|
|
Post by Admin on Dec 11, 2015 7:44:20 GMT
 Mitochondrial analyses the Kostenki genome, the oldest Homo sapiens DNA sample from Europe tested to date, confirmed Kostenki 14 belongs to mtDNA Haplogroup U2, which is commonly found among Mesolithic European hunter-gatherers. The Y chromosome of Kostenki 14 belongs to the Asian haplogroup C M130, which attains the highest frequencies among the indigenous populations of Mongolia and the Russian Far East. Genetically speaking, Kostenki 14, the oldest European man discovered to date, was a mixed-race person with the European haplogroup U2 (mtDNA) and the Asian haplogroup C (Y-DNA), which shows that he was born between a European mother and an Asian father.  Kostenki is a rural area of Khokholsky District of Voronezh Oblast in Russia and Siberia has been a racial melting pot for at least 30,000 years with a mixed-race metapopulation that stretched from Europe to central Asia. For example, the Khanty and Mansi populations in Northwest Siberia are genetically half Asian and half European. 53.6% of the Khanty and 32.0% of the Mansi individuals carry West Eurasian haplogroups (R1a1, U4, U5a), whereas 21.4% of the Khanty and 44.0% of the Mansi are represented by East Eurasian haplogroups (C*, D*). 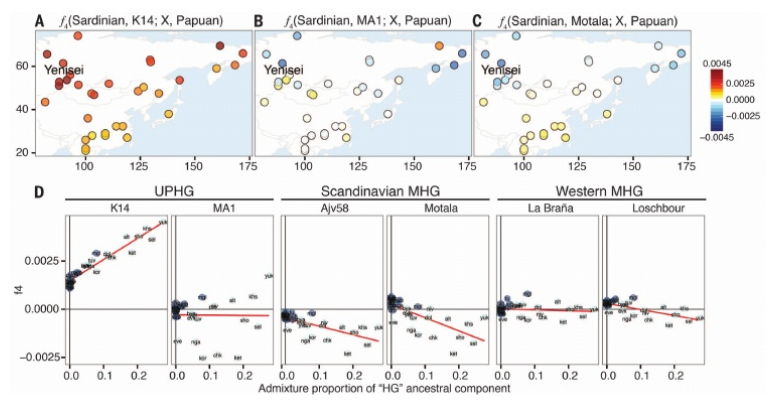 A ground-breaking new study on DNA recovered from a fossil of one of the earliest known Europeans - a man who lived 36,000 years ago in Kostenki, western Russia - has shown that the earliest European humans’ genetic ancestry survived the Last Glacial Maximum: the peak point of the last ice age. The study also uncovers a more accurate timescale for when humans and Neanderthals interbred, and finds evidence for an early contact between the European hunter-gatherers and those in the Middle East – who would later develop agriculture and disperse into Europe about 8,000 years ago, transforming the European gene pool. By cross-referencing the ancient man’s complete genome – the second oldest modern human genome ever sequenced – with previous research, the team discovered a surprising genetic “unity” running from the first modern humans in Europe, suggesting that a ‘meta-population’ of Palaeolithic hunter-gatherers with deep shared ancestry managed to survive through the Last Glacial Maximum and colonise the landmass of Europe for more than 30,000 years. While the communities within this overarching population expanded, mixed and fragmented during seismic cultural shifts and ferocious climate change, this was a “reshuffling of the same genetic deck” say scientists, and European populations as a whole maintained the same genetic thread from their earliest establishment out of Africa until Middle Eastern populations arrived in the last 8,000 years, bringing with them agriculture and lighter skin colour.  The Kostenki genome also contained, as with all people of Eurasia today, a small percentage of Neanderthal genes, confirming previous findings which show there was an ‘admixture event’ early in the human colonisation Eurasia: a period when Neanderthals and the first humans to leave Africa for Europe briefly interbred. The new study allows scientists to closer estimate this ‘event’ as occurring around 54,000 years ago, before the Eurasian population began to separate. This means that, even today, anyone with a Eurasian ancestry – from Chinese to Scandinavian and North American – has a small element of Neanderthal DNA.  |
|
|
|
Post by Admin on Dec 12, 2015 7:50:35 GMT
 A new paper in Science reports on the genome of Kostenki-14 (K14), an Upper Paleolithic European from Russia. This is now the third oldest Homo sapiens for which we have genetic data, after Ust'-Ishim (Siberia, 45 thousand years), Tianyuan (China, 40 thousand years), and now Kostenki (European part of Russia, 37 thousand years). Of these three genomes, the Ust'-Ishim is both the highest coverage and earliest (Siberia is the gift that keeps on givin'), Tianyuan only has its chromosome 21 known, and K14, a complete 2.42x coverage sequence (and, apparently, good teeth, after all these years; left). K14 may not be the older Upper Paleolithic human, but as of this writing it is the only Upper Paleolithic European that has been published so far, the next ones being the Loschbour, Motala, and La Brana Mesolithic Europeans who who have about 1/5 of its age. The new paper shows that K14 was definitely European (or more correctly West Eurasian or Caucasoid), as it was more similar to modern Europeans than to East Asians or other non-West Eurasian populations. Thus, the morphological description of the sample as "Australoid" by some early anthropologists did not reflect its ancestral makeup. Also, this proves that Caucasoids existed 37,000 years ago, which most physical anthropologists would believe, but it is nice to have direct confirmation. This pushes the lower bound from 24,000 years ago (because MA-1 was West Eurasian according to the results of Raghavan et al.). It will be nice to push the lower bound further to the past as there are much older bones (and plenty of teeth) from earlier Upper Paleolithic Europeans.  But there is a slight kink in the story, as K14 also belonged to Y-haplogroup C which is predominantly East Asian/Ocenian/Native American today. So, maybe there is some distant link to these populations in its ancestry. But, there is definitely a link to much more recent Europeans: the tiny percentage of living Europeans who have preserved K14's Y-chromosomal type (some of which were doubtlessly told a few years back that they were descendants of Genghis Khan, before the phylogenetic structure of C was known), the La Brana hunter-gatherer from Mesolithic Spain, as well as Neolithic Europeans from Hungary. 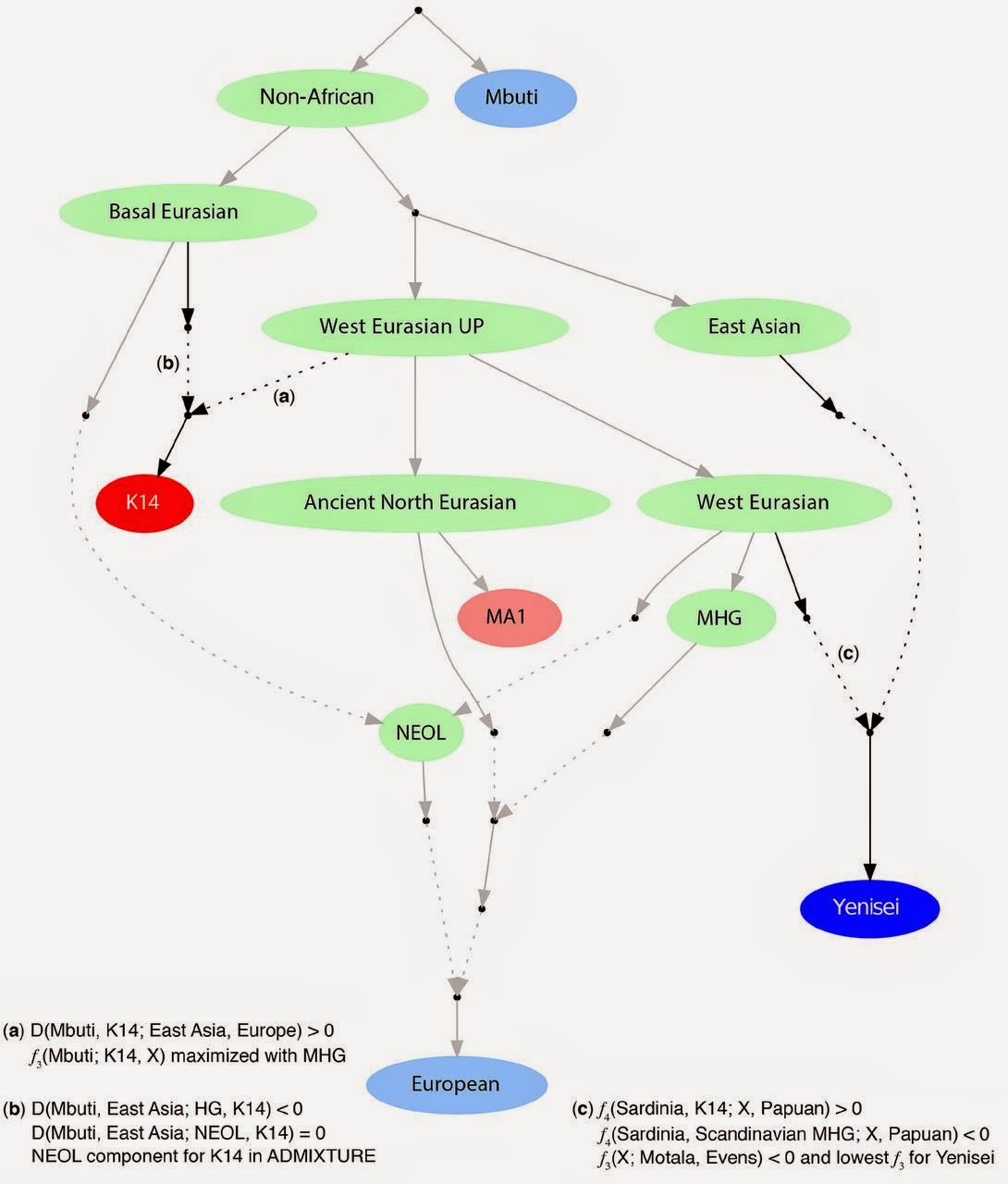 This model is not formally tested, but at least it seems to derive Europeans as a 3-way mixture that is basically identical to that of Lazaridis et al., with some relabeling of populations (MHG=WHG and NEOL=EEF). The model also includes Yeniseian Siberians as a mixture of MHG and East Asians (although it does not include actual East Asians). It's strange that Yeniseians apparently are given no ANE ancestry but only WHG/MHG. Both Raghavan et al. and Lazaridis et al. mentioned that ancestry related to MA-1 in living Siberians is diminished, but none at all? 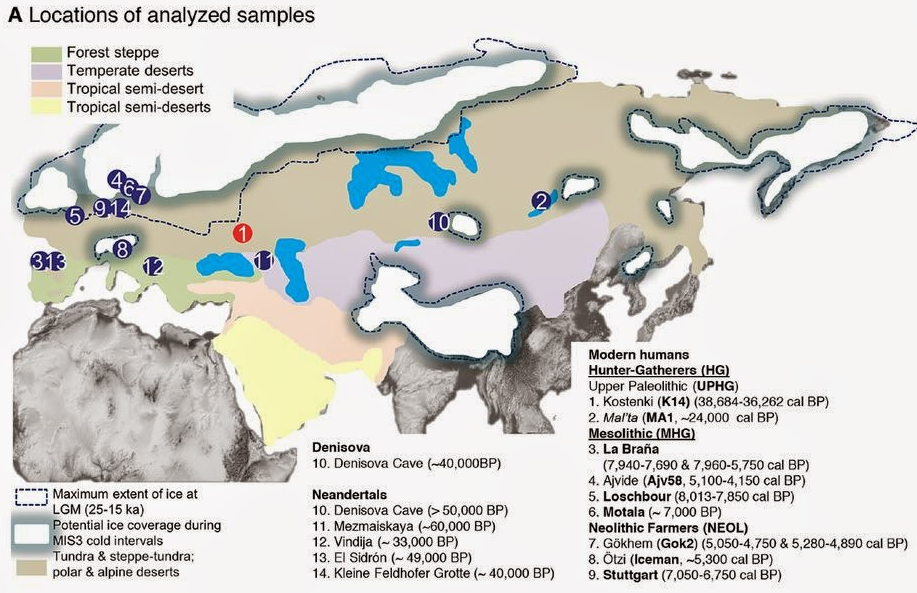 The major new finding of this paper, however, is that K14 had Basal Eurasian ancestry, which was first proposed for EEF from Germany 7,000 years ago, so now it postulated for Russian hunter-gatherers 37,000 years ago. I don't think many archaeologists would derive European farmers from Russia (Russia is actually one of the last places in Europe that became agricultural). So, maybe the hunter-gatherers from Russia had Basal Eurasian ancestry and this wasn't limited to the ancestors of the EEF? If they did, it's strange that Loschbour, La Brana, MA-1, Ust'-Ishim, Swedish Mesolithic (and maybe KO1?) didn't have it. So, either Kostenki was very unique or there is an alternative explanation for its strangeness. |
|
|
|
Post by Admin on Apr 3, 2019 21:16:36 GMT
 The study, reported in the journal Science, examined genetic information from the remains of anatomically modern humans who lived during the Upper Palaeolithic, a period when modern humans from Africa first colonised western Eurasia. The results suggest that people deliberately sought partners beyond their immediate family, and that they were probably connected to a wider network of groups from within which mates were chosen, in order to avoid becoming inbred. This suggests that our distant ancestors are likely to have been aware of the dangers of inbreeding, and purposely avoided it at a surprisingly early stage in prehistory.  The symbolism, complexity and time invested in the objects and jewellery found buried with the remains also suggests that it is possible that they developed rules, ceremonies and rituals to accompany the exchange of mates between groups, which perhaps foreshadowed modern marriage ceremonies, and may have been similar to those still practised by hunter-gatherer communities in parts of the world today. The study’s authors also hint that the early development of more complex mating systems may at least partly explain why anatomically modern humans proved successful while other species, such as Neanderthals, did not. However, more ancient genomic information from both early humans and Neanderthals is needed to test this idea.  The research was carried out by an international team of academics, led by the University of Cambridge, UK, and the University of Copenhagen, Denmark. They sequenced the genomes of four individuals from Sunghir, a famous Upper Palaeolithic site in Russia, which is believed to have been inhabited about 34,000 years ago. The human fossils buried at Sunghir represent a rare and highly valuable source of information because, very unusually for finds from this period, the people buried there appear to have lived at the same time and were buried together. To the researchers’ surprise, however, these individuals were not closely related in genetic terms; at the very most, they were second cousins. This is true even in the case of two children who were buried head-to-head in the same grave.  Professor Eske Willerslev, a Fellow at St John’s College, Cambridge, Prince Philip Professor of Ecology and Evolution in the Department of Zoology, and a Professor at the University of Copenhagen, was the senior author on the study. “What this means is that even people in the Upper Palaeolithic, who were living in tiny groups, understood the importance of avoiding inbreeding,” he said. “The data that we have suggest that it was being purposely avoided.” “This means that they must have developed a system for this purpose. If small hunter–gatherer bands were mixing at random, we would see much greater evidence of inbreeding than we have here.” Early humans and other hominins such as Neanderthals appear to have lived in small family units. The small population size made inbreeding likely, but among anatomically modern humans it eventually ceased to be commonplace; when this happened, however, is unclear. “Small family bands are likely to have interconnected with larger networks, facilitating the exchange of people between groups in order to maintain diversity,” Professor Martin Sikora, from the Centre for GeoGenetics at the University of Copenhagen, said.  Sunghir contains the burials of one adult male and two younger individuals, accompanied by the symbolically-modified incomplete remains of another adult, as well as a spectacular array of grave goods. The researchers were able to sequence the complete genomes of the four individuals, all of whom were probably living on the site at the same time. These data were compared with information from a large number of both modern and ancient human genomes. They found that the four individuals studied were genetically no closer than second cousins, while an adult femur filled with red ochre found in the children’s’ grave would have belonged to an individual no closer than great-great grandfather of the boys. “This goes against what many would have predicted,” Willerslev said. “I think many researchers had assumed that the people of Sunghir were very closely related, especially the two youngsters from the same grave.”  The people at Sunghir may have been part of a network similar to that of modern day hunter-gatherers, such as Aboriginal Australians and some historical Native American societies. Like their Upper Palaeolithic ancestors, these people live in fairly small groups of around 25 people, but they are also less directly connected to a larger community of perhaps 200 people, within which there are rules governing with whom individuals can form partnerships. “Most non-human primate societies are organised around single-sex kin where one of the sexes remains resident and the other migrates to another group, minimising inbreeding,” Professor Marta Mirazón Lahr, from the Leverhulme Centre for Human Evolutionary Studies at the University of Cambridge, said. “At some point, early human societies changed their mating system into one in which a large number of the individuals that form small hunter-gatherer units are non-kin. The results from Sunghir show that Upper Palaeolithic human groups could use sophisticated cultural systems to sustain very small group sizes by embedding them in a wide social network of other groups.”  By comparison, genomic sequencing of a Neanderthal individual from the Altai Mountains who lived around 50,000 years ago indicates that inbreeding was not avoided. This leads the researchers to speculate that an early, systematic approach to preventing inbreeding may have helped anatomically modern humans to thrive, compared with other hominins. This should be treated with caution, however: “We don’t know why the Altai Neanderthal groups were inbred,” Sikora said. “Maybe they were isolated and that was the only option; or maybe they really did fail to develop an available network of connections. We will need more genomic data of diverse Neanderthal populations to be sure.” |
|
|
|
Post by Admin on Apr 4, 2019 0:03:43 GMT
 Fig. 1 Sampling locations and genomic affinities of K14 and other ancient genomes. (A) Location of Kostenki and the samples analyzed in this study. Kostenki (K14) is shown in red, while comparative ancient samples are shown in blue. (B) Admixture proportions for the ancient genomes, assuming nine ancestral components for a clustering analysis in a set of modern worldwide populations. We labeled the components according to the modern populations in which they are maximized for all but one case: The yellow component that we label HG is maximized in eastern Europeans. NEOL: Neolithic farmers. (C) Shared drift between K14 and a set of worldwide populations. For every modern population X on the map, we compute f3(Mbuti Pygmy; K14, X). The warmer colors indicate increased shared ancestry. (D) Shared drift between K14 and a set of European populations. This figure is a close-up of (C). The ancestors of contemporary Eurasians are believed to have left Africa some 60,000 to 50,000 years ago (60 to 50 ka) (1, 2), possibly 30,000 to 40,000 years later than Australo-Melanesian ancestors (3). Despite controversies about routes out of Africa, the first Upper Paleolithic (UP) industries of Eurasia are found in the Levant from ~48 ka (4, 5). Expansion into Europe took place through multiple events that by ~40 ka had generated a spatially and culturally structured anatomically modern human (AMH) population—from Russia (6) to Georgia (7), Bulgaria (8), southern Europe (9, 10), and the United Kingdom (11). The few AMH fossils associated with these initial UP industries are morphologically variable (9, 12–17). In western Eurasia, the distinctive Aurignacian toolkit, first observed at Willendorf (Austria) by 43.5 ka (18), became predominant across the earlier range by 39 ka. Although analyses of ancient human genomes have advanced our understanding of the European past, revealing contributions from Paleolithic Siberians, European Mesolithic, and Near Eastern Neolithic groups to the European gene pool (19–23), the possible contribution of the earliest Eurasians to these later cultures and to contemporary human populations remains unknown. To investigate this, we sequenced the genome of Kostenki 14 (K14, Markina Gora) (Fig. 1A). The locality of Kostenki-Borshchevo on the Middle Don River, Russia, has one of the most extensive Paleolithic records in eastern Europe. The K14 human skeleton was excavated in 1954 (24) and was recently dated to 33,250 ± 500 radiocarbon years before the present (B.P.) (25), 38.7 to 36.2 thousand calendar years B.P. (ky cal B.P.), in agreement with the stratigraphic position of the burial that cuts into the Campanian Ignimbrite ash layer dated to ~39.3 ky cal B.P. (26). Below the skeleton, there is a distinctive early UP industry, with end scrapers, burins, prismatic cores, and bone artifacts (layer IV); the cultural layer above (layer III) has a regionally local character (27, 28) [supplementary materials (SM) S1 and S2]. We performed 13 DNA extractions from a total of 1.285 g of the left tibia (dorsal side of the shaft), using two extraction methods based on silica purification (29, 30). We first constructed seven Illumina libraries and validated the presence of typical signatures of postmortem DNA damage, using a fraction of DNA extracts (SM S3). The remaining extracts were built into 63 libraries after enzymatic uracil-specific excision reagent (USER) treatment to limit the effect of nucleotide misincorporations in downstream analyses (31) (table S2). Additionally, a limited fraction of two DNA extracts was purified for methylated DNA fragments using methyl binding domain (MBD) enrichment (32) before USER treatment and library building, for a total of eight DNA libraries. Following stringent quality criteria for read alignment, we identified a total number of 175.2 million unique reads aligning against the human reference genome hg19, representing an average depth of coverage of 2.84X (SM S4). The eight USER-treated DNA libraries that exhibited limited error rates and contamination levels were selected for further analyses. This restricted the data set to 148.9 million unique reads, representing a final depth of coverage of 2.42X. We exploited the fact that K14 was a male and used the heterozygosity levels present in the X chromosome to estimate overall levels of contamination around 2.0% (SM S5 and S6 and table S5). The population genetics analyses results are robust to contamination of that level. In particular, we replicated the main analyses with selected libraries with varying contamination levels and observed no qualitative effect on the results (see SM S9 for details).
Mitochondrial analyses confirmed the sequence previously reported for K14 [haplogroup U2 (33)], which supports data authenticity. The Y chromosome belongs to haplogroup C M130, the same as in La Braña—a late Mesolithic hunter-gatherer (MHG) from northern Spain (22) (SM S7).To identify patterns of shared ancestry and admixture among K14, other ancient genomes and contemporary Eurasians [based on a single-nucleotide polymorphism (SNP) array panel of 2091 individuals from 167 populations], we carried out a series of analyses—model-based clustering and principal component analysis (PCA)—to show the contribution of diverse genetic components within K14: D statistics to explore the affinity of K14 to pairs of populations (using Mbuti Pygmy as an outgroup); f4 statistics to test whether a given modern population is equidistant to an ancient individual and a particular recent group (here, Sardinians), given an outgroup (here, Papuans); and f3 statistics to explore both patterns of admixture (“admixture” f3) and shared ancestry (“outgroup” f3). Key results were also replicated using two whole-genome sequencing data sets of modern individuals from worldwide populations (23, 34).  Fig. 2 Relationships of the K14 sample and MA1, MHG, NEOL, modern Europeans, and the modern populations in the Yenisei region. Model-based clustering analyses (35) show that K14 has different genetic components of substantial size (Fig. 1B and SM S10), suggesting the sharing of sets of alleles with different Eurasian groups. The largest fraction of K14’s ancestry derives from a component that is maximized in European MHGs and also predominant in contemporary northern and eastern Europeans. The genetic affinity of K14 to contemporary Europeans is also observed using outgroup f3 statistics (36). Using Mbuti Pygmy as an outgroup, we find that among a panel of 167 contemporary populations, Europeans have the greatest affinity (i.e., the largest f3) to K14 (Fig. 1C). This conclusion is also formally supported by comparing pairs of populations to K14 using the D statistics of the form D(Mbuti Pygmy, K14; Population 1, Population 2). This statistic is expected to be equal to zero if K14 is symmetrically related to Population 1 and Population 2, whereas its expectation is negative (positive) if K14 is more closely related to Population 1 (Population 2). For pairs of populations involving East Asians (Population 1) and Europeans (Population 2), K14 is always significantly more closely related to Europeans [e.g., Z = 12.1 (Han and Lithuanians)], in all data sets analyzed (SM S9 and table S7). We also confirmed that these results are robust to possible contamination from a modern DNA source by filtering for reads with a high likelihood of ancient DNA using a model-based approach (37), as well as calculating contamination-corrected D statistics (23) (SM S9 and fig. S18). Within Europe, northern Europeans show the closest affinity to K14, based both on the f3 (Fig. 1D) and D statistics [e.g., Z = 6.7 for Sardinians and Lithuanians; table S7 and fig. S16]. This pattern closely resembles that of European MHGs (La Braña, Ajv58, Loschbour, and Motala) and Mal’ta (MA1) (figs. S14 and S15), with the exception of the latter’s strong genetic affinity with Native Americans, which is unique to that individual. Furthermore, a direct comparison to ancient genomes in the outgroup f3 statistics shows that K14 has a higher affinity with MHGs (Loschbour and La Braña) than any other ancient individual or contemporary population (fig. S14). Together with the rare Y chromosome lineage shared with La Braña, these results provide strong evidence of shared ancestry and extensive gene flow between UP West Eurasian people related to K14 and European MHGs and their contemporary European descendants. An interpretation of the above results would be that K14 is an early member of a lineage leading to western Eurasian MHGs after their split from the proposed ancestral northern Eurasian lineage, including MA1. However, D statistics of the form D(Mbuti Pygmy, Modern; Ancient, K14)—which test whether K14 and an ancient individual form a clade with respect to a modern population—reject this simple tree-like relationship. We find that all contemporary non-Africans, except Australo-Melanesians, are closer to either MA1 or MHGs than to K14 [e.g., Z = –5.3 for D (Mbuti and Han; Loschbour and K14); SM S9, table S10, and fig. S19]. This would suggest a basal position of K14 with respect to MHGs and ancient north Eurasians, which is also shown in admixture graphs using TreeMix (SOM S12 and figs. S24 and S25). In addition, a sizeable component of K14’s ancestry observed in the model-based clustering analyses is predominant in contemporary Middle Eastern/Caucasus (ME/C) populations and Neolithic ancient genomes (NEOL) (Gok2, Iceman, and Stuttgart) but absent in MA1 or MHGs (Fig. 1B and fig. S20). This component has been associated with a suggested “basal Eurasian” lineage contributing to NEOL to explain an observed increase in allele sharing between MHGs/MA1 and East Asians compared with NEOL (21). Because K14 shows the same pattern as NEOL, a parsimonious explanation would be that K14 also derives some ancestry from a related basal Eurasian lineage. Consistent with this hypothesis, we find that East Asians are equally distant to NEOL and K14, using D statistics as described above [e.g., Z = 0.0 for D (Mbuti, Han; Stuttgart, K14); tables S10 and S11]. This suggests that the main ancestral components proposed for contemporary Europeans, including the Middle Eastern component commonly attributed to the expansion of early farmers within Europe, were likely already genetically differentiated and related through complex gene flow by the time of K14, at least 36.2 ka (Fig. 2). |
|
|
|
Post by Admin on Apr 4, 2019 18:18:37 GMT
We further investigated the relationship of K14 and the other ancient genomes to East Asian and Siberian populations using f4 statistics f4(Sardinian, Ancient; Modern, Papuan), which measure whether a modern population shares more alleles with contemporary Europeans or an ancient genome. We find that all Siberian and East Asians are equally distant from western MHGs (all |Z| < 1.9) (Fig. 3D and table S12), supporting the postulated early split between East Asians and western Eurasians. In contrast to MHGs and MA1, all Siberian populations are genetically closer to contemporary Europeans (Sardinians) than to K14 (3.1 < |Z| < 9.9) (table S12), particularly those from the Yenisei and Ob’ basins (e.g., Shors, Z = 8.0) (Fig. 3A). Furthermore, these populations derive parts of their ancestry from a European hunter-gatherer (HG) component inferred in the clustering analysis (Fig. 1D and fig. S20), with populations showing a higher HG ancestry proportion also being closer to contemporary Europeans, using the f4 statistic (Spearman ρ = 0.96; P = 3.0 × 10−18) (Fig. 3D and table S13). Notably, the opposite pattern is observed with Scandinavian MHGs (Ajv58 and Motala), where the same populations tend to share more alleles with MHGs than contemporary Europeans and the HG component is negatively correlated with f4 (e.g., Motala ρ = –0.85; P = 6.2 × 10−10) (Fig. 3, C and D). Calculating admixture f3 statistics, we find significant evidence for admixture in those populations, with a variety of Siberian and European source populations. The best pair of source populations (i.e., the most negative f3 statistic) involves Swedish MHGs (Motala) and Evens (a northeast Siberian population) [e.g., f3 (Shors; Evens, Motala) = –0.012; Z = –9.1] (table S14). Altogether, these results suggest that contemporary Siberian populations from the Yenisei basin derive part of their gene pool from a Eurasian HG population that shares ancestry with K14 but is more closely related to Scandinavian MHGs than to either MA1 or western European MHGs, indicating gene flow between their ancestors and Scandinavian Europe after K14 but before the Mesolithic (between 36.3 and 7 ky B.P.).  Fig. 3 Relationships of K14 and other HG genomes with contemporary East Asian and Siberian populations. (A) Values of the f4 statistic for a set of Siberian and East Asian populations and K14. We compute the f4 statistic for a topology (Sardinian, K14; X, Papuan). Warmer values indicate departure from the topology (Sardinian, K14; X, Papuan) with increased ancestry between the modern population X and the Sardinian. The Yenisei region includes the Selkup, Shor, and Ket populations. (B) Values of the f4 statistic for a set of Siberian and East Asian populations and MA1. We compute the f4 statistic for a topology (Sardinian, MA1; X, Papuan). (C) Values of the f4 statistic for a set of Siberian and East Asian populations and Scandinavian hunter gatherers (Motala). (D) Relationship between the HG admixture proportion and the f4(Sardinian, K14; X, Papuan) shown in (A). The red lines are linear regressions for each case. Finally, we estimated levels of Neandertal ancestry in K14 using f4-ratio statistics (38). Our estimates are consistent with previous analyses (34) showing a Neandertal contribution lower than 2% for most individuals (Fig. 4A). However, both La Braña and K14 show slightly elevated levels, with an estimated 2.4 ± 0.4% in K14 (tables S15 and S16). Restricting this analysis to genomic regions without evidence for Neandertal introgressed haplotypes in contemporary humans (38, 39) results in 0% estimated ancestry for most individuals except K14, where 0.9 ± 0.4% Neandertal ancestry is still detected (tables S17 and S18). The difference between K14 and modern genomes could be caused by several factors, including sampling effects and genetic drift, natural selection as argued in (38, 39), or by the effects of additional Neandertal admixture not represented in the modern gene pool. We next compared the size distribution of genomic tracts of archaic hominin origin in K14 and other ancient individuals (Fig. 4B) by identifying genomic regions with high frequencies of archaic alleles at sites where all modern Africans carry the ancestral allele. The length of Neandertal tracts was higher in K14 than in other ancient individuals, with the longest tract totaling ~3 Mb on chromosome 6 (Fig. 4C). This is consistent with K14 being closer to the time of the admixture event with Neandertals, and carrying longer archaic tracts that have been affected by less recombination, than in the other ~11,000- to 30,000-year-old younger ancient genomes. We then used the length distribution of shared ancestry to estimate the admixture time of Neandertals and humans based on the K14 sample and obtained an estimate of ~54,000 years (S15). We note that genomic data from a 45,000-year-old modern human from Siberia, which was published during the review process of this study, also shows longer segments of Neandertal ancestry, further supporting our conclusions (40). Because of the divergent position of the K14 sample, we also examined whether it contained any fragments of introgressed DNA from other previously unsampled hominins. However, the distribution of tracts of divergent DNA provides no evidence for additional divergent introgressed DNA (S14).  Fig. 4 Neandertal admixture in K14 and other ancient genomes. (A) Neandertal admixture proportions for the modern and ancient individuals from Eurasia. (B) Ancestry tract length distribution for tracts identified as Neandertal through a sliding-window approach. The sites are ascertained to be ancestral in the African populations. For each non-African, the tracts are identified as the regions where sites are derived in Neandertal and the individual shown in X. (C) The longest Neandertal haplotype identified in K14 through a sliding-window approach. Individuals were clustered using hierarchical clustering on the genotype matrix for the region. Missing data are shown in white; gray indicates homozygous ancestral, blue heterozygote-derived, and black homozygous-derived. Several studies have reported on the basal genetic distinctiveness between western Eurasian and eastern Asian populations, as well as between all Eurasians and Australo-Melanesians (41–43). Our results show no close genetic relationship between K14 and Australo-Melanesians and support earlier studies that suggest Australo-Melanesians derive part of their ancestry from an early population divergence that predates the separation of Europeans and East Asians (3). The K14 genome shows that this early UP individual was clearly part of a western Eurasian lineage that had already diverged from eastern Asians, thus establishing a minimum date for that separation at least 36.2 ka. The fact that the limited genomic information on the ~40 ka Tianyuan modern human from China clusters with contemporary East Asian populations (44) suggests an even earlier date. Our results further suggest that the early stages of the western Eurasian lineage were already complex (see also Fig. 2). Besides its core affinities with subsequent European groups, K14 also shares alleles with European Neolithic farmers and contemporary people from the Middle East/Caucasus, which are not found in MA1 and western European MHGs, indicating genetic exchange between K14 and a Basal Eurasian Lineage (which eventually contributed to Neolithic groups) after the ancestors of MA1 and subsequent European MHGs had diverged. This implies that early AMH populations became structured early in their history, but in the UP already contained the major genetic components found in Europeans today. As such, our findings show the existence of a metapopulation structure in Europe from the Upper Paleolithic onward, remnants of which are still found today despite migrations to and from Europe since the UP. The early UP contribution is greater among northern than southern Europeans, in agreement with the southeast to west and north gene flow cline resulting from the expansion of Neolithic famers 9 to 6 ky cal B.P. (20, 45). However, descendants of the early UP population represented by K14 likely also contributed genes to western Siberian groups living around the mouth of the Yenisei River. Therefore, our findings support the view that these Uralic-speaking populations represent an ancient admixture between European and East Asian lineages. The recently proposed Holocene gene flow from East Asians into northern Europeans (21) can, in our view, be equally well explained by population structure of the hunter-gatherer metapopulation within Europe. As such, our results paint an increasingly complex picture of colonization history of Europe from the UP to today. Instead of inferring a few discrete migration events from Asia into Europe, we now see evidence that humans in Western Eurasia formed a large metapopulation with gene flow in multiple directions occurring repeatedly and perhaps continuously. Science 28 Nov 2014: Vol. 346, Issue 6213, pp. 1113-1118 |
|




















