|
|
Post by Admin on Mar 22, 2022 22:09:28 GMT
Principal Component Analysis
We performed principal component analysis (PCA) with the smartpca program of the EIGENSOFT package (Patterson et al., 2006) using the default settings with additional parameters: lsqproject: YES and numoutlieriter: 0. Population data of modern East Asia were used to reconstruct the genetic background of PCA, in which modern samples were mainly sampled from Altaic, Sino-Tibetan, Hmong-Mien, Austronesian, Austroasiatic, and Tai-Kadai language families. Ancient genomes were projected onto the first two components. The projected ancient populations included eight individuals from Nepal (Jeong et al., 2016) (Chokhopani, Samdzong, and Mebrak cultures), eighty-four samples from the Yellow River (Ning et al., 2020; Yang et al., 2020; Wang C. C. et al., 2021), Amur River and West Liao River in the coastal and inland northern East Asia (including Houli, Yangshao, Longshan, Qijia, Hongshan, Yumin and other cultures), fifty-eight individuals (Ning et al., 2020; Yang et al., 2020; Wang C. C. et al., 2021) belonging to Tanshishan and other cultures in the coastal southeast East Asia (Fujian and Taiwan).
FST Calculation and TreeMix Analysis
We used the Plink 1.9 and an in-house script to estimate the pairwise FST genetic distance (Purcell et al., 2007) among 82 modern populations with a sample size large than five. We also calculated FST values among 31 ancient populations. We ran TreeMix v.1.13 (Pickrell and Pritchard, 2012) with migration events ranging from 0 to 8 to construct the topology among eastern Eurasians with the maximum likelihood tree.
ADMIXTURE Analysis
We carried out the model-based clustering analysis using the ADMIXTURE (v.1.3.0) (Alexander et al., 2009) after pruning SNPs with a strong linkage disequilibrium via the PLINK v.1.9 (Chang et al., 2015) with the parameters of – indep-pairwise 200 25 0.4. We ran ADMIXTURE with the 10-fold cross-validation (−cv = 10). The predefined number of ancestral populations ranging from K = 2 to K = 20 with 100 bootstraps and different random seeds were used. We chose the best-fitted model with the minimum cross-validation errors. The smallest cross-validation error was obtained (0.4176) when we used 10 predefined ancestral sources.
F-Statistics and Admixture Modeling Graph
We conducted two different forms of the three-population tests using the qp3Pop program implemented in the ADMIXTOOLS (Reich et al., 2009; Patterson et al., 2012). Outgroup-f3-statistics were performed in the form of f3(Reference Eurasians, targeted Tibetans; Mbuti) to assess the shared genetic drift between our focused Tibetans and their reference populations. A central African population Mbuti was used as the outgroup. Admixture-f3(Surrogate population1, Surrogate population2; Targeted populations) were performed to test whether our targeted population was an admixture of two sources related to our used surrogate populations. Negative f3-values with a Z-score smaller than −3 indicated that two source populations were admixed to form the targeted populations. Four-population comparisons were conducted using qpDstat programs implemented in the ADMIXTOOLS (Reich et al., 2009; Patterson et al., 2012) with the additional parameter (f4: YES) in three different forms. The first one was conducted in the form of f4(Tibetan1, Tibetan2; Eurasian reference, Mbuti) to test whether two Tibetans form one clade relative to the used Eurasian reference. Non-statistically significant f4 values showed two left populations formed one clade. Other two f4-statistics in the forms f4(Eurasian, Source; Eurasian2, Mbuti) and f4(Eurasian1, Eurasian2; Source, Mbuti) were conducted to examine whether the used ancestral source shared more alleles with one of the Eurasians compared with others. We assessed standard errors using the weighted block jackknife approach. We next used the qpGraph program implemented in the ADMIXTOOLS (Reich et al., 2009; Patterson et al., 2012) to reconstruct the deep population history of modern Tibetans and other modern and ancient East Asians based on the combined results of the f2, f3 and f4-statistics. The absolute Z-scores smaller than 3 indicated better-fitted models.
Streams of Ancestry and Inference of Mixture Proportions
We used the qpWave/qpAdm programs implemented in the ADMIXTOOLS (Haak et al., 2015) to estimate mixture coefficient and corresponding standard errors according to a basic set of outgroup populations: Mbuti, Ust_Ishim, Russia_Kostenki14, Papuan, Australian, Mixe, MA1, and Mongolia_N_East.
|
|
|
|
Post by Admin on Mar 23, 2022 19:34:22 GMT
Results
Close Genetic Affinity Between Ancient/Modern Tibetans With NEAs
Descriptive analyses of PCA and ADMIXTURE were first used to provide an overview of the genetic structure. All modern Tibetans and Neolithic-to-historic East Asians were grouped in the East-Asian genetic cline along with the second component in the Eurasian-PCA. To focus on the genetic variations of East Asians, we constructed East-Asian-PCA among 106 modern populations (Figure 1B) and found that modern East Asians grouped into four genetic clines or clusters: Mongolic/Tungusic genetic cline consisting of populations from northeast Asia; south-China/Southeast-Asian genetic cluster comprising of Austronesian, Austroasiatic, Tai-Kadai, and Hmong-Mien speakers; Sinitic-related north-to-south genetic cline, and Tibeto-Burman cluster, which were consistent with the linguistic/geographical divisions. Tibetan populations were grouped and showed a relatively close relationship with some of the Mongolic/Tungusic speakers in northern China, and they were also grouped closely with northern Han and other lowland Tibeto-Burman speakers. Focused on the population substructures within Tibetans, we further observed three different sub-clusters: the high-altitude Tibet-Ü-Tsang cluster (Lhasa, Nagqu, Shannan and Shigatse), Gan-Qing-Ando cluster in northeastern TP (Xunhua, Gangcha and Gannan) and Tibetan-Yi-corridor cluster (Chamdo, Xinlong, Yajiang and Yunnan), which were also consistent with the geographical positions of sampling places and cultural backgrounds.
We subsequently explored the patterns of genomic affinity between ancient populations and modern East Asians by projecting all included ancient individuals (243 eastern Eurasian ancients) onto the genetic background of modern populations. Here, we found four ancient population genetic clusters. Neolithic-to-historic SEAs (including Hanben and Gongguan from Taiwan, Late-Neolithic mainland Tanshishan and Xitoucun people) grouped together and clustered with modern Tai-Kadai, Austronesian, and Austroasiatic speakers. Neolithic-to-Iron Age NEAs (both coastal Shandong Houli and inland Yangshao, Longshan, and Qijia people) grouped together and were projected closely to the juncture position of three main East Asian genetic lines and the northmost end of Han Chinese genetic cline. We observed a close genetic relationship between early Neolithic Houli individuals associated with the main subsistence strategy of hunter-gathering and the Henan Middle/Late-Neolithic Yangshao/Longshan farmers, which indicated the genetic continuity in the Neolithic transition from foragers to millet farmers in the early Neolithic northern China. We also identified the subtle genetic differences within these Neolithic-to-Iron Age individuals from northern China. These Shandong Houli individuals were localized closely with modern Mongolic-speaking Baoan, Tu, Yugur, and Dongxiang, while the early Neolithic Xiaogao individuals were posited closely with modern Tungusic-speaking Hezhen and Xibo. All Shandong Neolithic ancient populations were localized distantly from the modern Shandong Han Chinese and shifted to modern northern Chinese minorities, which indicated that modern northern Han received additional gene flow from SEA related ancestral lineage or ancient Houli individuals harbored more Siberian-associated ancestry. Late-Neolithic Longshan individuals (Pingliangtai, Haojiatai, and Wadian) and Bronze/Iron Age individuals (Haojiatai, Jiaozuoniecun, and Luoheguxiang) in Henan province were grouped together and shifted to the Han Chinese genetic cline and partially overlapped with Han Chinese from Shanxi and Shandong provinces. This observed genetic similarities among the Late Neolithic to present-day NEAs from the Central Plain (Henan, Shanxi, and Shandong) indicated a genetic stability in the core region of Chinese civilization since the Late-Neolithic period. Middle-Neolithic Yangshao individuals (Xiaowu and Wanggou) in Henan province grouped with some of the Wuzhuangguoliang_LN individuals collected from Shaanxi province and were shifted to more northern modern minorities. The inland Middle/Late-Neolithic NEAs from Shaanxi (Shimao), Inner Mongolia (Miaozigou) and upper Yellow River (Lajia and Jinchankou) clustered together and were shifted toward modern Tibetans and ancient Nepal samples (Mebrak, Samdzong and Chokhopani).
For ancient populations from the West Liao River, three genetic-affinity clusters could be identified in the projected PCA results: northern cluster (Haminmangha_MN and Longtoushan_BA_O) showed a genetic affinity with Shamanka and Mongolia Neolithic people; middle Hongshan cluster was localized between Mongolia minorities and modern Gangcha Tibetan; southern cluster (Upper Xiajiadian Longtoushan_BA and Erdaojingzi_LN) possessed close relationship with the Yellow River farmers, which suggested that both Neolithic ancients associated with steppe pastoralists from Mongolia Plateau and millet farmers from Yellow River Basin had participated in the formation of the Late Neolithic and subsequent populations in the West Liao River Basin. These population movements, interactions, and admixture processes have recently been fully elucidated by Ning et al. (2020). Here, we observed that the Late Neolithic populations in the southern cluster were localized between the coastal early Neolithic NEAs and inland Neolithic Yangshao and Longshan individuals, which indicated that millet farmers from the middle/lower Yellow Rivers (Henan and Shandong) had played an important role in the formation of Hongshan people or their descendants via both inland and coastal northward migration routes. For ancient populations from Mongolia Plateau, Russia Far East, Trans-Baikal-Region, and Amur River Basin, all included forty-six individuals (Neolithic-to-Bronze Shamanka, Mongolian, DevilsCave, Boisman, and others) clustered closely to modern Tungusic language speakers (Nanai and Ulchi) and also to some Mongolic speakers. Jomon individuals were grouped together in the intermediate position between the northern Russian coastal Neolithic people and southern Iron Age Taiwan Hanben and coastal Neolithic SEAs, but localized far away from modern Japanese populations.
Patterns of genetic relationship revealed from the top two components (extracting 1.42% variation: PC1: 1.03% and PC2: 0.39%) showed a genomic affinity between modern Tibetans, ancient Nepal populations, and ancient/modern East Asians and Siberians. To further explore the genetic structure and corresponding population relationships, we estimated the ancestry composition and cluster patterns according to the model-based maximizing likelihood clustering algorithm (Figure 1C and Supplementary Figure 1). We observed two northern and two southern East Asian dominant ancestries. The coastal NEA ancestry (light green) maximized in Neolithic northeast Asians (Boisman_MN, Wuqi_EN, Zhalainuoer_EN, Mongolia_N_North, Mongolia_N_East, DevilsCave_N and Shamanka_EN) and modern Tungusic speakers (Ulchi and Nanai). This light green ancestry also existed in the Bronze Age to present-day populations from northeastern China and Russia, and reached at a high proportion in the coastal Early Neolithic NEAs from Shandong. The other type of northern ancestry was enriched in modern highland Tibetans and Qijia culture-related Late Neolithic Lajia and Jinchankou populations, which also maximized in Nepal Bronze Age to historic individuals and ancient NEAs, as well as the lowland modern Sino-Tibetan speakers, inland Hmong-Mien and Tai-Kadai language speakers. We named this Tibetan-associated ancestry as inland NEA ancestry, which was the direct indicator of the close genetic affinity between Tibetan and ancient/modern NEAs. Dark green ancestry was enriched in the coastal Early Neolithic SEAs, Iron-Age Hanben, and modern Austronesian Ami and Atayal. Therefore, we referred to this dark green component as the coastal SEA ancestry. The blue component maximized in LaChi samples as the counterpart of the coastal ancestry that was widely distributed in Hmong-Mien and Tai-Kadai-speaking populations. This blue inland SEA ancestry also existed in the lowland Tibetans with a relatively high proportion in all Kham and Ando Tibetans except for Chamdo Tibetans. Besides, we found that Tibetans collected from the northeast TP harbored more coastal NEA ancestry. Some Austroasiatic-associated dark pink ancestry maximized in Mlabri also appeared in Yajiang, Xinlong Kham, and Xunhua and Gannan Ando Tibetans. The Steppe pastoralist-like red component was enriched in Bronze Age Afanasievo and Yamnaya, which was also identified in Qinghai and Gansu Ando Tibetans.
|
|
|
|
Post by Admin on Mar 23, 2022 21:39:20 GMT
Population Differentiation Between Highland and Lowland East Asians and Substructure Among Tibetans To further explore the genetic differentiation between eleven modern Tibetan populations and ancient/modern reference populations, we first calculated the pairwise FST genetic distances among 82 modern populations (Supplementary Table 1, modern dataset) and 32 ancient/modern populations (Supplementary Table 2, ancient dataset). We found a strong genetic affinity among geographically close populations. As shown in Supplementary Figures 2, 3, the high-altitude Tibetans from the south (Shigatse and Shannan), central (Lhasa), north or northeast (Nagqu and Chamdo) of Tibet Autonomous Region had the smallest FST genetic distances with their geographical neighbors, followed by lowland Ando Tibetans from the northeastern TP (Qinghai and Gansu) and the Kham Tibetans from the southeastern region of the TP (Sichuan and Yunnan) and other Tibeto-Burman-speaking populations (Qiang, Tu and Yi). For Ando Tibetans from the Ganqing region, Gangcha Tibetan harbored a close genetic affinity with northern or northeastern Tibet Tibetans (Chamdo and Nagqu) with the smallest FST genetic distances, followed by Qiang, Yugur, and Tu or other geographically close Tibetans (Supplementary Figure 4). Different patterns were observed in Gangcha and Xunhua Tibetans, which showed the closest relationship with each other, and then followed by Tu and Yugur. We also found relatively small genetic distances between Tibetans (Gannan and Xunhua) and the Turkic-speaking Kazakh population, suggesting a western Eurasian affinity of Tibetans from the northeastern region of the TP relative to the Tibetans from the central region. Supplementary Figure 5 presented the patterns of genetic differentiation between lowland Kham Tibetans and their reference populations. We found that Yajiang and Xinlong Tibetans from Sichuan province harbored a close genetic affinity with the geographically close populations (Tibetan, Qiang, Yugur and Tu). Yunnan Tibetans had the smallest genetic distance with Gangcha and Chamdo Tibetans, followed by Qiang, Yi, and Tu. Among Tibetans and Neolithic to Iron Age East Asians (Supplementary Figure 6), we also found Iron Age Hanben population from Taiwan and some southern Siberian ancients showed a closer relationship with modern Tibetans relative to other ancient East Asians. We should note there might be statistical bias in the FST-based analyses because of the different sample sizes in different populations. Phylogenetic relationships were further reconstructed based on the genetic variations of modern Eurasian populations and ancient eastern Eurasians using TreeMix software based on genetic distances. As shown in Figure 2, a phylogenetic tree with no migration events showed that modern populations from similar language families tended to cluster into one clade. Altaic-speaking (Turkic and Mongolic) populations clustered with Uralic speakers. Southern Austronesians first clustered with Tai-Kadai speakers and then clustered with Hmong-Mien and Austroasiatic speakers. Tibetans first clustered with each other, especially for high-altitude Ü-Tsang Tibetans, and then clustered with the lowland East Asians. The observed geographical affinity showed that the genetic differentiation between modern highland Tibetans and lowland East Asians could be identified although they both derived majority of their ancestry from Neolithic Yellow River farmers. We further analyzed the population splits and gene flow events between modern Tibetans and 26 ancients from eastern Eurasia (except for Anatolia_N from Near East) with three predefined admixture events. Modern Tibetans (except for Gannan and Xinlong Tibetans) first clustered with the highland Nepal ancients and then clustered with the lowland Neolithic-NEAs and Neolithic to Bronze Age southern Siberians. The cluster patterns also showed a distant relationship between northern and southern East Asians, as well as the genetic distinction between the highland ancient/modern Tibetans and the lowland SEAs, which further provided evidence for some special connections or close genetic relationships between Tibetans and NEAs. FIGURE 2 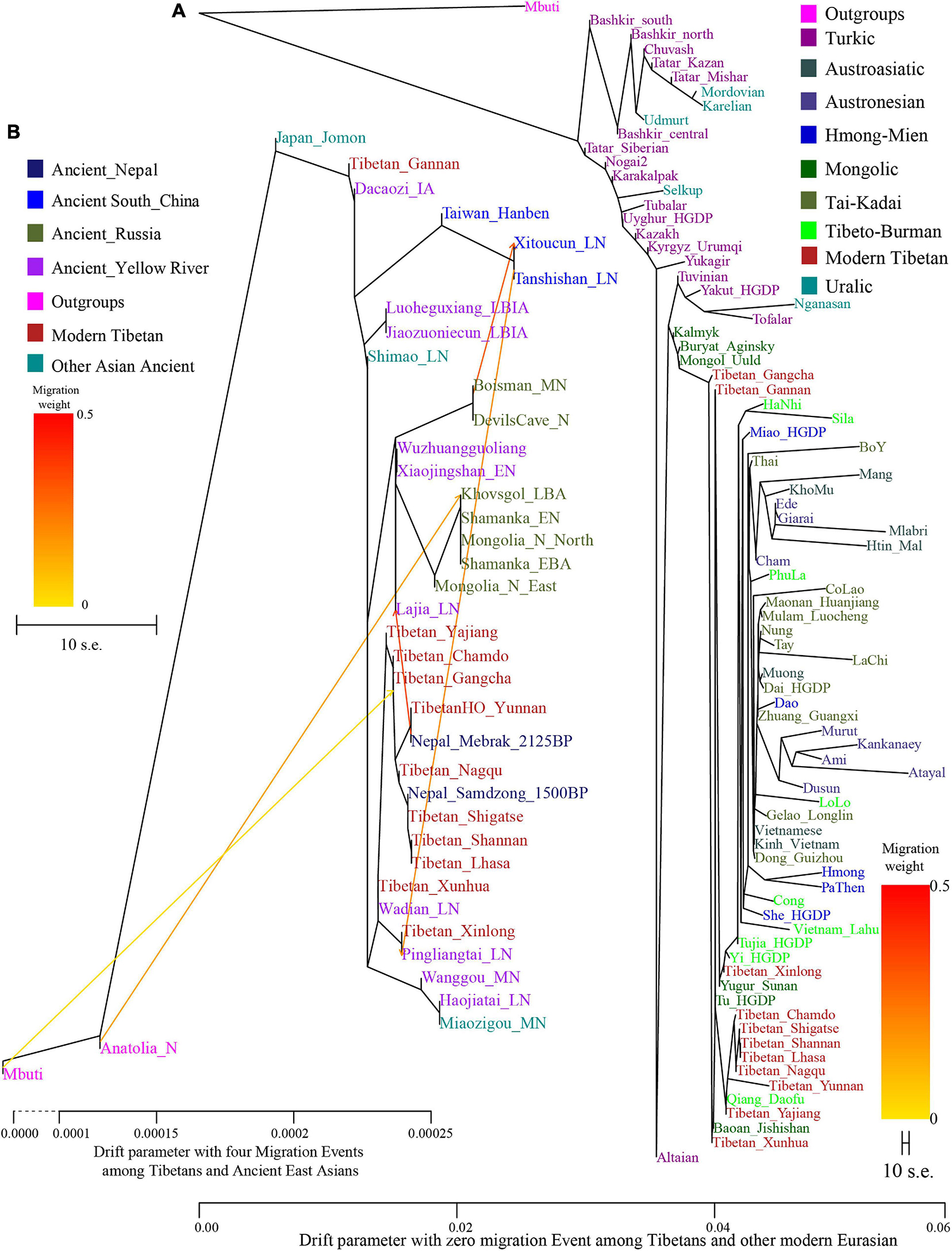 Figure 2. Maximum likelihood phylogeny reconstruction based on the genetic variation from both modern Tibetan and Eurasian modern reference populations. (A), modern Tibetan and Neolithic-to-historic East Asian (B). Mbuti was used as the root. Focused on the phylogenetic relationship among all modern populations, we used the patterns of genetic relationship with zero migration events. And evaluating the evolutionary history among modern Tibetan and ancient Chinese, we included three migration events. To better present our result, the drift branch length of Mlabri was shortened as the third of the truth drift branch length due to the strong genetic drift that occurred in Mlabri. Genetic affinity was further evaluated via the outgroup-f3-statistics in the form f3(modern Tibetans, ancient/modern Eurasians, Mbuti). We found a close genetic affinity within Tibetan populations and identified the genetic connection between Tibetan and Han Chinese. Among 184 modern populations (Figure 3 and Supplementary Table 3), the top allele sharing population for each Tibet Tibetan was another geographically close Tibetan group. Shannan Tibetan shared the most alleles with Lhasa/Shigatse/Nagqu Tibetans, and similar patterns of population affinity were identified in southern Shigatse Tibetan and central Lhasa Tibetan. However, Nagqu Tibetan shared the most alleles with the northeastern Chamdo Kham Tibetan (followed by Tibetan-Burman-speaking Qiang from Sichuan province and other Tibetans or Sherpa), and these patterns of genetic affinity were consistent with that of Chamdo Tibetan and others. Following the genomic affinity within Tibetans, we also found that these five Tibet Tibetans shared the strongest genetic affinity with the lowland Han Chinese, which was consistent with the common origin of Sino-Tibetan speakers from the Upper and Middle Yellow River Basin (YRB). For Sichuan/Yunnan lowland Kham Tibetans, Xinlong Tibetan shared the most genetic drift with Han Chinese and other lowland Tibeto-Burman-speaking Qiang and Tujia. Being different from Xinlong Tibetan, geographically close Yajiang and Yunnan Tibetans shared the most genetic drifts with Qiang and geographically close Tibetans (Chamdo and Xinlong), followed by Han Chinese and other Tibetans. These lowland Han/SEA affinities of Kham Tibetans suggested that lowland Tibetans from southwestern China harbored ancestry that derived from SEAs via the massive migrations and admixtures in the prehistoric/historic times. Gangcha Ando Tibetan not only showed the genetic affinity with Sinitic and Tibeto-Burman speakers but also showed the signals of genetic affinity with Turkic-speaking populations. Allele sharing results from Gannan and Xunhua Tibetans showed that the Han Chinese groups shared the most ancestry components with them. |
|
|
|
Post by Admin on Mar 24, 2022 1:06:58 GMT
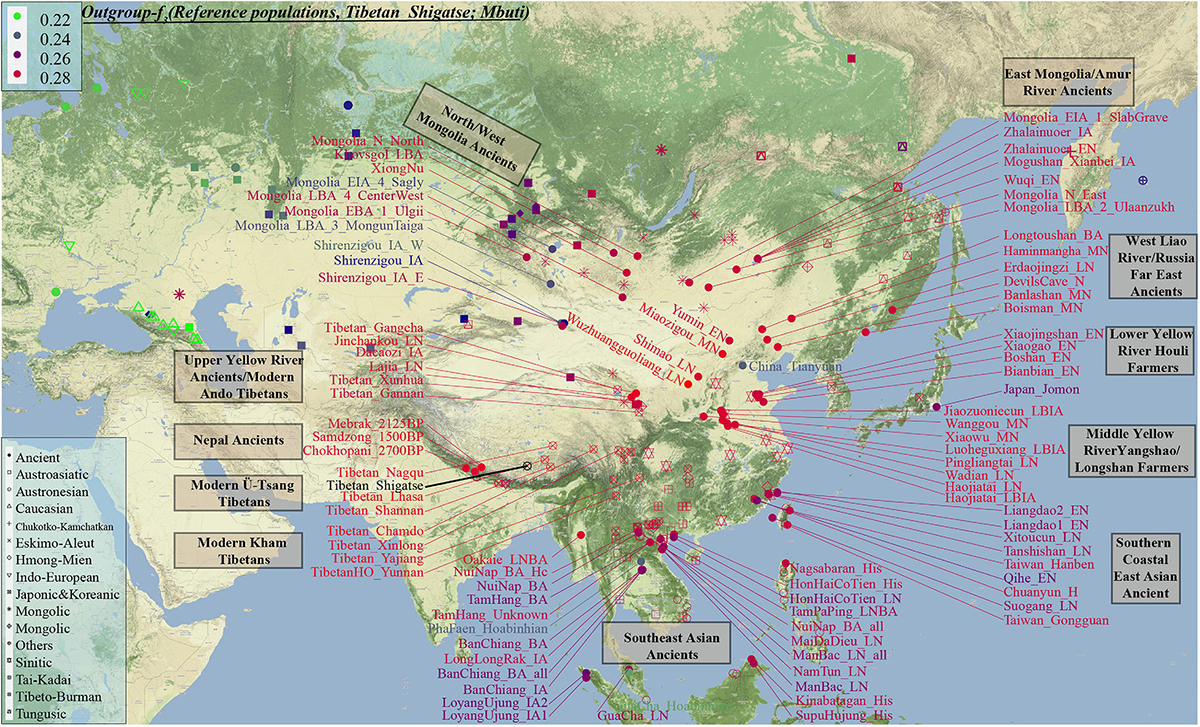 Figure 3. The genomic affinity between our Shigatse Tibetan populations and other modern and ancient spatial-temporally different eastern Eurasian populations. The red color denoted a stronger genetic affinity with Shigatse Tibetans, and the blue color showed a lower genetic affinity. Levels of allele sharing between modern Tibetans and 106 Paleolithic to historic Eurasian ancients (including 33 populations from Russia, 41 from China, 29 from Mongolia, and 3 from Nepal) inferred from the outgroup-f3-statistics showed that modern Tibetans had a clear connection with ancient Neolithic to Iron Age NEAs, which was consistent with the patterns observed in the PCA, FST, ADMIXTURE and modern population-based affinity estimations (Supplementary Table 3). Middle-altitude Chamdo Tibetan shared the most genetic drift with Neolithic Wuzhuangguoliang_LN (low coverage samples), upper Yellow River Late Neolithic farmers (Jinchankou and Lajia, which are the represented typical source populations for Qijia culture), followed by Iron Age Dacaozi people, Shimao people from Shaanxi, Middle-Neolithic Banlashan associated with Hongshan culture in northern China and other NEAs from lower and middle YRB (Supplementary Figure 7). Neolithic people from Russia and Mongolia and Bronze to historic Nepal ancients showed a relatively distant genetic relationship with modern Chamdo Tibetan (Supplementary Table 3). Different from the pattern of Chamdo Tibetan, southern and central Ü-Tsang Tibetans showed increased ancestry associated with Nepal ancient people, and northern Nagqu Tibetan showed the intermediate trend of population affinity with 2700-year-old Chokhopani. As showed in Supplementary Figures 8, 9, lowland Tibetans from southwestern China and northeastern China showed a similar population affinity with NEA ancients. The genomic affinity between modern Tibetans and some southern East Asians (such as Oakaie_LNBA) could be also identified in Figure 3. Admixture Signatures of Modern Tibetans and Ancient Populations From Tibetan Plateau We carried out admixture-f3-statistics in the form f3(source population1, source population2; Targeted Tibetan) to detect the signals of recent genetic admixture in Tibetans. We also re-evaluated the admixture signatures in the eight ancient individuals from Nepal and eleven ancient individuals from Qinghai province using this three-population comparison testing and our comprehensive ancient/modern reference dataset. We found different patterns of admixture signals and source populations in the highland/lowland ancient/modern Tibetans (Supplementary Tables 4–18). Besides, we also identified small but significant differences within geographically/culturally different Tibetans. By setting the statistically significant threshold at Z-score < −3, no admixture signals were observed in southern Tibetans (Shannan and Shigatse) over forty thousand tested pairs, and only four pairs in central Lhasa Tibetan with one source from 1500-year-old Samdzong and other from Kham Tibetan/Qiang, or the combination of southern Tibet Tibetan with Neolithic-NEAs or Baikal ancients (Supplementary Tables 4–6). It was interesting to find that 188 tested population pairs showed statistically significant f3-statistic values with one source from Tibeto-Burman speakers and the other from Western Eurasian Steppe pastoralists (Alan, Andronovo, Sintashta, Poltavka, and Yamnaya) in f3(Source1, Source2; Nagqu Tibetan). Tibetans from southern and central Tibet combined with the lowland modern East Asians, but not with ancient East Asians, could also produce significant admixture signals for Nagqu Tibetan (Supplementary Table 7). Chamdo Tibetan at the junction regions between Ü-Tsang Tibetan and Kham Tibetan had the potential possibility of cultural contact and population admixture, but only one pair of source populations could give a significant admixture signal in Chamdo Tibetans: f3(Lhasa Tibetan, Yajiang Tibetan; Chamdo Tibetan) = −3.49∗SE (Supplementary Table 8). Three Tibetans from the Gansu-Qinghai region possessed admixture signatures from over several thousand population pairs with one from modern or ancient East Asians and the other from Western Eurasians (Supplementary Tables 9–11). Results from f3(Yumin_EN, Austronesian/Tai-Kadai; Gansu-Qinghai Tibetan) showed that the combination of inland Neolithic NEA of Yumin_EN as northern ancestral source with Austronesian/Tai-Kadai speakers as the southern ancestral source could produce significant negative f3-values, and these admixture signals could also be identified in f3(Neolithic NEAs, Neolithic-Russian/modern Turkic/Mongolic/Indo-European speakers; Gansu-Qinghai Tibetan). Tibetans from Sichuan province only showed significant signals as an admixture between northern and southern East Asians or the highland Tibeto-Burman speakers and lowland East Asians, i.e., f3(highland Tibeto-Burman speakers, lowland Tibeto-Burman speakers; Sichuan Tibetan) < −3∗SE (Supplementary Tables 12, 13). Similar to the southern Tibet Tibetans, no obvious admixture signals were observed in Yunnan Tibetans, which may be caused by the genetic isolation or obvious genetic drift that occurred recently (Supplementary Table 14). The statistics focused on the ancient populations from the TP showed seven pairs can give admixture signals for modeling Qinghai Iron Age Dacaozi samples (Supplementary Tables 15–18), which are the pairs of ancient NEAs and modern SEAs, or Chamdo Tibetan-related source and Taiwan Iron Age Hanben-like populations. |
|
|
|
Post by Admin on Mar 24, 2022 20:14:16 GMT
Intra Population Differentiation Amongst High-Altitude and Low-Altitude Residing Tibetans Inferred From f4-Statistics To gain insights into the population substructures among modern Tibetans, we first conducted symmetry-f4-statistics in the form f4(modern Tibetan1, modern Tibetan2; modern Tibetan3, Mbuti), in which we expected to observe the non-significant f4-values if no significant differences existed between different Tibetan groups. As shown in Supplementary Table 19 and Supplementary Figure 10, we observed that Chamdo Tibetan formed a clade with Nagqu/Yunnan Tibetans compared with others in f4(Tibetan1, Chamdo Tibetan; Tibetan2, Mbuti) and all included Tibetans shared more alleles with Chamdo Tibetan compared with Ando Tibetans. Compared to the low-altitude Sichuan Tibetans, Chamdo Tibetan had more high-altitude Tibetan-related ancestry, while Gannan Tibetan shared more alleles with Xinlong Tibetan compared with Chamdo Tibetan. Compared with high-altitude Tibetans, Chamdo Tibetan shared more alleles with other low-altitude Tibetans. Results from the symmetry-f4(Shigatse/Shannan/Lhasa Tibetans, Shigatse/Shannan/Lhasa Tibetans; Tibetan2, Mbuti) with non-significant Z-scores showed clear genetic homogeneity among Tibet central/southern-Ü-Tsang Tibetans (Supplementary Figures 11, 12). Negative-f4-values in f4(Gansu-Qinghai Ando Tibetans, Shigatse/Shannan/Lhasa Tibetan; Tibetans, Mbuti) showed that all included Tibetans shared more alleles with southern Tibet Tibetans relative to Gansu-Qinghai Ando Tibetans. However, northern Tibet Tibetans formed a clade with Chamdo and Yunnan Tibetans and received more high-altitude Tibetan-related derived alleles compared with Gansu-Qinghai and Sichuan Tibetans. For lowland Tibetans, northwestern Tibetans in Gangcha and Xunhua formed one clade, i.e., all absolute Z-scores of f4(Gangcha, Xunhua Tibetan; Tibetan2, Mbuti) were less than three (Supplementary Figure 13). Compared with Gannan Tibetans, Qinghai Tibetans had more ancestry sharing with Tibet Tibetans. We did not find Tibetan populations shared more alleles with Gannan Tibetans relative to other Tibetans, as all values in f4(Tibetan1, Gannan Tibetan; Tibetan2, Mbuti) were larger than zero. Southwestern Yunnan Tibetan formed one clade with Chamdo/Xinlong/Yajiang Tibetans, all of them belonged to Kham Tibetans (Supplementary Figures 14, 15). Lowland Sichuan/Yunnan Tibetans harbored increased Tibetan-related derived alleles compared with Gansu-Qinghai Tibetans and more ancestry related to highland Tibetans compared with other highland Tibetans. We additionally explored genetic affinity and population substructure among highland and lowland Tibetans using ancient Eurasian populations via f4(Modern Tibetan1, Modern Tibetan2; Ancient Eurasians, Mbuti). The non-significant Z-scores in f4(Ü-Tsang Tibetans1, Ü-Tsang Tibetans2; Ancient Eurasians, Mbuti) confirmed the genomic homogeneity within the four high-altitude Ü-Tsang Tibetans. We could also identify the more allele sharing between the Nepal ancients and Ü-Tsang Tibetans compared to Ando and Kham Tibetans (Supplementary Figures 16–19). Compared with Shannan Tibetan, Nagqu Tibetan harbored increased ancestry associated with the lowland ancient populations. Compared to Qinghai Ando Tibetans, Nagqu Tibetan possessed both increased Nepal ancients-related ancestry and increased Late Neolithic Lajia-related ancestry relative to Xunhua Tibetan. Nagqu Tibetan also harbored additionally increased ancestry related to the coastal Late Neolithic SEAs, middle Yellow River Middle-Neolithic to Iron Age ancient populations, Upper Xiajiadian culture-related Bronze Age populations, inland Neolithic NEAs and other upper Yellow River Late Neolithic and Iron Age populations. Significant negative-f4-values were observed in Ando Tibetans via f4(modern Tibetan1, Gansu-Qinghai Ando Tibetans; Bronze Age stepped pastoralists, Mbuti), which suggested that Ando Tibetans harbored increased ancestry related to steppe pastoralists, such as Sintashta, Yamnaya, Afanasievo, Srubnaya, Andronovo and Xinjiang Iron Age Shirenzigou populations. Although strong genetic affinity within Ando Tibetans was confirmed with the similar patterns of f4-based sharing alleles and non-significant statistical results in symmetry-f4 statistics. Statistically significant negative f4-values in f4(Gangcha Tibetan, Gannan Tibetan; Ami/Atayal/Hanben/Gongguan/Tanshishan_LN/Qihe_EN, Mbuti) showed that Gannan Tibetan harbored increased SEA ancestry related to modern Austronesian or Proto-Austronesian-related Neolithic to present-day southeastern coastal/island populations (Supplementary Figures 20–22). A similar SEA affinity of Gannan Tibetan was also identified compared with Tibet Ü-Tsang Tibetans. Results of the four-population comparison analysis focused on Kham Tibetans are presented in Supplementary Figures 23–25, which suggested that Kham Tibetans had increased both northern and SEA ancestry. Spatiotemporal Comparison Analysis Among Modern Tibetans and All Paleolithic-to-Historic East Asians Showed the Genetic Admixture and Continuity of Modern Tibetans We nest used f4-statistics to elucidate the patterns of genomic structure and population dynamic of East Asians and provide new insights into the origin of culturally/geographically diverse Tibetans. Focused on four early coastal Neolithic NEAs from Shandong province, f4(coastal Neolithic NEA1, coastal Neolithic NEA2; Modern Tibetans/Ancient East Asians, Mbuti) revealed the similar genetic relationship between modern Tibetans and these different Neolithic NEAs (Supplementary Figure 26). Results from f4(Bronze/Iron Age Henan populations, Neolithic-to-Iron-Age Henan populations; Eastern Modern Tibetan/Ancient East Asians, Mbuti) only revealed Luoheguxiang people had increased ancestry associated with modern Austronesian-speaking Ami (Supplementary Figures 27–29) relative to Wanggou_MN. The Late Neolithic Haojiatai population had more SEA-like ancestry related to Xitoucun_LN and Iron Age Hanben people compared with Wanggou_MN (Supplementary Figure 30). The genetic affinity with southern coastal populations (Ami/Atayal/Hanben-related) was also observed in Pingliangtai_LN, but not in Wadian_LN and Middle Neolithic Wanggou_MN and Xiaowu_EN (Supplementary Figures 31–34). Focused on ancients from Shaanxi and Inner Mongolia, we found that modern Tibetans and northern and southern EAs from the Yellow River and south China shared more alleles with Late Neolithic Shimao populations (Supplementary Figure 35). Temporal analysis among upper Yellow River ancients showed all modern Tibetans showed a similar relationship with them, although Iron Age Dacaozi people harbored more SEA ancestry. These results suggested that population movements from southern China have a significant influence on the gene pool formation of northeastern populations on the TP at least from the Iron Age (Supplementary Figure 36). Symmetrical relationships among East Asians with temporally different Nepal ancient populations were shown in Supplementary Figure 37. Next, we also explored the similarities and differences of the shared genetic profiles related to northern Neolithic East Asians via the spatial comparison analysis with modern Tibetans and all available ancient East Asians as reference. We conducted a series of symmetry f4-statistics to compare all eleven modern Tibetan populations and other ancient East Asians against the geographically different ancient NEAs and ancient Tibetans. Figure 4 and Supplementary Figures 38–41 showed the shared alleles between the targeted populations and the lowland early Neolithic NEAs and others. The f4(NEAs, Chokhopani; Modern Tibetan/Neolithic-to-historic East Asians, Mbuti) was used to determine the lowland and highland East Asian affinity. Compared with four coastal Neolithic Shandong populations, we found that Ü-Tsang Tibetans had a strong highland East Asian affinity. Besides, comparison against the coastal and inland ancients revealed that modern Tibetans had a strong inland-NEA-affinity, especially with Late Neolithic Lajia people from the upper Yellow River. This Lajia-affinity or inland-NEA-affinity persisted when we substituted inland Yumin_MN with the coastal Neolithic NEAs (Supplementary Figure 42), but disappeared when we substituted the latter Neolithic groups with the early Neolithic NEAs (Supplementary Figures 43–48). We summarized the overall highland/lowland East Asian affinities of Tibetans in Supplementary Figure 49, which showed the Ando and Kham Tibetans had lowland NEA affinity, and Ü-Tsang Tibetans possessed additional Nepal ancient affinity. FIGURE 4 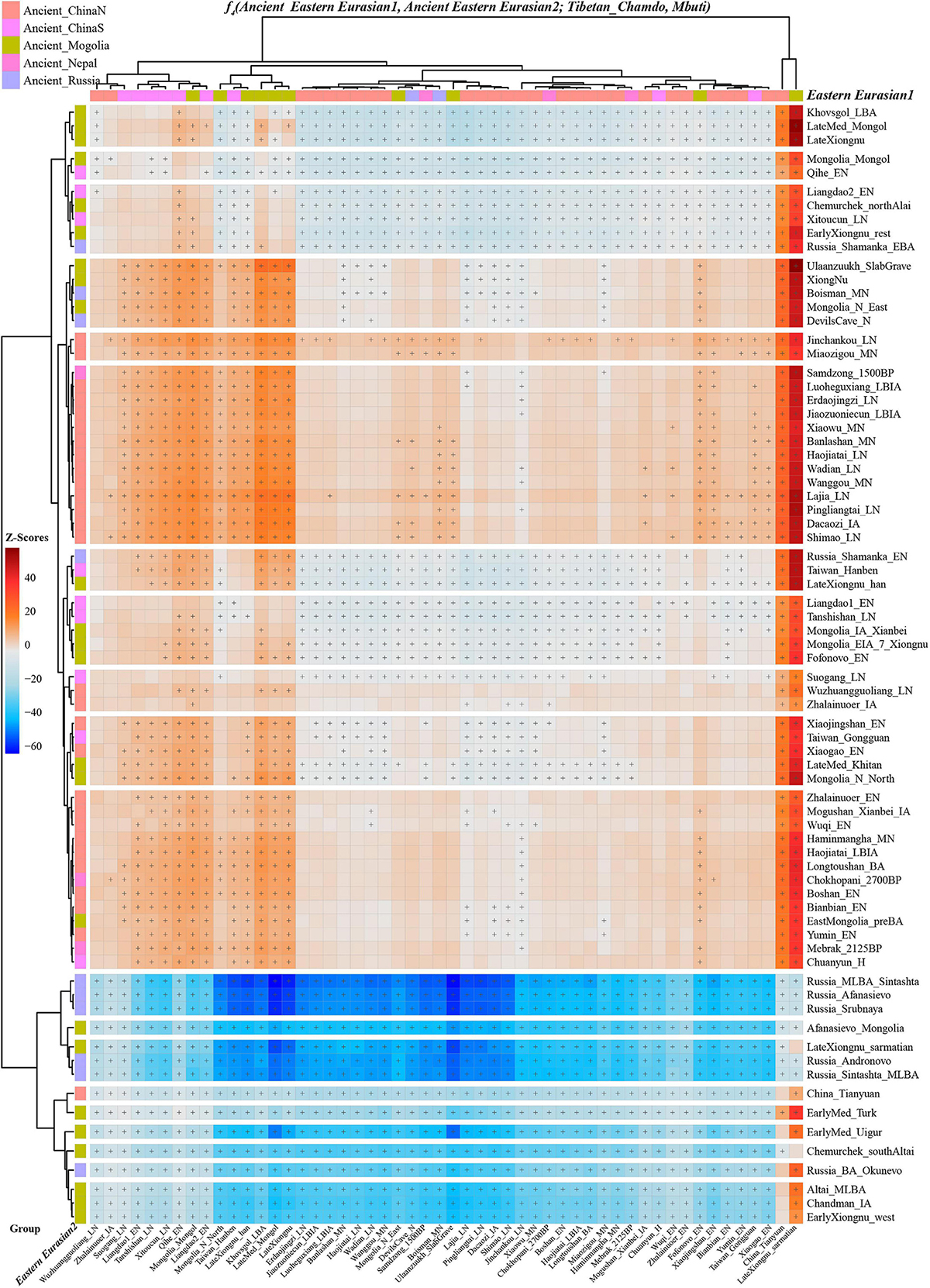 Figure 4. The genomic affinity between Chamdo Tibetans and other eastern Eurasian ancient populations inferred from four population affinity-f4 statistics of the form f4(Ancient Eastern Eurasian1, Ancient Eastern Eurasian; Tibetan_Chamdo, Mbuti). Red color with statistically significant f4-values (marked with “+”) demoted Chamdo Tibetans shared more derived alleles with Ancient Eastern Eurasian1 (right population lists) compared with Ancient Eastern Eurasian2 (bottom population lists). Blue color with significant f4-values denoted Chamdo Tibetans shared more Ancient Eastern Eurasian1-related derived alleles relative to their counterpart. Our genomic studies have identified population substructures within modern Tibetans. Modern Tibetans can be classified into three subgroups by their different affinities with NEAs, SEAs and Siberians, which were confirmed by the negative values in f4(Reference populations, modern Tibetans; northern/southern EAs and Siberians, Mbuti). We further tested if one single source could explain the observed genetic variations in Tibetans. We first assumed that modern Tibetans were the direct descendants of SEAs which is associated with the Yangtze Rice farmers. As shown in Supplementary Figures 50–58, we observed significant negative f4-values in f4(SEAs, modern Tibetans; Reference populations, Mbuti) when we used NEAs/Siberians as the reference populations, which indicated obvious gene flow events from these reference populations into modern Tibetans. We then assumed that Tibetans’ direct ancestor was coastal Neolithic-NEAs, we conducted f4(Shandong ancients, modern Tibetans; Neolithic-to-historic East Asians, Mbuti) and found only Nepal ancients showed the negative-f4-values, which was consistent with the common origin of the Sino-Tibetan speakers from YRB (Supplementary Figures 59–62). The patterns were confirmed when we assumed Yangshao and Longshan farmers or their related populations (Supplementary Figures 63–71), Shaanxi ancients (Supplementary Figures 72–74), and other ancient NEAs and southern Siberians (Supplementary Figures 75–88) as the direct ancestor of modern Tibetans. As shown in Supplementary Figures 75–88, when assuming Yumin or Ulchi as the direct ancestor of Tibetans, we identified additional gene flows from the SEAs (Hanben and Tanshishan et al.) and Yellow River farmers into Tibetans. Assuming the Nepal ancients as direct ancestors, we detected obvious additional gene flow from the lowland ancient East Asians to Kham Tibetans (Supplementary Figures 89–91). Additional predefined ancestral populations from Russia and Chinese Xinjiang further confirmed the strong northern East Asian affinity (Supplementary Figures 92–104). Thus, f4-statistics showed that the formation of modern Tibetans had involved multiple admixture events. |
|



