specific regions that the liberated people were from, says Fatimah Jackson, a biological anthropologist at Howard University in Washington DC. The key to identifying the origins of enslaved people, she says, will be expanding data sets of modern Africans, as well as sequencing more remains. She and her colleagues have skeletal material from all 325 people that were recovered from the St Helena burial and hope to generate genome data soon. David Eltis, a historian at Emory University in Atlanta, Georgia who co-founded a database that collects information on 36,000 slaving voyages between 1514 and 1866, notes that most people captured in the transatlantic slave trade originated from south of the equator — where a paucity of genome data from modern inhabitants makes it difficult to trace the origins of enslaved individuals with any accuracy. Reburial plan Although working with human remains can be ethically fraught, particularly when there are no known direct descendants to consult, the work can have value when carried out with sensitivity, says Jada Benn Torres, an anthropologist at Vanderbilt University in Memphis, Tennessee. (Several hundred of the liberated Africans later integrated into St Helena’s population, but it is not clear if they left any descendants.) Studies like this add another layer to the historical record, bringing to life the moving personal stories behind the slave trade, she says. Remains of the 325 liberated Africans that were excavated are in storage on St Helena. In 2018, the territory’s government endorsed plans to rebury them in the valley where they were first uncovered and to create a memorial at the site. Nature 575, 18 (2019) |
|
|
|
Post by Admin on Nov 6, 2019 22:05:42 GMT
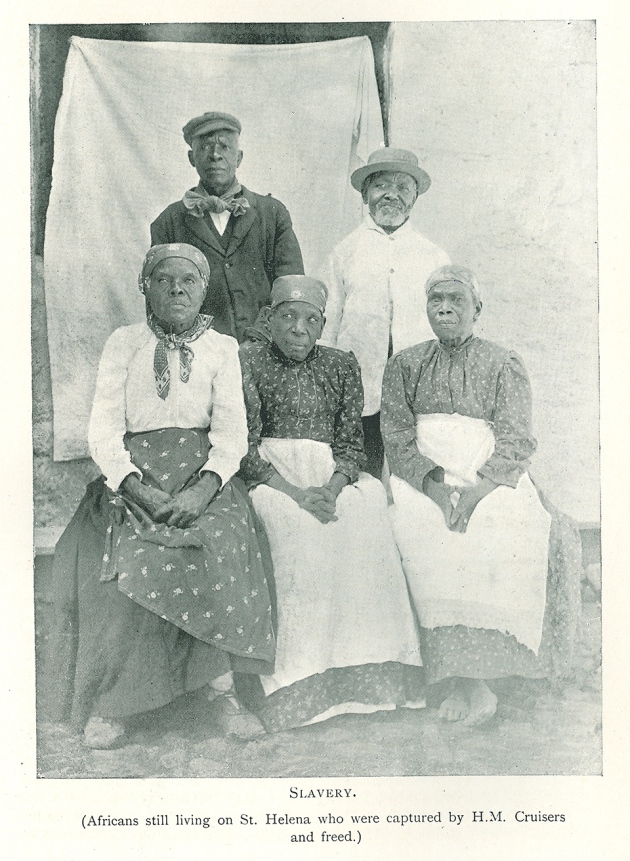 Valley of lost souls Rupert's Valley is a seaside gorge on the remote isle where Napoleon Bonaparte spent his final days in exile. Driving through it, you pass warehouses, a small diesel power station and a fish-processing plant, as well as a few houses and a lone church. The valley is also dotted with crumbling reminders of its past, such as a metre-wide defensive wall built by the British East India Company and a late-nineteenth-century desalination plant that supported a prison camp during the Second Boer War. But aside from a nondescript stone building, there is little visible reminder of the thousands of former slaves who once resided here.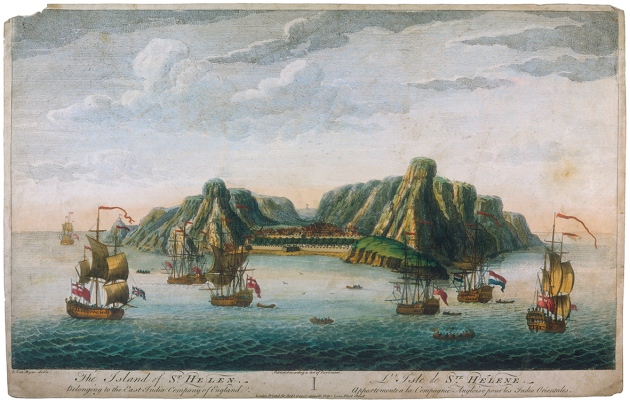  Historical records illustrate the dire condition in which many newly liberated slaves arrived. “Their arms and legs were worn down to about the size of a walking-stick,” wrote one witness of an 1861 slave-ship landing. “Many died as they passed from the ship to the boat … there was no time to separate the dead from the living.” Diseases such as smallpox and dysentery killed scores more soon after they landed, and there are oblique references to suicides. Between 1840 and 1849, one-third of the freed slaves, nearly 5,000 people, died. Many of those who survived the camp's miserable conditions were sent to Jamaica, Trinidad and other British colonies, where they served as indentured workers on sugar plantations. After Shilston found the first remains of St Helena's Liberated African Graveyard, as the site is now known, he learned that the burials hadn't been so much forgotten as ignored by many of St Helena's residents. His discovery made the front page of a local newspaper. “There is no doubt that numerous remains of human bodies will be found,” the story read. “Many believe that the souls of the dead slaves still are haunting Rupert's.” The remains were stored in small caskets at a local church, and later reburied in a cemetery in the island's capital, Jamestown. 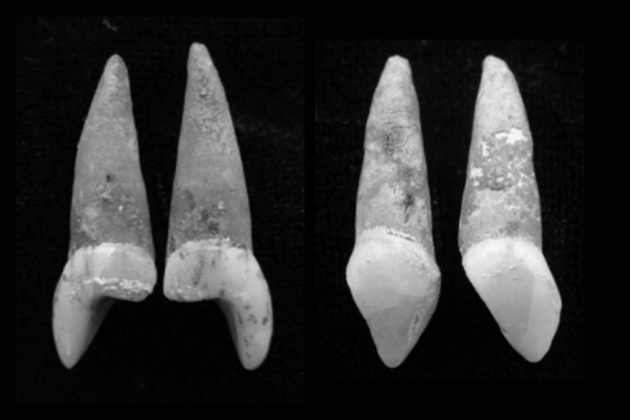 North and South America and the Caribbean islands are full of burial sites related to the transatlantic slave trade, but these tend to hold multiple generations of slaves: some born in Africa and others in the Americas. Rupert's Valley, by contrast, contains physical evidence of the individuals who were sold into slavery and then freed. It therefore has the potential to indicate not only where slaves came from and their condition aboard the ships, but also how the slave trade might have reshaped their identities. “It's literally people who are kidnapped in Africa, weeks before,” says Pearson. “So you actually have a snapshot of the Middle Passage, which in other ways is completely intangible.” Comparisons with DNA from 11 contemporary African groups suggested that one of the St Martin individuals was most closely related to members of a Bantu-speaking ethnic group in northern Cameroon called the Bamoun, whereas the other two had DNA in common with ethnic groups that now live in Nigeria and Ghana, including the Igbo and Brong. Schroeder is hesitant to conclude that they belonged to these groups, but the findings, published last year (H. Schroeder et al. Proc. Natl Acad. Sci. USA 112, 3669–3673; 2015), gave him hope that ancient DNA could offer genuine insights into the genetic ancestry of slaves, as well as how the Middle Passage reshaped their identities. “These three individuals — despite the fact that they were found buried together and may have arrived on the same vessel — had different ethnic backgrounds” and probably spoke distinct languages, he says. “That makes you think: how did they communicate with each other and what does this mean for the formation of new identities in the Americas?”  Rupert's Valley was probably even more polyglot. Shipping records suggest that slave ships that landed in St Helena had embarked from ports across Central and West Africa, including present-day Angola and the Congo region (see 'The route to Rupert's Valley'). There are indications that some individuals came from much further away, including Mozambique and even Madagascar. One 1840s observer noted a group of 40 “natives of the interior”, who had travelled for several months to reach the Atlantic coast. “They are of so many different tribes and districts that it would be curious, if one knew the languages, to trace them out,” said another visitor to Rupert's Valley. The former slaves' DNA backs up the observation. Schroeder and Marcela Sandoval Velasco, a palaeogeneticist at the University of Copenhagen, collected DNA from the teeth of 63 individuals and sequenced partial genomes from 20 of the best-preserved samples. Comparisons with contemporary African populations suggested that the liberated slaves came from diverse African backgrounds. A few individuals shared ancestry with contemporary West and Central African ethnic groups such as the Bamoun and Kongo, but for most, none of the African groups that the team compared them with was an especially close match. Schroeder attributes that to a lack of genomic data from places such as Angola and Mozambique. |
|
|
|
Post by Admin on Nov 7, 2019 18:33:38 GMT
Historians have long been interested in the origins of the millions of enslaved Africans who were transported to the Americas, but there is little direct evidence to go on (1, 2). Although much is known about the volume and changing demographic trends of the Atlantic slave trade, the African origins of the enslaved remain largely unknown (3). Genome-wide analyses of SNPs provide a powerful tool for estimating individual ancestry, and several studies (e.g., refs. 4 and 5) have shown that they can be used to infer an individual’s geographic origin with great accuracy. In the present study, we used genome-wide data to trace the origins of three enslaved Africans whose remains were recovered in the Zoutsteeg area of Philipsburg on the Caribbean island of Saint Martin (Materials and Methods). Previous reports (6) suggest that the “Zoutsteeg Three,” as they became known locally, were likely born in Africa as opposed to the New World. But they did not reveal where in Africa they originated. Bayesian analysis of individual calibrated radiocarbon dates suggests that the burials date between A.D. 1660 and 1688 (for more details, see SI Appendix, Section 2). During this period, Saint Martin—like other islands—relied to a large extent on African slave labor, but there are no records about the slaves’ origins in Africa. The Transatlantic Slave Trade Database (slavevoyages.org), a large online repository containing information on over 35,000 slaving voyages, lists only a single vessel that arrived in Saint Martin between A.D. 1650 and 1700, although there were undoubtedly more for which there are no records. Moreover, the lone entry does not give the African port of embarkation or any information on the slave cargo. Given the lack of documentation, we embarked on a genomic study with the goal of identifying the genetic origins of the Zoutsteeg Three in Africa.
Table 1.
Modeled radiocarbon dates and sequencing results for the Zoutsteeg Three
Sample Modeled 14C age* Sex† Nuclear coverage Contamination (%)‡ Damage (%)§ Mt coverage Mt haplogroup Y haplogroup
STM1 A.D. 1660–1688 M 0.3× 0.63 16.8 641× L3b1a R1b1c-V88
STM2 A.D. 1660–1688 M 0.1× 0.22 23.2 543× L3d1b —
STM3 A.D. 1660–1688 F 0.5× 0.15 14.9 651× L2a1f —
↵* Modeled age range of the samples based on Bayesian analysis of individual calibrated radiocarbon dates.
↵† Biological sex inferred from the ratio of reads mapping to X and Y chromosomes (44).
↵‡ Likelihood-based contamination estimate based on mtDNA reads (10).
↵§ Frequency of C-to-T misincorporations at 5' ends of sequencing reads.
Results Initial shotgun sequencing revealed that the DNA in the samples was very poorly preserved (SI Appendix, Section 8). This result was expected because DNA preservation in the Caribbean is known to be poor (7). The fraction of nonredundant reads mapping to the human reference genome varied between 0.3% and 7.6%, and the sequences showed all features typical of ancient DNA, including short average read lengths (∼67 bp), characteristic fragmentation patterns, and an increased frequency of apparent C-to-T substitutions toward the 5′ ends of molecules (see Table 1 and SI Appendix, Fig. S7). To increase sequencing efficiency and lower cost, we enriched the ancient DNA libraries using two recently developed whole-genome capture methods (8, 9). Following enrichment, we generated between 0.1- and 0.5-fold genome-wide coverage for the three individuals, which proved to be sufficient to infer their likely origins within Africa. Although the presence of characteristic damage patterns and short average read lengths suggest that authentic ancient molecules were sequenced, it is possible that some degree of modern contamination could be present in the data. Therefore, we used a previously published Bayesian method (10) to detect contamination and found very low (i.e., <1%) levels overall (see Table 1 and SI Appendix, Section 9). We then merged our ancient sequence data with genotype data from the Human Genome Diversity Cell Line Panel (HGDP) reference panel (11) and used principal component analysis (PCA) (12) to confirm the individuals’ African ancestry (for more details, see SI Appendix, Section 12). Because of low depth, we randomly sampled a single allele from both ancient and modern individuals, as done in ref. 13. As expected, all three individuals fell within African variation as defined by PC1 and PC2 (SI Appendix, Fig. S17). This clustering was retained upon restricting the analysis to sequences showing signs of ancient DNA damage (14) (SI Appendix, Fig. S18), indicating that the signal was not driven by modern DNA contamination.  Fig. 1. Genetic affinities of the Zoutsteeg individuals. (A) D-statistic test results for STM3. Error bars correspond to 3 SEs of the D-statistic. Results for STM1 and STM2 are plotted in SI Appendix, Fig. S18. (B) Sampling locations for the 11 African populations in our reference panel (17). (C) Procrustes-transformed PCA plot of the Zoutsteeg individuals with African reference panel samples. (D) Ancestry proportions for the Zoutsteeg individuals and those of 188 African individuals in the reference panel, as inferred by ADMIXTURE analysis (19) D-Statistic Test. We then tested the relationships between each sample (henceforth referred to as STM1, STM2, and STM3) and 11 populations from across the world for which whole genome data were available (15), using a D-test of the form D (chimpanzee, STM; Yoruba, X), where X stands for a population other than Yoruba (for more details, see SI Appendix, Section 13). We found that the STMs were significantly more closely related to the Yoruba than to any non-African population (Fig. 1A), again confirming their African origins. Within Africa, the STMs appeared significantly more closely related to the Yoruba than to hunter-gatherer populations (San, Mbuti Pygmies). This was not surprising, as the San and the Mbuti were not represented in the transatlantic slave trade. For the Mandenka and Dinka, the D-test results were not significant, suggesting that these populations are equally closely related to the STMs as are the Yoruba. The lack of rejection for the Dinka was surprising, as this population—from southern Sudan—is not known to have been involved in the Atlantic slave trade (16). |
|
|
|
Post by Admin on Nov 7, 2019 23:32:49 GMT
Principal Component Analysis. To refine our assignment within Africa, we compared our samples to another reference panel, consisting of genotype data from 11 West African populations (Fig. 1B) (17). We intersected 294,651 sites from this reference panel with our sequence data (SI Appendix, Section 14) and conducted PCA to determine whether the individuals showed close affinity to a particular population within the panel. For each of the three individuals, we merged the sequence data with the reference panel genotypes and calculated PC1 and PC2 based on the overlapping sites (SI Appendix, Fig. S19). We then combined the three analyses using Procrustes transformation, as done in ref. 13. Interestingly, the samples clustered with different populations: Bantu-speaking groups in the case of STM1 (specifically, Bamoun) and non-Bantu–speaking groups for STM2 and STM3 (Fig. 1C). We observed similar patterns using the probabilistic model of population splits and divergence implemented in TreeMix (18) (SI Appendix, Fig. S20). ADMIXTURE Analysis. To further explore the genetic ancestry of the STMs, we used the maximum-likelihood–based clustering algorithm ADMIXTURE (19). When assuming three ancestral populations (K = 3), the clusters in the reference panel mirror the grouping of individuals in the space defined by PC1 and PC2: a cluster predominating in Bantu-speaking populations, a cluster for non-Bantu West African populations, and a third restricted mostly to Kaba, Mada, and Bulala (Fig. 1D). The distribution of these components in our samples indicates that STM1 has a higher proportion of Bantu-specific ancestry, whereas STM2 and STM3 carry higher proportions of the component prevalent among the non-Bantu–speaking Yoruba, Brong, and Igbo. Notably, STM2 also shows a slightly higher proportion of the component prevalent among the Kaba, Mada, and Bulala, perhaps suggesting closer affinity with Chadic or Sudanic speakers (Fig. 1D).  Uniparental Markers. Furthermore, we determined the individuals’ mitochondrial DNA (mtDNA) haplogroups and the Y-chromosome haplogroup of STM1 (SI Appendix, Section 11). The mtDNAs were assigned to haplogroups L3b1a, L3d1b2, and L2a1f, respectively. Tracing these lineages to particular regions in Africa is challenging because of their pan-continental distribution, which is the result of thousands of years of population movements (e.g., the Bantu migrations) and continued gene flow (20⇓–22). Nevertheless, we note that haplogroup L3b1a is one of the most common lineages found in the Lake Chad Basin (23). This finding is noteworthy, because the Y-chromosome lineage of this individual (STM1) was identified as belonging to haplogroup R1b1c-V88, which—although quite rare in Africa on the whole—occurs at extremely high frequency in the Lake Chad Basin, rising to 95% in one population of northern Cameroon (24).  Discussion Taken together, the genetic data suggest that STM1 may have originated among Bantu-speaking groups in northern Cameroon, whereas STM2 and STM3 more likely originated among non-Bantu speakers living in present-day Nigeria and Ghana. To our knowledge, these findings provide the first direct evidence for the ethnic origins of enslaved Africans, with the important caveat that the modern reference populations might not be the same as the historical populations who lived in the same locations at the time of the Atlantic slave trade. Nevertheless, the data suggest that the Africans who reached Saint Martin were drawn from diverse cultures and societies. This finding highlights interesting questions regarding the formation of Creole communities and the survival of African cultures in the Americas. Chief among these is to what extent Africans were able to maintain African cultural beliefs and practices following their arrival in the New World (see, for example, refs. 25⇓–27). Genome-wide analyses clearly bear great potential to predict a person’s place of origin, but we caution that there are also limitations. First, accuracy is limited by the number of markers used, although new methods (e.g., ref. 28) are constantly being developed, claiming to achieve greater accuracy using relatively small sets of makers. Unfortunately, however, many of these methods rely on called genotypes, which makes them unsuited to low-coverage ancient genome studies. Second, accuracy also depends on the appropriate samples being represented in our reference panels, highlighting the need for more comprehensive sampling and sequencing of human populations across Africa. Third, tracing the ancestry of admixed individuals is more complicated, but advances are also being made in the study of the ancestral composition of admixed genomes (see, for example, refs. 29 and 30). Many of these limitations will be overcome, as more data are being generated and new analytical methods are being developed. PNAS March 24, 2015 112 (12) 3669-3673; first published March 9, 2015 |
|










