|
|
Post by Admin on Nov 10, 2021 0:22:15 GMT
Results Authenticity of results A total of 23 reproducible mtDNA fragments (360 bp) were obtained from 30 individual sets of the Xiaohe human remains, after discarding seven samples due to failed amplification or irreproducible results. Three of the 23 DNA fragments were similar to the DNA of two people involved in the study and were also removed from the study, even though they yielded consistent results through three or more independent extractions. The remains of 20 individuals were selected for cloning analysis. Eight to twelve clones from two independent amplifications of the 20 individuals were selected for automated DNA sequencing. Although a few positions were different from direct sequencing of the PCR products, which could be due to random Taq misincorporation or DNA damage, the consensus sequence from cloning was consistent (see Additional File 2). In order to validate the results generated in our laboratory, human remains from three Xiaohe individuals (100, 119, and 127) were sent to the ancient DNA laboratory of Fudan University for further analysis and identical results were obtained. The following facts further confirm the authenticity of our results: (1) an inverse correlation between the size of the PCR amplicons and the amplification efficiency for the Xiaohe samples (209 bp > 235 bp > 399 bp) was found; (2) molecular sexual identification results were in accordance with morphological sex assignments; (3) HVR-I sequences match well with the key coding region SNPs according to the well-defined mtDNA phylogenetic tree [16, 17]; and (4) parallel studies were also performed on Xiaohe animal samples, and no human DNA was detected. Collectively, 20 mtDNA fragments from 30 human remains were obtained that were inferred to be authentic (Table 3). The sequences have been submitted to GenBank with accession numbers FJ719792-FJ719811. This high success rate suggests that the DNA from the Xiaohe human remains is well preserved, as would be expected in samples originated from dry environmental conditions. MtDNA haplogroup profile and distribution The 20 mtDNA fragments containing eight different haplotypes that can be further assigned to five haplogroups, which belong to subhaplogroups of macrohaplogroups M and N (Figure 2). Figure 2 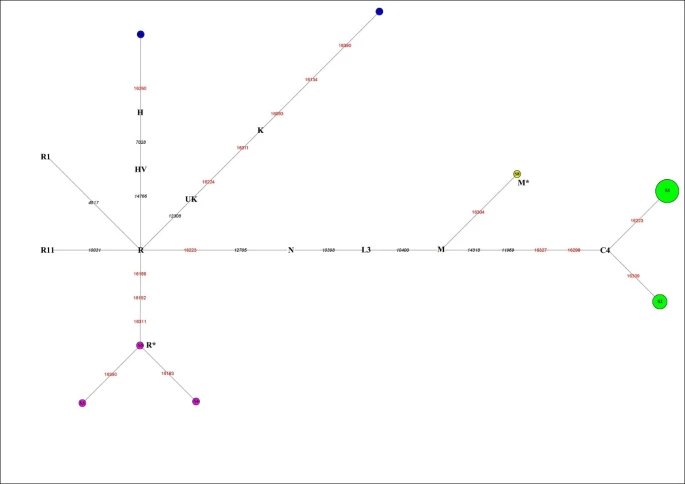 Reduced median network of Xiaohe sequences. Node size is proportional to frequencies. HVR1 positions are numbered relative to Cambridge reference sequence (CRS). Single nucleotide polymorphism diagnostic positions are in black italics; green represents East Eurasian lineage C, containing 14 individuals. Xiaohe R* is the cluster under the macrohaplogroup R. The dominant haplogroup in the Xiaohe people was the East Eurasian lineage C, shared by 14 Xiaohe individuals who were associated with two different mtDNA haplotypes (S1 and S2). According to the coding region 11969 G to A, all lineage C found in the Xiaohe people was further changed to subhaplogroup C4, which had D-loop group-specific polymorphisms at nucleotide positions (np) 16298 (T to C) and 16327 (C to T) [18]. Interestingly, the haplotype S1 shared by these 10 of the 14 Xiaohe individuals did not have the 16223T, a mutation had been found in the majority of modern lineage C populations. This haplotype was found merely in modern Evenks and Udegeys of southeastern Siberia but the frequency is low [18]. It is interesting to note that our study found a mutation, unique to the Xiaohe people but rare among lineage C, 16309 G (see Additional File 3). To the best of our knowledge, the lineage C with 16309 G was observed only in three people from modern central Asia to date, who also possess another two mutations in HVRI region [19]. Besides the East Eurasian lineage, two West Eurasian mtDNA haplogroups H and K were found among the Xiaohe people. H lineage is the most common mtDNA haplogroup in West Eurasia [20], but haplogroup H with a 16260T was shared by only nine modern people in Genbank, including one Italian, one German, one Hungarian, one Portuguese, one Icelander and four English people. Haplogroup K, a western Eurasian-specific haplogroup, is mainly distributed in Europe, central Asia, and Iran [20, 21]. However, haplogroup K with 16134T, found in the Xiaohe people, has not been found in modern people to our knowledge. Among the Xiaohe people, three sequences with the unique HVRI motif 16189-16192-16311 formed a subcluster (Figure 2) and were not shared by modern people. They are identified as macrohaplogroup R through sequencing the PCR amplicons at np10400 and np12705 in the coding region. The np12308, np14766, np10031, np4917, np3970, and 9 bp deletion, which are the diagnostic sites for the main subhaplogroups of R, were further examined [15]. The results showed that they are related neither to the West Eurasian haplogroups UK, TJ, HV, R11 and R1, nor to the East Eurasian haplogroups B and F. So we designated them as haplogroup R* temporarily. Another sequence with motif 16223-16304, shared by some people from East Asia, India, and Europe, was assigned to haplogroup M*. Y chromosome haplogroup profiling and distribution Fifteen individuals' AMG amplicons were obtained from the 20 Xiaohe individuals (whose mtDNA was successfully amplified), among which seven individuals were identified as male and eight as female. The Y chromosome haplogroup of the seven males were all assigned to haplogroup R1a1a through screening the Y-SNPs at M89, M9, M45, M173 and M198 successively. Haplogroup R1a1a is widely distributed in Eurasia: it is mainly found in Eastern Europe, Central Asia, South Asia, Siberia, ancient Siberia, but rare in East Asia [22–24]. |
|
|
|
Post by Admin on Nov 10, 2021 3:25:21 GMT
Discussion The Xiaohe cemetery is the oldest archeological site with human remains discovered in the Tarim Basin to date. Our genetic analyses revealed that the maternal lineages of the Xiaohe people were originated from both the East and the West, whereas paternal lineages discovered in the Xiaohe people all originated from the West. The East Eurasian lineage C, which was widely distributed in modern Asian populations, was the dominant haplogroup in the remains recovered from the lowest layer of the Xiaohe cemetery. This lineage is most frequently found in modern Siberian populations (Evenks, Yakut, Evens, Tuvinian, Buryat, Koryak and Chukchi) and to a lesser extent in modern East Asian (Mongolian, Daur and Korean) and Central Asian populations [25–29] (Figure 3). It was also found in the ancient Qinghai (4000BP) of China [30] and ancient South Siberian populations [31, 32]. In order to trace the original wellspring of lineage C in the Xiaohe population, a phylogenetic tree was constructed using 14 ancient Xiaohe samples and 522 modern haplogroup C samples from surrounding areas of the Xiaohe cemetery, including Siberian, Mongolian, Central Asian, northern Chinese and northern minorities of East Asia (see Additional File 3). The phylogenetic network displays a star-like distribution within the South Siberian population, which has an ancestral haplotype motif 16223-16298-16327. The ancestral haplotype was found mainly in South Siberian whose diversity of haplotypes C is very high (Figure 4). Therefore, the original source of haplogroup C was inferred to South Siberian. It is important to note that the C haplotypes of the Xiaohe people had only a single mutation compared with the ancestral haplotype. The shared sequences of the Xiaohe C haplotype (S1) were distributed in southeastern Siberia. It implies that the east Eurasian component in the Xiaohe people originated from the Siberian populations, especially the southern or eastern Siberian populations. Figure 3 Map of Eurasia, showing the approximate distribution of haplogroup C. Black sections reflect the frequency of haplogroup C data taken from references listed in Additional File 3. 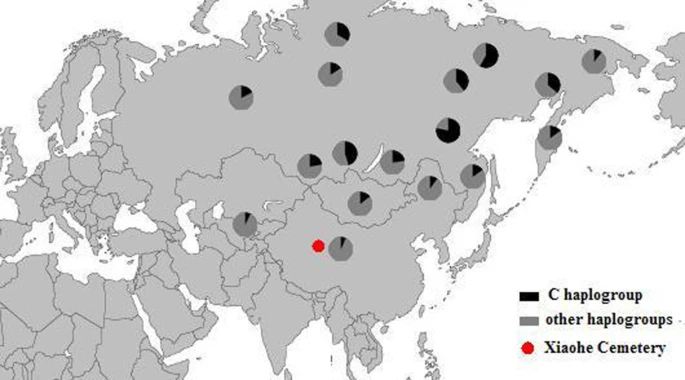 Figure 4 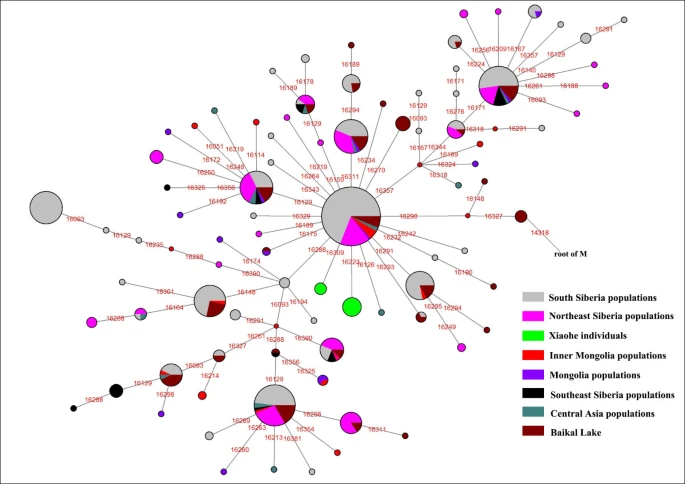 Phylogenetic tree of haplogroup C, based on HVS-I sequences in the region 16050-16391. For references for the mitochondrial DNA sequences in this study see Additional File 3; the length difference mutations were excluded from this analysis. The mtDNA haplogroup H is the most common mtDNA haplogroup in Europe, especially in northwestern Europe, and its frequency can be as high as 65% in Iberia. Frequencies gradually decrease from the northwest to the southeast of Europe. By contrast, the frequency of haplogroup H rises to only 20% in the Near East, and to less than10% in Central Asia, and is very low in East Asia [33, 34]. All of the shared sequences of the Xiaohe H haplotype, however, were distributed in Western Europe. Haplogroup K is also common in Europe, particularly around the Alps and the British Isles. It is found with less frequency in North Africa, the Middle East, and South Asia [21, 35–37]. Considering the presence of haplogroups H and K in the Xiaohe people and the geographical distribution of shared sequences, we conclude that the west Eurasian component observed in the Xiaohe people originated from western European, and maternal ancestry of the Xiaohe people might have close relationship with western European. Regarding the Y chromosomal DNA analyses, the seven males identified all belonged to haplogroup R1a1a. It is most frequently found in Eastern Europe, South Asia and Siberia. In contrast, it is relatively uncommon in Middle Easterners and rare in East Asian [22–24]. It is thought to be a trace of the migration events of early Indo-European [38, 39]. The presence of haplogroup R1a1a in the ancient Xiaohe people implies that the parental ancestry of the Xiaohe people originated from somewhere in Siberia or Europe, which is consistent with the origin of maternal ancestry. It is generally agreed that the origin of modern populations in Xinjiang and Central Asia is the result of the admixture of people from the West and the East [19, 25, 40]. When and where this admixture first occurred has long been of interest to geneticists and archaeologists [41–44]. The year 132 BC is often considered to be the beginning of contact between the East and the West along the Great Silk Road, since the Chinese explorer Zhang Qian went westward into Central Asia at that time. However, Mair has suggested that the date should be even earlier, based on the fact that silk appeared in Europe at 1000 BC [1]. In this study, the East and West Eurasian lineages are seen to coexist in the Xiaohe people, implying that the East had contacted the West during the early Bronze Age. It is noteworthy that the maternal lineage of five male individuals (106, 111, 115, 136 and 139) originated from East Eurasian, whereas their paternal lineage originated from the West Eurasian, implying that the Xiaohe population had been an admixture of people from both the West and the East. Given the unique genetic haplotypes and the particular archaeological culture, the time of this admixture could be much earlier than the time at which the Xiaohe people were living at the site. This means that the time of their mingling was at least a 1000 years earlier than previously proposed. However, the mtDNA haplogroups H, K and C all are very ancient lineages, over 10,000 years old in vast north Eurasia, whereas the civilization of the Tarim Basin, according to the archaeological materials, arose very late. The admixture therefore probably occurred elsewhere, before immigration into the Tarim Basin. The Xiaohe people might well have been an admixture at the time of their arrival. Where did the initial admixture occur? The admixture from people of the West and the East was also found in ancient Central Asia, Siberia, and Mongolia [16, 38, 45, 46]. The extent of the admixture varied in different regions and at different periods (Figure 5). Central Asia has always been the crossroads of contact between the West and the East. Lalueza-Fox et al. proved that Eastern lineages coexisted with Western lineages in Central Asia after 700 BC [41], whereas the West had met the East in south Siberia in the Bronze Age [38], and even earlier at Lake Baikal [45]. Xinjiang and the surrounding areas, especially south Siberia, were places at which the contact between western and eastern populations occurred earlier than in Central Asia. Given the fact that the mtDNA haplogroup C was distributed mainly in south Siberia, and that haplogroups H, K and R1a1a already had spread eastward into south Siberia during the Bronze Age, it is possible that the initial admixture occurred somewhere in southern Siberia. Considering that the cultural characteristics of the Xiaohe cemetery are similar to those of the Andronovo or Afanasevo culture that appeared throughout the southern Russian steppe, Kazakhstan, and western Central Asia during the second millennium BC [1, 46], the admixed population might have had relationship with populations settled South Siberia during the Bronze Age. Conclusions Our results demonstrated that the Xiaohe people was an admixture from populations originating from both the West and the East, implying that the Tarim Basin had been occupied by an admixed population since the early Bronze Age. Considering the unique genetic haplotypes and particular archaeological culture, the admixed population might have had relationship with populations settled South Siberia during the Bronze Age. To our knowledge, this is the earliest genetic evidence of an admixed population settled in the Tarim Basin. |
|
|
|
Post by Admin on Dec 19, 2021 18:59:09 GMT
The population history of northeastern Siberia since the Pleistocene Northeastern Siberia has been inhabited by humans for more than 40,000 years but its deep population history remains poorly understood. Here we investigate the late Pleistocene population history of northeastern Siberia through analyses of 34 newly recovered ancient genomes that date to between 31,000 and 600 years ago. We document complex population dynamics during this period, including at least three major migration events: an initial peopling by a previously unknown Palaeolithic population of ‘Ancient North Siberians’ who are distantly related to early West Eurasian hunter-gatherers; the arrival of East Asian-related peoples, which gave rise to ‘Ancient Palaeo-Siberians’ who are closely related to contemporary communities from far-northeastern Siberia (such as the Koryaks), as well as Native Americans; and a Holocene migration of other East Asian-related peoples, who we name ‘Neo-Siberians’, and from whom many contemporary Siberians are descended. Each of these population expansions largely replaced the earlier inhabitants, and ultimately generated the mosaic genetic make-up of contemporary peoples who inhabit a vast area across northern Eurasia and the Americas.Northeastern Siberia (the modern Russian Far East) is one of the most remote and extreme of the environments that were colonized by humans in the Pleistocene epoch. Extending from the Taimyr Peninsula in the west to the Pacific Ocean in the east, and north from the border between China and Russia to the Arctic Ocean, the region is currently home to dozens of diverse ethnolinguistic groups. Recent genetic stud-ies of the indigenous peoples of this land have revealed complex pat-terns of admixture, which are argued to have occurred largely within the past 10,000years1–3. Humans have been in the region for far longer4–6, but their origins and the demographic processes of this deeper popu-lation history are largely unknown. The earliest, most secure archae-ological evidence for human occupation in this region comes from an artefact-rich, high-latitude (approximately 70°N) site on the Yana River (Siberia) named Yana RHS, which dates to 31,600calibrated years before present (taken as 1950)4 (Fig.1). Yana RHS yielded a flake-based stone tool industry and sophisticated bone and ivory artefacts, which are reminiscent of technologies seen in the Upper Palaeolithic in Eurasia and southern Siberia5,7 (Extended Data Fig.1). By the time of the Last Glacial Maximum (LGM) at about 23–19thousand years ago (ka)8, the archaeological culture seen at Yana RHS had disappeared from northeastern Siberia. Instead, archaeological lithic assemblages at this time are dominated by a distinctive microblade technology, which spread in a time-transgressive manner north and east out of the Amur region9,10. This tradition did not reach Chukotka or cross the Bering land bridge (Beringia) until the end of the Pleistocene—later than the earliest known sites in the Americas. Changes in material culture in northeastern Siberia continued into the late Holocene epoch, but it remains debated whether these successive cultural complexes repre-sent insitu technological evolution or migrations of distinct groups of people. In the case of the latter, it is unclear how these groups may have been related to each other, to contemporary Siberians, or to Native Americans, whose ancestors may have emerged in this region (or, at the very least, traversed it en route to Beringia).  Fig. 1 | Genetic structure of ancient northeast Siberians. a, Sampling locations of newly reported individuals, as well as select previously published individuals with dates in parentheses. b, Sample dates. c, Principal component (PC) analysis of 257 ancient individuals projected onto a set of 1,541 modern Eurasian and American individuals. Abbreviations in group labels: UP, Upper Palaeolithic; LP, Late Palaeolithic; M, Mesolithic; EN, Early Neolithic; MN, Middle Neolithic; LN, Late Neolithic; N, Neolithic; EBA, Early Bronze Age; LBA, Late Bronze Age; IA, Iron Age; PE, Palaeo-Eskimo; MED, Medieval; HG, huntergatherer. |
|
|
|
Post by Admin on Dec 19, 2021 20:20:20 GMT
To investigate these questions, we used single-end shotgun sequenc-ing to generate whole genomes of 34 ancient individuals with associ-ated radiocarbon ages that range from 31,600 to 600calibrated years before present (Fig.1, Supplementary Information1, 2, Supplementary Tables1, 2). Our data include samples from ancient individuals that are key for understanding Siberian population history: two high-qual-ity genomes sequenced from fragmented milk teeth (Supplementary Information2) that were recovered from Yana RHS (Yana1 and Yana2, which produced genomes at 25× and 7× coverage, respectively), which are the oldest Pleistocene human remains found to date at such high latitude; a high-coverage genome (14× coverage) of an individual from the Duvanny Yar site at the Kolyma River (Kolyma1), dated to about 9.8ka; 14 genomes from ancient individuals from sites in far eastern Chukotka (Ekven and Uelen) and the northern coast of the Sea of Okhotsk (Ol’skaya, near the city of Magadan), ranging in age between 3 and 2ka; 6 individuals from the approximately 7,600-year-old site of Devil’s Gate Cave in northern East Asia11; 6 genomes of individuals from southern Siberia (Ust’Belaya in the Lake Baikal region) dating to between 6.5ka and 0.6ka; an individual from the Yana River but from a different locality than Yana RHS, here named ‘Young Yana’ (dated to 0.8ka); as well as 4 individuals from the Levänluhta site in southwestern Finland, dated to approximately 1.5ka. We analysed these data in the context of large panels of previously published ancient and present-day individuals (Supplementary Information3, Supplementary Tables3, 4).
Upper Palaeolithic peoples at Yana RHSThe Yana RHS human remains represent the earliest direct evidence of human presence in northeastern Siberia, a population that we refer to as Ancient North Siberians (ANS). The two Yana RHS individuals were unrelated males who carry mitochondrial haplogroup U (which is predominant among ancient West Eurasian hunter-gatherers) and Y chromosome haplogroup P1, which is ancestral to haplogroups Q and R—which are widespread among present-day Native Americans and Eurasians, respectively12,13 (Extended Data Fig.2, Supplementary Information4, 5, Supplementary Table1). Genetic clustering using outgroup-f3 statistics demonstrates broad genetic similarities with a wide range of present-day populations across Northern Eurasia and the Americas. This contrasts with other Upper Palaeolithic Eurasians, such as those from Sunghir14 and Tianyuan15—who share similar amounts of genetic drift with present-day populations that are geographi-cally more restricted to western Eurasia and East Asia, respectively (Extended Data Fig.3). Despite their extreme northeastern Siberian geographical location, the Yana RHS individuals are genetically closer to West Eurasians, although symmetry tests using f4 statistics reject tree-like clade relationships between them, early West Eurasians (Sunghir) and East Asians (Tianyuan) (Extended Data Fig.3d,e, Extended Data Table1, Supplementary Information6). Using admix-ture graphs and outgroup-based estimation of mixture proportions, we find that ANS can be modelled as early West Eurasian with an approximately 22% contribution from early East Asians (Extended Data Fig.3f, Supplementary Information6). Demographic modelling of the high-coverage individual Yana1 using a site-frequency-spectrum-based framework indicates an early divergence of the ANS lineage at about 39ka (95% confidence interval (CI) 32.2–45.8ka), concomitant with substantial gene flow (approximately 29%; 95% CI 21.3–40.1%) from East Asians, an event that probably occurred very soon after the latter diverged from West Eurasians 43.1ka (95% CI 33.4–48.6ka) (Fig.2a, Supplementary Information7). Thus, the ANS population represents a distinct lineage with affinities to both early West Eurasians and early East Asians, albeit in a 2:1 ratio. These complex relationships among early Eurasian groups are also supported by the presence of East Asian ancestry and mitochondrial haplogroup M in Western Europe by 35ka16,17. Finally, we estimate about 2% Neanderthal ancestry in the Yana RHS genomes, which is contained in longer genomic tracts than in present-day individuals, as also shown for other Upper Palaeolithic Eurasians14,16 (Supplementary Information6).
We next investigated how the Yana RHS individuals relate to an ancient Siberian population represented by the Mal’ta individual (dated to 24ka) from the Lake Baikal region (previously termed ‘Ancestral North Eurasians’), from which Native Americans derive around 40% of their ancestry18. We find that the Mal’ta individual shares more alleles with the Yana RHS individuals than with other West Eurasian hunter-gatherers (for example, f4(Mbuti, Mal’ta; Sunghir, Yana)> 0; Z=3.99) (Extended Data Table1, Supplementary Information6). The Mal’ta and Yana RHS individuals also exhibit a similar pattern of genetic affinities to both early West Eurasians and East Asians, consistent with previous studies19,20. In admixture graphs, the Mal’ta individual can be successfully fit as a descendant of the ANS lineage, with a minor contribution from an early Eurasian lineage that is ances-trally related to Late Palaeolithic hunter-gatherers from the Caucasus (Extended Data Fig.3e, f). The Ancestral North Eurasian lineage of the Mal’ta individual can thus be considered a descendant of the ANS line-age, and our results therefore suggest that—by 31.6ka—people related to the ANS were probably widespread across northeastern Eurasia.The two Yana RHS individuals were contemporaneous, which pro-vides an opportunity to investigate relatedness and levels of inbreeding
at this remote Upper Palaeolithic settlement. We find that the two indi-viduals were not closely related and did not exhibit signatures of recent inbreeding, with a moderately large estimate of recent effective-popula-tion size of up to 500 individuals (Extended Data Fig.4, Supplementary Information4, 5). Our results mirror those observed at Sunghir, an early (approximately 34,000-year-old) European Upper Palaeolithic site located about 4,500km southwest of Yana RHS, which reinforces the view that wide-ranging mate exchange networks were present among Upper Palaeolithic foragers across the pre-LGM landscape14.
|
|
|
|
Post by Admin on Dec 20, 2021 2:00:40 GMT
Ancient Palaeo-Siberians and Native AmericansFollowing the occupation at Yana RHS, there is an absence of archaeo-logical sites in northeastern Siberia. The area is re-occupied in the later part of the LGM, about 20ka, when sites preserving a very distinctive stone tool technology become widespread. This gap in the archaeo-logical sequence is critical, as it was within the intervening period that the population ancestral to Native Americans emerged18,21, although no genomes from individuals of this age have been recovered in north-eastern Siberia to date. We find that the Kolyma1 individual (dated to 9.8ka)—who represents a lineage that formed after about 30ka, which we name ‘Ancient Palaeo-Siberian’—documents the first major genetic shift that we observe in the region (Extended Data Fig.5). Principal component analysis, outgroup-f3statistics and mitochondrial DNA and Y chromosome haplogroups (G1b and Q1a1b, respectively) demon-strate a close affinity between Ancient Palaeo-Siberians and present-day Koryaks, Itelmen and Chukchis, as well as with Native Americans (Extended Data Fig.5, Supplementary Information6). Admixture graph modelling shows that the Kolyma1 individual derives from a mixture of East Asian and ANS-related ancestry, similar to that found in Native Americans, although the East Asian contribution is greater in Kolyma1 than in Native Americans (75% versus 63%) (Extended Data Fig.3f, Supplementary Information6). For both Ancient Palaeo-Siberians and Native Americans, ANS-related ancestry is more closely related to Mal’ta than to the Yana individuals (Extended Data Fig.3f), which rejects the hypothesis that the Yana lineage contributed directly to later Ancient Palaeo-Siberians or Native American groups.  We then estimated the demographic parameters of population history models that included the Ancient Palaeo-Siberian Kolyma1 and Native Americans, represented either by Ancient Beringians21 (the Upward Sun River 1 (USR1) individual) or present-day Native Americans (Karitiana individuals; Supplementary Information7). We find that their ancestors diverged at about 30ka (95% CI 26.8–36.4ka) from present-day East Asians (represented by Han individuals), con-sistent with previous results21; the subsequent divergence of Ancient Palaeo-Siberians from the population ancestral to Ancient Beringians or Native Americans occurred at about 24ka (95% CI 20.9–27.9ka) (Fig.2, Supplementary Information7). We infer that both Ancient Palaeo-Siberians and Ancient Beringians received ANS-related gene flow at a similar time (Kolyma1 at 20.2ka (95% CI 15.5–23.7ka) and USR1 at 19.7ka (95% CI 13.3–23.5ka)). This gene flow amounts to 16.6% (95% CI 7.5–22.2%) of ANS ancestry into the Kolyma1 individ-ual and 18.3% (95% CI 9.8–20.3%) of ANS ancestry into USR1, which is comparable to the estimates we obtained using admixture graphs. An alternative model with a single admixture pulse in the ancestral popula-tion of the Kolyma1 and USR1 individuals showed a comparable likeli- hood (Supplementary Information7), but differences in the estimated proportions of ANS-related ancestry between Kolyma1 and USR1 favour the two-independent-pulses model. The Kolyma1 individual thus represents the closest relative to the ancestral Native American population in northeastern Siberia that has been found to date. Changes in climatic conditions are commonly put forward as principal drivers of Pleistocene population movement and regional abandonment in Siberia. We used palaeoclimatic modelling to infer geographical locations in Siberia that were suitable for human occupa-tion between 48 and 12ka to further investigate this hypothesis. When humans were present at Yana RHS, interstadial climatic conditions meant that a large stretch of the Arctic coast of northeastern Siberia was suitable for human occupation (Extended Data Fig.6, Supplementary Information8). Conditions in the region became harsher during the LGM, which is consistent with the absence of archaeological sites in the area at this time. However, our models suggest the existence of a refugium across southern Beringia during the LGM (for example, see Alaska in the ‘22ka’ panel of Extended Data Fig.6a), consistent with previous reports22. Therefore, a possible scenario for gene flow during the formation of the early Native American and Ancient Palaeo-Siberian gene pools may have involved early ANS-related groups occu-pying southern Beringia during the LGM, and subsequently admixing with EastAsian-related peoples who expanded northwards towards the end of the LGM. This scenario would also be consistent with a divergence of Ancient Beringians from ancestral Native Americans in eastern Beringia rather than in Siberia, which is supported by genetic data (scenario 2 in ref. 21). Alternatively, the closer affinity of both Kolyma1 and Native Americans to Mal’ta, rather than the Yana RHS individuals, could suggest a more-southwesterly location (Lake Baikal region) for the admixture, with a northward expansion after the LGM. The alternative scenario is supported by archaeological evidence for a movement south during the LGM, although the genetic isolation that is observed between Asians and ancestral Native Americans after about 23ka would have required the maintenance of a structured population during the LGM—this, in turn, implies that Ancient Palaeo-Siberians and ancestral Native Americans occupied different refugia. Regardless, our results support the broader implication that glacial and post-glacial climate change was a major driver of human population history across northern Eurasia. |
|






