|
|
Post by Admin on Sept 3, 2020 19:31:06 GMT
SARS-COV-2 INFECTION IN QUARANTINE Of the 1797 qPCR-positive Icelanders, 1088 (61%) were in quarantine when SARS-CoV-2 infection was diagnosed by qPCR. We tested for antibodies among 4222 quarantined persons who had not tested qPCR-positive (they had received a negative result by qPCR or had simply not been tested). Of those 4222 quarantined persons, 97 (2.3%; 95% CI, 1.9 to 2.8) were seropositive (Table 1). Those with household exposure were 5.2 (95% CI, 3.3 to 8.0) times more likely to be seropositive than those with other types of exposure (Table 3); similarly, a positive result by qPCR for those with household exposure was 5.2 (95% CI, 4.5 to 6.1) times more likely than for those with other types of exposure. When these two sets of results (qPCR-positive and seropositive) were combined, we calculated that 26.6% of quarantined persons with household exposure and 5.0% of quarantined persons without household exposure were infected. Those who had symptoms during quarantine were 3.2 (95% CI, 1.7 to 6.2) times more likely to be seropositive and 18.2 times (95% CI, 14.8 to 22.4) more likely to test positive with qPCR than those without symptoms. We also tested persons in two regions of Iceland affected by cluster outbreaks. In a SARS-CoV-2 cluster in Vestfirdir, 1.4% of residents were qPCR-positive and 10% of residents were quarantined. We found that none of the 326 persons outside quarantine who had not been tested by qPCR (or who tested negative) were seropositive. In a cluster in Vestmannaeyjar, 2.3% of residents were qPCR-positive and 13% of residents were quarantined. Of the 447 quarantined persons who had not received a qPCR-positive result, 4 were seropositive (0.9%; 95% CI, 0.3 to 2.1). Of the 663 outside quarantine in Vestmannaeyjar, 3 were seropositive (0.5%; 95% CI, 0.1 to 0.2%).  SARS-COV-2 SEROPREVALENCE IN ICELAND None of the serum samples collected from 470 healthy Icelanders between February 18 and March 9, 2020, tested positive for both pan-Ig antibodies, although four were positive for the pan-Ig anti-N assay (0.9%), a finding that suggests that the virus had not spread widely in Iceland before March 9. Of the 18,609 persons tested for SARS-CoV-2 antibodies through contact with the Icelandic health care system for reasons other than Covid-19, 39 were positive for both pan-Ig antibody assays (estimated seroprevalence by weighting the sample on the basis of residence, sex, and 10-year age category, 0.3%; 95% CI, 0.2 to 0.4). There were regional differences in the percentages of qPCR-positive persons across Iceland that were roughly proportional to the percentage of people quarantined (Table S6). However, after exclusion of the qPCR-positive and quarantined persons, the percentage of persons who tested positive for SARS-CoV-2 antibodies did not correlate with the percentage of those who tested positive by qPCR. The estimated seroprevalence in the random sample collection from Reykjavik (0.4%; 95% CI, 0.3 to 0.6) was similar to that in the Health Care group (0.3%; 95% CI, 0.2 to 0.4) (Table S6). We calculate that 0.5% of the residents of Iceland have tested positive with qPCR. The 2.3% with SARS-CoV-2 seroconversion among persons in quarantine extrapolates to 0.1% of Icelandic residents. On the basis of this finding and the seroprevalence from the Health Care group, we estimate that 0.9% (95% CI, 0.8 to 0.9) of the population of Iceland has been infected by SARS-CoV-2. Approximately 56% of all SARS-CoV-2 infections were therefore diagnosed by qPCR, 14% occurred in quarantine without having been diagnosed with qPCR, and the remaining 30% of infections occurred outside quarantine and were not detected by qPCR. DEATHS FROM COVID-19 IN ICELAND In Iceland, 10 deaths have been attributed to Covid-19, which corresponds to 3 deaths per 100,000 nationwide. Among the qPCR-positive cases, 0.6% (95% CI, 0.3 to 1.0) were fatal. Using the 0.9% prevalence of SARS-CoV-2 infection in Iceland as the denominator, however, we calculate an infection fatality risk of 0.3% (95% CI, 0.2 to 0.6). Stratified by age, the infection fatality risk was substantially lower in those 70 years old or younger (0.1%; 95% CI, 0.0 to 0.3) than in those over 70 years of age (4.4%; 95% CI, 1.9 to 8.4) (Table S7).  AGE, SEX, CLINICAL CHARACTERISTICS, AND ANTIBODY LEVELS SARS-CoV-2 antibody levels were higher in older people and in those who were hospitalized (Table 4, and Table S8 [described in Supplementary Appendix 1 and available in Supplementary Appendix 2]). Pan-Ig anti–S1-RBD and IgA anti-S1 levels were lower in female persons. Of the preexisting conditions, and after adjustment for multiple testing, we found that body-mass index, smoking status, and use of antiinflammatory medication were associated with SARS-CoV-2 antibody levels. Body-mass index correlated positively with antibody levels; smokers and users of antiinflammatory medication had lower antibody levels. With respect to clinical characteristics, antibody levels were most strongly associated with hospitalization and clinical severity, followed by clinical symptoms such as fever, maximum temperature reading, cough, and loss of appetite. Severity of these individual symptoms, with the exception of loss of energy, was associated with higher antibody levels. We estimate that during the first wave of the SARS-CoV-2 pandemic, the incidence of infection in Iceland was 0.9% (95% CI, 0.8 to 0.9) and the infection fatality risk was 0.3% (95% CI, 0.2 to 0.6). Our estimate of the infection fatality risk is lower than but consistent with estimates described by others.6-8 We estimate that of the infected persons, 56% had cases previously diagnosed by qPCR, 14% had been in quarantine (but either had not been qPCR-tested or had tested negative), and 30% neither were known to be qPCR-positive nor had been placed in quarantine. We therefore conclude that, despite extensive screening by qPCR, a substantial fraction of infections were not detected, which indicates that many infected persons did not have substantial symptoms. The case fatality risk is straightforward to estimate but may differ across countries and over time. An accurate calculation of infection fatality risk requires an accurate estimate of the number of infections, both diagnosed and undiagnosed. In Iceland, the high percentage of infections identified through qPCR (56%) as compared with that of other countries (for example, approximately 9% in Spain4) renders a commensurately accurate estimate of the total number of infections. Each of the pan-Ig SARS-CoV-2 antibody assays that we used has high specificity (99.8%, according to the manufacturers’ literature), which raises the question of whether using a single pan-Ig assay would have sufficed. One sample obtained in 2017 was positive on only one pan-Ig antibody assay, a finding that supports the use of two separate assays to determine seroprevalence, if the infection rate is below 1%, as in Iceland. By April 30, a total of 20,766 Icelanders had been placed in quarantine. Of the 1797 Icelanders who tested positive by qPCR, 1088 (61%) were in quarantine when tested. Despite substantial qPCR testing of persons in quarantine, 2.3% of persons in quarantine who did not receive qPCR-positive result (i.e., a diagnosis of infection) developed SARS-CoV-2 antibodies. Household exposure was more likely to lead to infection than other types of exposure, which suggests that people who share a household with an infected person should not have contact during quarantine and that contacts of household members should be quarantined. Seroprevalence in the two regional hot spots (Vestfirdir and Vestmannaeyjar) was absent or low outside quarantine, which indicates that most infections were detected by qPCR screening and that quarantine, social distancing, contact tracing, and limits on public gatherings were effective in limiting spread. Over 90% of qPCR-positive persons tested positive with both pan-Ig SARS-CoV-2 antibody assays and remained seropositive 120 days after diagnosis, with no decrease of antibody levels as detected by the two pan-Ig assays. We observed some diminution of antibody titer with some of the single-Ig assays. Previous smaller studies reported reduction of IgG antibodies against the N protein and a peptide representing the S protein within 21 to 28 days5 and against trimeric S protein within 56 days14 after a positive test by qPCR. These discrepancies may be explained partly by differences in the specificity and sensitivity of the assays used as well as differences in the design and performance of the semiquantitative assays used, including the antigen targeted and the analytic sensitivity and range, as well as differences in the study populations. For example, because of widespread qPCR testing and screening, it is likely that the Icelandic qPCR-positive persons were healthy, as compared with the participants in other studies. Repeated SARS-CoV-2 exposure is unlikely to affect the persistence of antibody levels in Iceland, given the low prevalence of infection. Comparative studies using validated quantitative SARS-CoV-2 antibody assays are needed; those described in the published literature are based on small sample sizes.9-12 Of the 22 recovered persons who had a negative result (using the combined pan-Ig antibody tests) for an early sample and who had another sample tested at least a month later, 19 (86%) received a second negative result. Thus, either some persons infected by SARS-CoV-2 produce no antibodies or undetectable levels of antibodies reactive to the S1 and N proteins, even 3 months after infection, or some qPCR delivered false positive results. Among recovered persons, antibody levels are higher in older persons and in those more severely affected by SARS-CoV-2 infection. Women, who tend to become less sick than men, had lower antibody levels in two spike protein antibody assays. SARS-CoV-2 antibody levels were lower in smokers. Smoking increases the probability of severe Covid-19 illness among young adults,15 and smoking has been reported to increase the expression of ACE2,16 the receptor for cellular entry of the SARS-CoV-2 virus. The humoral immune response is critical for the clearance of cytopathic viruses and is generally important for the prevention of viral reinfection.17 A relationship between a humoral immune response to SARS-CoV-2 infection and protection against reinfection by this virus has been shown in rhesus macaques18 but has yet to be established in humans. Regardless of the relationship or lack thereof between seropositivity against SARS-CoV-2 and protection against reinfection, the low SARS-CoV-2 antibody seroprevalence in Iceland indicates that the Icelandic population is vulnerable to a second wave of infection. September 1, 2020 DOI: 10.1056/NEJMoa2026116 |
|
|
|
Post by Admin on Nov 4, 2020 5:50:44 GMT
Cellular (T cell) immunity against SARS-CoV-2 is likely to be present within most adults six months after primary infection, a new pre-print on bioRxiv suggests. The research from the UK Coronavirus Immunology Consortium (UK-CIC), Public Health England and Manchester University NHS Foundation Trust demonstrates robust T cell responses to SARS-CoV-2 virus peptides at this timepoint in all participants following asymptomatic or mild/moderate COVID-19 infection. A key question is whether previous infection with SARS-CoV-2 results in immunity to reinfection, and if so for how long. The immune system is extremely complex and there are many different potential routes whereby it can generate immunity to a disease post-infection. This study examines the role of T cells in contributing to immunity against SARS-CoV-2 at six months post infection. 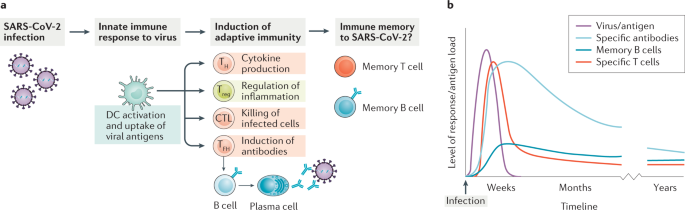 As part of UK-CIC, researchers from the University of Birmingham, Public Health England, Manchester University NHS Foundation Trust (MFT) and NIHR Manchester Clinical Research Facility collected serum and blood samples from a cohort of more than 2,000 clinical and non-clinical healthcare workers including 100 individuals who tested sero-positive for SARS-CoV-2 in March/April 2020 (average age 41 (range 22–65); 23 men, 77 women). All 100 individuals experienced either mild/moderate symptoms or were asymptomatic (56 versus 44 people) and none were hospitalised for COVID-19. Serum samples were collected monthly to measure antibody levels, and blood samples were taken after six months to assess the cellular (T cell) response. A range of analyses were carried out to assess different aspects of the T cell response including the magnitude of response and the response to different proteins from SARS-CoV-2. Carrying out these cellular analyses is much more complex than antibody studies – but this study of 100 individuals is one of the largest in the world to date in this field. T cell responses were present in all individuals at six months after SARS-CoV-2 infection. The cellular immune response was directed against a range of proteins from the virus, including the Spike protein that is being used in most vaccine studies. However, comparable immunity was present against additional proteins, such as nucleoprotein, which suggests that these may be of value for incorporation in future vaccine protocols. This indicates that a robust cellular memory against the virus persists for at least six months.  The size of T cell response differed between individuals, being considerably (50%) higher in people who had experienced symptomatic disease at the time of infection six months previously. Further research will be needed to determine the significance of this finding. It is possible that heightened cellular immunity might provide increased protection against re-infection in people with initial symptomatic infection, or that asymptomatic individuals are simply able to fight off the virus without the need to generate a large immune response. Antibodies are also a crucial component of immune defence and cellular immunity was strongly correlated with the peak level of the antibody response. Furthermore, larger cellular responses appeared to protect against antibody ‘waning’ over time, again suggesting the need to ensure that cellular immune responses are elicited in vaccine regimens. Overall, these findings indicate a robust cellular (T cell) immune response against SARS-CoV-2 at six months post-infection. These findings will feed not only into our understanding of how immunity to SARS-CoV-2 works but also help inform future vaccine strategies. Further research is now needed to assess whether this immune response is maintained over the longer term and to better understand how strength of cellular immune response corresponds to likelihood of reinfection. The study of 100 people showed that all had a cellular immune response against SARS-CoV-2 six months after infection although the size of response was 50% higher in those who had experienced symptomatic disease. There has been concern that the cellular immune response following covid-19 infection may not be sustained. “This data is reassuring,” lead study author Paul Moss, from the University of Birmingham, told a Science Media Centre briefing on 2 November. “However, it does not mean that people cannot be re-infected. We need to have much larger population studies to show that.” Moss also added that the findings “can’t be taken as confirmation that an ‘immunity passport’ would be feasible.” The study from the UK Coronavirus Immunology Consortium and Public Health England, which is published as a preprint and has not yet been peer reviewed, is believed to be the first in the world to show that a robust cellular memory against the virus persists for at least for six months. The researchers collected serum and blood samples from a cohort of more than 2000 clinical and non-clinical healthcare workers, including 100 who tested seropositive for SARS-CoV-2 in March and April 2020. The average age of the donors was 41 (range 22 to 65 years old); 23 were men and 77 were women. None of them were hospitalised with covid-19—56 people had mild or moderate symptoms and 44 were asymptomatic. Serum samples were collected monthly to measure antibody levels and blood samples were taken after six months to measure the T cell response using an ELISPOT and ICS analysis. The study found that virus specific T cells were detectable in all donors at six months. Antibody levels fell by around 50% during the first two months after infection but then plateaued. The magnitude of the T cell response at six months was strongly correlated with the magnitude of the peak antibody response, the study found. Moss said the finding that the T cell response was 50% higher in those who had experienced symptoms did not necessarily mean that asymptomatic people may be more susceptible to reinfection as they may just be better at fighting off the virus without the need to generate a large immune response. The findings have implications for vaccine development. The cellular response was directed against a range of proteins from the virus, including the spike protein that is being used as a target in most vaccine studies. The study authors suggested that as T cell responses were also directed against additional nucleoprotein and membrane proteins these could also be valuable targets for future vaccines strategies. “This is promising news—if natural infection with the virus can elicit a robust T cell response then this may mean that a vaccine could do the same,” said Fiona Watt, executive chair of the Medical Research Council. Charles Bangham, chair of immunology at Imperial College London, said, “This excellent study provides strong evidence that T cell immunity to SARS-CoV-2 may last longer than antibody immunity.” He added, “These results provide reassurance that, although the titre of antibody to SARS-CoV-2 can fall below detectable levels within a few months of infection, a degree of immunity to the virus may be maintained. However, the critical question remains: do these persistent T cells provide efficient protection against re-infection?” References ↵Zuo J, Dowell A, Pearce H, et al. Robust SARS-CoV-2-specific T-cell immunity is maintained at 6 months following primary infection. BioRxiv 2020.11.01.362319v1 [Preprint]. 2 November 2020. www.biorxiv.org/content/10.1101/2020.11.01.362319v1. |
|
|
|
Post by Admin on Nov 16, 2020 20:09:42 GMT
Selective and cross-reactive SARS-CoV-2 T cell epitopes in unexposed humans Science 02 Oct 2020: Vol. 370, Issue 6512, pp. 89-94 DOI: 10.1126/science.abd3871 Preexisting immune response to SARS-CoV-2 Robust T cell responses to the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) virus occur in most individuals with coronavirus disease 2019 (COVID-19). Several studies have reported that some people who have not been exposed to SARS-CoV-2 have preexisting reactivity to SARS-CoV-2 sequences. The immunological mechanisms underlying this preexisting reactivity are not clear, but previous exposure to widely circulating common cold coronaviruses might be involved. Mateus et al. found that the preexisting reactivity against SARS-CoV-2 comes from memory T cells and that cross-reactive T cells can specifically recognize a SARS-CoV-2 epitope as well as the homologous epitope from a common cold coronavirus. These findings underline the importance of determining the impacts of preexisting immune memory in COVID-19 disease severity. Science, this issue p. 89 Abstract Many unknowns exist about human immune responses to the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) virus. SARS-CoV-2–reactive CD4+ T cells have been reported in unexposed individuals, suggesting preexisting cross-reactive T cell memory in 20 to 50% of people. However, the source of those T cells has been speculative. Using human blood samples derived before the SARS-CoV-2 virus was discovered in 2019, we mapped 142 T cell epitopes across the SARS-CoV-2 genome to facilitate precise interrogation of the SARS-CoV-2–specific CD4+ T cell repertoire. We demonstrate a range of preexisting memory CD4+ T cells that are cross-reactive with comparable affinity to SARS-CoV-2 and the common cold coronaviruses human coronavirus (HCoV)-OC43, HCoV-229E, HCoV-NL63, and HCoV-HKU1. Thus, variegated T cell memory to coronaviruses that cause the common cold may underlie at least some of the extensive heterogeneity observed in coronavirus disease 2019 (COVID-19) disease. The emergence of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in late 2019 and its subsequent global spread has led to millions of infections and substantial morbidity and mortality (1). Coronavirus disease 2019 (COVID-19), the clinical disease caused by SARS-CoV-2 infection, can range from mild, self-limiting disease to acute respiratory distress syndrome and death (2). The mechanisms underlying the spectrum of COVID-19 disease severity states and the nature of protective immunity against COVID-19 remain unclear. Studies investigating the human immune response against SARS-CoV-2 have begun to characterize SARS-CoV-2 antigen-specific T cell responses (3–8), and multiple studies have described marked activation of T cell subsets in acute COVID-19 patients (9–13). Unexpectedly, antigen-specific T cell studies performed with five different cohorts reported that 20 to 50% of people who had not been exposed to SARS-CoV-2 had significant T cell reactivity directed against peptides corresponding to SARS-CoV-2 sequences (3–7). The studies were from geographically diverse cohorts (the United States, the Netherlands, Germany, Singapore, and the United Kingdom), and the general pattern observed was that the T cell reactivity found in unexposed individuals was predominantly mediated by CD4+ T cells. It was speculated that this phenomenon might be due to preexisting memory responses against human “common cold” coronaviruses (HCoVs) such as HCoV-OC43, HCoV-HKU1, HCoV-NL63, and HCoV-229E. These HCoVs share partial sequence homology with SARS-CoV-2, are widely circulating in the general population, and are typically responsible for mild respiratory symptoms (14–16). However, the hypothesis of cross-reactive immunity between SARS-CoV-2 and common cold HCoVs still awaits experimental trials. This potential preexisting cross-reactive T cell immunity to SARS-CoV-2 has broad implications because it could explain aspects of differential COVID-19 clinical outcomes, influence epidemiological models of herd immunity (17, 18), or affect the performance of COVID-19 candidate vaccines. Epitope repertoire in SARS-CoV-2–unexposed individuals To define the repertoire of CD4+ T cells recognizing SARS-CoV-2 epitopes in previously unexposed individuals, we used in vitro stimulation of peripheral blood mononuclear cells (PBMCs) for 2 weeks with pools of 15-mer peptides. This method is known to be robust for detecting low-frequency T cell responses to allergens and bacterial or viral antigens (19, 20), including naive T cells (21). For screening SARS-CoV-2 epitopes, we used PBMC samples from unexposed subjects collected between March 2015 and March 2018, well before the global circulation of SARS-CoV-2 occurred. The unexposed subjects were confirmed to be seronegative for SARS-CoV-2 (fig. S1A). SARS-CoV-2–reactive T cells were expanded, with one pool of peptides spanning the entire sequence of the spike protein (CD4-S) and the other a nonspike “megapool” (CD4-R) of predicted epitopes from the nonspike regions (i.e., “remainder”) of the viral genome (4). In total, 474 15-mer SARS-CoV-2 peptides were screened. After 14 days of stimulation, T cell reactivity against intermediate “mesopools,” each encompassing ~10 peptides, was assayed using a FluoroSPOT assay (e.g., 22 CD4-R mesopools; fig. S2A). Positive mesopools were further deconvoluted to identify specific individual SARS-CoV-2 epitopes. Representative results from one donor show the deconvolution of mesopools P6 and P18 to identify seven different SARS-CoV-2 epitopes (fig. S2B). Intracellular cytokine-staining assays specific for interferon γ (IFN-γ) determined whether antigen-specific T cells responding to the SARS-CoV-2 mesopools were CD4+ or CD8+ T cells (fig. S2C). Results from the 44 donors/CD4-R mesopool and 40 donors/CD4-S mesopool combinations yielding a positive response are shown in fig. S2, D and E, respectively. In 82/88 cases (93.2%), the cells responding to SARS-CoV-2 mesopool stimulation were clearly CD4+ T cells, as judged by the ratio of CD4/CD8–responding cells; in four cases (4.5%), the responding cells were CD8+ T cells; and in two cases (2.3%), the responses were mediated by both CD4+ and CD8+ T cells. The fact that CD8+ T cells were rarely detected was not surprising because the peptides used in CD4-R encompassed predicted class II epitopes and the CD4-S is composed of 15-mer peptides (9- to 10-mer peptides are optimal for CD8+ T cells). Furthermore, the 2-week restimulation protocol was originally designed to expand CD4+ T cells (20). Overall, these results indicated that the peptide-screening strategy used mapped SARS-CoV-2 epitopes recognized by CD4+ T cells in unexposed individuals. A total of 142 SARS-CoV-2 epitopes were identified, 66 from the spike protein (CD4-S) and 76 from the remainder of the genome (CD4-R) (table S1). For each combination of epitope and responding donor, potential human leukocyte antigen (HLA) restrictions were inferred on the basis of the predicted HLA-binding capacity of the particular epitope for the specific HLA alleles present in the responding donor (22). Each donor recognized an average of 11.4 epitopes (range 1 to 33, median 6.5; fig. S3A). Forty of the 142 epitopes were recognized by two or more donors (fig. S3B), accounting for 55% of the total response (fig. S3C). These 142 mapped SARS-CoV-2 epitopes may prove useful in future studies as reagents for tracking CD4+ T cells in SARS-CoV-2–infected individuals and in COVID-19 vaccine trials. Epitope distribution by ORF of origin Although a broad range of different SARS-CoV-2 antigens were recognized, several of the epitopes yielding the most frequent (i.e., recognized in multiple donors) or most vigorous [i.e., the most spot-forming cells (SFCs)/106 cells] responses were derived from the SARS-CoV-2 spike antigen (table S1). We therefore assessed the overall distribution of the 142 T cell epitopes mapped among all SARS-CoV-2 proteins compared with the relative size of each SARS-CoV-2 antigen (Fig. 1, A and B). Fifty-four percent of the total positive response was associated with spike-derived epitopes [Fig. 1A; 11% for receptor-binding domain (RBD), and 44% for the non-RBD portion of spike]. Of relevance for COVID-19 vaccine development, only 20% of the spike responses were derived from the RBD region (Fig. 1A; comparing 11 versus 44%, as described above), and the RBD region accounted for only 11% of the overall CD4+ T cell reactivity (Fig. 1A). Mapped epitopes were fairly evenly distributed across the SARS-CoV-2 genome in proportion to the size of each protein (Fig. 1B; P = 0.038, r = 0.42). In addition to the strong responses directed to spike, responses were also seen for open reading frame 6 (ORF6), ORF3a, N, ORF8, and within Orf1a/b, where nsp3, nsp12, nsp4, nsp6, nsp2, and nsp14 were more prominently recognized. These mapped epitope results at the ORFeome level partially overlap with the ORFs targeted by CD4+ T cells in COVID-19 cases (4). No epitopes derived from the membrane protein (M) were identified in unexposed individuals (Fig. 1B), but M is robustly recognized by SARS-CoV-2–specific CD4+ T cell responses in COVID-19 cases (4). The lack of quality class II epitopes in M was unsurprising based on M molecular biology: M is a small protein with three transmembrane domains. Combined, the data indicate that class II epitopes are relatively broadly available across the SARS-CoV-2 genome but that SARS-CoV-2 memory CD4+ T cells preferentially target proteins highly expressed during infection, as exemplified by M and S (spike) epitope-mapping results.  Fig. 1 Characteristics of SARS-CoV-2 epitopes identified in unexposed donors. Reactivity was determined by FluoroSPOT assay after 17 days of in vitro stimulation of unexposed donor PBMCs (n = 18) with one pool of peptides spanning the entire sequence of the spike protein (CD4-S) or a nonspike “megapool” (CD4-R) of predicted epitopes from the nonspike (i.e., “remainder”) regions of the viral genome. (A) Summary of the responses as a function of the protein of origin. (B) Spearman correlation of positive responses per SARS-CoV-2 protein size. (C) Percent similarity of the identified epitopes with common cold coronavirus peptides as a function of the number of responding donors. (D) Each dot shows the reactivity of a donor-epitope combination derived from either nonspike (CD4-R) or spike (CD4-S) protein. Black bars indicate the geometric mean and geometric SD. Red indicates donor-epitope combinations with sequence identity >67% with common cold coronaviruses, and blue indicates highly reactive donor-epitope combinations (>1000 SFCs*106) with sequence identity ≤67%. In (C) and (D), statistical comparisons were performed with a two-tailed Mann–Whitney test. ***P < 0.001, ****P < 0.0001. Sequence homology of the identified SARS-CoV-2 epitopes to other common HCoVs When this epitope-mapping study was initiated, an assumption was that the in vitro T cell culture epitope mapping would reveal an epitope repertoire associated with de novo generation of responses from naïve T cells. However, while these epitope-mapping studies were in progress, we and others detected significant ex vivo reactivity against bulk pools of SARS-CoV-2 peptides (3–7) and speculated that this might reflect the presence of memory T cells cross-reactive between HCoVs and SARS-CoV-2. These other HCoVs circulate widely in human populations and are typically responsible for mild, usually undiagnosed, respiratory illnesses such as the common cold (14–16). However, there is currently a lack of experimental data addressing whether memory CD4+ T cells that are cross-reactive between SARS-CoV-2 and other HCoVs do indeed exist. We therefore next determined the degree of homology for all four widely circulating HCoVs for all 142 SARS-CoV-2 epitopes identified herein. For the analysis, we split the peptides into three groups based on immunogenicity as follows: (i) never immunogenic, (ii) immunogenic in one individual, or (iii) immunogenic in two or more individuals (Fig. 1C). There was significantly higher sequence similarity in peptides recognized by more than one individual compared with peptides recognized by a single individual or not recognized at all (P < 0.0001, two-tailed Mann–Whitney test). Additionally, almost all donors from the unexposed cohort used for the epitope screen were seropositive for three widely circulating HCoVs (HCoV-NL63, HCoV-OC42, and HCoV-HKU1) (fig. S1B). Thus, epitope homology and seropositivity data suggest that T cell cross-reactivity is plausible between SARS-CoV-2 and HCoVs already established in the human population. To select the epitope subsets to be analyzed in more detail, we plotted the T cell response magnitude of each positive epitope per donor (Fig. 1D). This analysis confirmed the dominance of the spike antigen over the epitopes derived from the remainder of the genome (P < 0.001, two-tailed Mann–Whitney test). Next, we selected two categories of SARS-CoV-2 epitopes of interest. The first category was epitopes with potential cross-reactivity from HCoVs. We initially selected the 67% arbitrary cutoff because we reasoned that a 9-mer is the epitope region involved in binding to class II (23) and that one or two residues in addition to the 9-mer core region are often required for optimal recognition (24) (Fig. 1D, red). Second, we independently filtered for any epitopes associated with high responses (top ~30%; Fig. 1D, blue). This resulted in the selection of 31 epitopes from spike (six with high homology and 25 for dominant responses) organized in a new CD4-[S31] pool. Similarly, we generated a new CD4-[R30] pool composed of 30 epitopes from the remainder of the genome (nine with high homology and 21 associated with strong responses; Fig. 1D). These epitope pools were then used for further CD4+ T cell studies. |
|
|
|
Post by Admin on Nov 17, 2020 7:34:10 GMT
Direct evidence of reactivity to HCoV epitopes homologous to SARS-CoV-2 epitopes To directly address whether reactivity against SARS-CoV-2 in unexposed donors could be ascribed to cross-reactivity against other HCoVs, we designed a peptide pool encompassing peptides homologous to CD4-R30 epitopes derived from HCoV-229E, HCoV-NL63, HCoV-OC43, HCoV-HKU1, and several other HCoVs (see the materials and methods), for a total of 129 HCoV homologs (HCoV-R129; table S2). Similarly, we synthesized a pool that encompassed peptides homologous to the SARS-CoV-2 CD4-S31 epitope pool consisting of potential epitopes derived from other HCoVs, for a total of 124 HCoV homologs (HCoV-S124; table S3). Next, we used an activation-induced marker assay (25–27) to detect virus-specific T cells in a new set of unexposed donors not used for the epitope identification studies (Fig. 2A and table S4) and a set of convalescent COVID-19 patients (table S5). We detected significant ex vivo CD4+ T cell responses against the SARS-CoV-2 nonspike (CD4-R) and spike (CD4-S) peptides compared with the negative control [dimethyl sulfoxide (DMSO)] (Fig. 2, B and C; P < 0.0001 and P < 0.0001, respectively, two-tailed Mann–Whitney test). These responses were increased in COVID-19 cases compared with unexposed subjects (Fig. 2D; P = 0.0015 and P = 0.0022, respectively, two-tailed Mann–Whitney test), as previously reported (4). In the unexposed subjects, significant frequencies of CD4+ T cells were detected against the CD4-R30 and CD4-S31 SARS-CoV-2 epitope pools compared with the negative control (Fig. 2B; P = 0.0063 and P = 0.0012, respectively, two-tailed Mann–Whitney test). Significant CD4+ T cell reactivity was also seen against the corresponding HCoV-R129 and HCoV-S124 pools of matching homologous peptides from other HCoVs (Fig. 2D; P < 0.0001 and P < 0.0001, two-tailed Mann–Whitney test). Detection of CD4+ T cells with peptide pools selected on the basis of homology was consistent with the hypothesis that cross-reactive CD4+ T cells between SARS-CoV-2 and other HCoVs exist in many individuals.  Fig. 2 CD4+ T cells in SARS-CoV-2–unexposed and recovered COVID-19 patients against HCoV epitopes homologous to SARS-CoV-2 epitopes. (A) Example of flow cytometry gating strategy for antigen-specific CD4+ T cells based on activation-induced marker assays (OX40+ and CD137+ double expression) after stimulation of PBMCs with HCoV or SARS-CoV-2 peptides. (B to D) Antigen-specific CD4+ T cells measured as the percentage of activation-induced marker assay–positive (OX40+CD137+) CD4+ T cells after stimulation of PBMCs with HCoV epitopes homologous to SARS-CoV-2 epitopes. Samples were derived from SARS-CoV-2–unexposed donors (n = 25) and recovered COVID-19 patients ( n = 20). Black bars indicate the geometric mean and geometric SD. Each dot is representative of an individual subject. Statistical pairwise comparisons [(B) and (C)] were performed with the Wilcoxon test. P values related to comparisons with the DMSO controls are listed at the bottom of the graphs, and any significant P values related to intergroup comparisons are listed on top of the graphs. Statistical comparisons across cohorts were performed with the Mann–Whitney test (D). See also figs. S5 and S6. Reactivity against CD4-R30 and CD4-S31 (Fig. 2D; P = 0.0008 and P = 0.0026, respectively), but not against HCoV-R129 and HCoV-S124, was increased in COVID-19 cases compared with unexposed individuals (Fig. 2C). Thus, preexisting CD4+ T cell reactivity to HCoV epitopes is modulated by COVID-19 and exposure to cross-reactive SARS-CoV-2 epitopes in COVID-19. These data from COVID-19 cases do not support the hypothesis that the HCoV exposure might induce an original antigenic sin phenomenon, impairing subsequent T cell responses to SARS-CoV-2 epitopes (28, 29), at least for COVID-19 cases of average disease severity. Next, we examined the ex vivo memory phenotype of the T cells responding to the various epitope megapools. Results from one representative unexposed donor are shown in Fig. 3A. Responding cells in unexposed donors were predominantly found in the effector memory CD4+ T cell population (CD45RAnegCCR7neg), followed by the central memory T cells (CD45RAnegCCR7pos) (30) (Fig. 3, A, B, and D). Comparable patterns of effector and central memory cells were observed among the antigen-specific CD4+ T cells detected in the COVID-19 cases (Fig. 3, C and D). The CD4+ T cells in unexposed donors that recognize SARS-CoV-2 epitopes and epitopes from other HCoVs have a memory phenotype. Overall, these data are consistent with the SARS-CoV-2–reactive CD4+ T cells in unexposed subjects being HCoV-specific memory CD4+ T cells with cross-reactivity to SARS-CoV-2.  Fig. 3 Phenotypes of antigen-specific CD4+ T cells from SARS-CoV-2–unexposed and recovered COVID-19 patients responding to HCoV epitopes homologous to SARS-CoV-2 epitopes. (A) Example of flow cytometry gating strategy for antigen-specific CD4+ T cell subsets after overnight stimulation of PBMCs with HCoV or SARS-CoV-2 peptides ex vivo. (B and C) Phenotype of antigen-specific CD4+ T cells (OX40+CD137+) responding to the indicated pools of SARS-CoV-2 and HCoV epitopes in unexposed subjects and recovered COVID-19 patients. Data are shown as mean ± SD. Each dot represents an individual subject. Statistical pairwise comparisons in (B) and (C) were performed with the Wilcoxon test. (D) Overall averages of antigen-specific CD4+ T cell subsets detected in unexposed subjects and recovered COVID-19 patients. See also fig. S5. Identification of SARS-CoV-2 epitopes cross-reactive with other common HCoVs The epitopes derived from the CD4-R30 and CD4-S31 pools were used to generate short-term T cell lines derived by stimulation of PBMCs from unexposed subjects. PBMCs were stimulated with an individual SARS-CoV-2 cognate epitope demonstrated to be recognized by T cells from that subject (Fig. 1 and table S1). Overall, T cell lines could be derived that were specific for a total of 42 SARS-CoV-2 epitopes. These T cell lines were next tested for cross-reactivity against various coronavirus homologs, analogous to an approach previously successful in flavivirus studies (31). Cross-reactivity between SARS-CoV-2 epitope recognition and other HCoV epitope recognition was detected for 10/42 (24%) of the T cell lines (Fig. 4, A to J). Cross-reactivity was associated with epitopes derived from SARS-CoV-2 spike, N, nsp8, nsp12, and nsp13. In three cases, HCoV analogs were better antigens than the SARS-CoV-2 peptide, suggesting that they may be the cognate immunogen (Fig. 4, E, I, and J). One SARS-CoV-2 spike epitope was tested in two different donors with similar findings, suggesting that HCoV cross-reactivity patterns are recurrent across individuals. Non–cross-reactive SARS-CoV-2 T cell lines are also shown (Fig. 4, K to L, and fig. S4). It is possible that cross-reactivity to these epitopes might be detected if T cell lines from additional individuals were to be tested. In addition, these epitopes might be homologous to some other, as yet unidentified viral sequence or be recognized by cognate naive T cells expanding in the in vitro culture (32). In addition, only 3/18 cases of strong response epitopes (defined in Fig. 1D) were cross-reactive compared with 4/5 of weaker epitopes (P = 0.02, Fisher’s exact test). To further demonstrate that the cross-reactive responses in unexposed donors are indeed derived from memory T cells, we stimulated purified memory and naïve CD4+ T cells with the CD4-[S31] epitope pool. After 14 days, we detected responses to the CD4-[S31] peptide pool from cultures of memory CD4+ T cells but not naïve CD4+ T cells (fig. S8). These data demonstrate that memory CD4+ T cells recognizing common cold coronaviruses including HCoV-OC43, HCoV-HKU1, HCoV-NL63, and HCoV-229E can exhibit substantial cross-reactivity to the homologous epitope in SARS-CoV-2.  Fig. 4 Cross-reactivity of SARS-CoV-2 and homologous HCoV peptides. Twelve short-term cell lines were generated using specific SARS-CoV-2 donor-epitope combinations selected on the basis of the primary screen. After 14 days of in vitro expansion, each T cell line was tested with the SARS-CoV-2 epitope used for stimulation and peptides corresponding to analogous sequences from other HCoVs at six different concentrations (1, 0.1, 0.01, 0.001, 0.0001, and 0.00001 μg/ml). SFCs/106 PBMCs are plotted for T cell lines stimulated with each peptide. See also fig. S7. Next we examined, for each SARS-CoV2:HCoV epitope pair, the degree of amino acid sequence homology and any relationship between homology and T cell cross-reactivity, considering different ranges of potentially relevant homology. Only 1% (1/99) of peptide pairs with 33 to 40% homology were cross-reactive. In the 47 to 60% epitope homology range, we observed cross-reactivity in 21% of cases (7/33). Epitope homology ≥67% was associated with cross-reactivity in 57% of cases (21/37; P = 0.0001 or P = 0.0033 by two-tailed Fisher’s exact test compared with the 33 to 40% range epitopes or the 47 to 60% range, respectively). A relationship was observed between epitope homology and CD4+ T cell cross-reactivity. The data demonstrated that the arbitrary selection used as described in Fig. 1D was indeed supported by the experimental data. Thus, ~67% amino acid homology appears to be a useful benchmark for consideration of potential cross-reactivity between class II epitopes. In summary, we have identified more than 140 human T cell epitopes derived from across the genome of SARS-CoV-2. We provide direct evidence that numerous CD4+ T cells that react to SARS-CoV-2 epitopes actually cross-react with corresponding homologous sequences from any of the many different commonly circulating HCoVs, and that these reactive cells are largely canonical memory CD4+ T cells. These findings of cross-reactive HCoV T cell specificities are in stark contrast to HCoV-neutralizing antibodies, which are HCoV species specific and did not show cross-reactivity against SARS-CoV-2 RBD (33–35). On the basis of these data, it is plausible to hypothesize that preexisting cross-reactive HCoV CD4+ T cell memory in some donors could be a contributing factor to variations in COVID-19 patient disease outcomes, but at present this is highly speculative (36). |
|
|
|
Post by Admin on Nov 17, 2020 23:49:38 GMT
Epidemiology, Genetic Recombination, and Pathogenesis of Coronaviruses Shuo Su Gary Wong Weifeng Shi Wenjun Liu Yuhai Bi George F. Gao Published:March 21, 2016 DOI:https://doi.org/10.1016/j.tim.2016.03.003 Human coronaviruses (HCoVs) were first described in the 1960s for patients with the common cold. Since then, more HCoVs have been discovered, including those that cause severe acute respiratory syndrome (SARS) and Middle East respiratory syndrome (MERS), two pathogens that, upon infection, can cause fatal respiratory disease in humans. It was recently discovered that dromedary camels in Saudi Arabia harbor three different HCoV species, including a dominant MERS HCoV lineage that was responsible for the outbreaks in the Middle East and South Korea during 2015. In this review we aim to compare and contrast the different HCoVs with regard to epidemiology and pathogenesis, in addition to the virus evolution and recombination events which have, on occasion, resulted in outbreaks amongst humans. A Brief Introduction to Human and Animal Coronaviruses Coronaviruses (CoVs), of the family Coronaviridae, are enveloped viruses with a single-strand, positive-sense RNA genome approximately 26–32 kilobases in size, which is the largest known genome for an RNA virus [1]. The term ‘coronavirus’ refers to the appearance of CoV virions when observed under electron microscopy, in which spike projections from the virus membrane give the semblance of a crown, or corona in Latin [2, 3]. All coronaviruses share similarities in the organization and expression of their genome, in which 16 nonstructural proteins (nsp1 through nsp16), encoded by open reading frame (ORF) 1a/b at the 5′ end, are followed by the structural proteins spike (S), envelope (E), membrane (M), and nucleocapsid (N), which are encoded by other ORFs at the 3′ end. CoVs are separated into four genera based on phylogeny: alpha-CoV (group 1), beta-CoV (group 2), gamma-CoV (group 3) and delta-CoV (group 4) (http://ictvonline.org/proposals/2008.085-122 V.v4.Coronaviridae.pdf). Within the beta-CoV genus, four lineages (A, B, C, and D) are recognized. Distinct from other beta-CoV lineages, lineage A viruses also encode a smaller protein called hemagglutinin esterase (HE), which is functionally similar to the S protein [4].  Figure 1 Global Distribution of Human Coronaviruses. (A) Green, blue, brown, and purple represent the global distribution of the NL63, HKU1, OC43, and 229E human coronaviruses, respectively. (B) Red and yellow represent the global distribution of MERS-CoV and SARS-CoV, respectively. Since the early 1970s, a variety of pathological conditions in domestic animals have been attributed to CoV infections [5]. With the exception of infectious bronchitis virus (IBV), which causes avian infectious bronchitis in chickens [6], canine respiratory coronavirus (CRCoV), which causes respiratory disease in dogs [7], and mouse hepatitis virus (MHV), which can cause a progressive demyelinating encephalitis in mice [3, 8], other CoV infections typically result in gastrointestinal symptoms. For instance, transmissible gastroenteritis coronavirus (TGEV) [9, 10], bovine coronavirus (BCV) [11], feline coronavirus (FCoV) [12], canine coronavirus (CCoV) [13], and turkey coronavirus (TCV) [14] are known to cause enteritis in their respective hosts [1]. In humans, CoV infections primarily involve the upper respiratory tract and the gastrointestinal tract, and vary from mild, self-limiting disease, such as the common cold, to more severe manifestations, such as bronchitis and pneumonia with renal involvement [15]. The first human coronavirus (HCoV) was isolated during 1965 from the nasal discharge of patients with the common cold and termed B814 [16]. Currently, six different CoV strains are known to infect humans. These include: HCoV-229E (229E), HCoV-OC43 (OC43), severe acute respiratory syndrome coronavirus (SARS-CoV), HCoV-NL63 (NL63), HCoV-HKU1 (HKU1), and Middle East respiratory syndrome coronavirus (MERS-CoV) [17]. 229E and OC43 are the prototype viruses from the two main HCoV lineages (Alpha and Beta, respectively) that cause 15–29% of all common colds, and are the best characterized [18]. SARS-CoV is the aetiological agent that was behind an outbreak of severe respiratory disease through China during 2002–2003 [19], and MERS-CoV is the pathogen responsible for an ongoing outbreak of severe respiratory disease centered in the Middle East since 2012 [20]. In this review, we first compare and contrast the epidemiology and pathogenesis of the known CoVs infecting humans, in which CoVs that cause mild to severe disease in humans are newly emerged from a zoonotic source. Second, we describe the ecology of CoVs, and highlight evidence for viral recombination between the different CoVs within animal populations, which may result in the generation of novel CoVs that are transmissible and lethal to humans. Finally, we summarize the phylogeny, evolution, and genetic recombination of CoVs in detail. Epidemiology and Pathogenesis of HCoV The epidemiology and pathogenesis of each HCoV are discussed within the following sections and summarized in Table 1 and Figure 1, respectively. In general, 229E, OC43, and NL63 are distributed globally (Figure 1A) and tend to be transmitted predominantly during the winter season in temperate-climate countries [21], while NL63 showed a spring–summer peak of activity from a study in Hong Kong [22]. |
|











