|
|
Post by Admin on Jul 19, 2018 18:19:39 GMT
Instead of a tree, a better metaphor would be a trellis, branching and remixing far back into the past, says Reich, whose work indicates that the idea of race is a very fluid, ephemeral concept. However, he is adamant that it is a very real one and takes issue with those geneticists who argue that there are no substantial differences in traits between populations. “This is a strategy that we scientists can no longer afford and that in fact is positively harmful,” he argues. Plenty of traits show differences between populations: skin colour, susceptibility to disease, the ability to breath at high altitudes and the ability to digest starch. More to the point, uncovering these differences is only just beginning. Many more will be discovered over the decades, Reich believes. Crucially, we need to be able to debate the implications of their presence at varying levels in different populations. That is not happening at present and that has dangerous implications.  “If as scientists we wilfully abstain from laying out a rational framework for discussing human differences, we will leave a vacuum that will be filled by pseudoscience, an outcome that is far worse than anything we could achieve by talking openly,” says Reich. The genome revolution provides us with a shared history, he adds. “If we pay proper attention, it should give us an alternative to the evils of racism and nationalism and make us realise that we are all entitled equally to our human heritage.” • Who We Are and How We Got Here: Ancient DNA and the New Science of the Human Past by David Reich is published by Oxford University Press (£20). To order a copy for £17 go to guardianbookshop.com or call 0330 333 6846. Free UK p&p over £10, online orders only. Phone orders min p&p of £1.99 |
|
|
|
Post by Admin on Oct 27, 2018 18:20:19 GMT
 The genomic revolution is transforming our understanding of modern humans. Geneticists like David Reich have made astounding advances in genomics, which is proving to be as important a field as archeology or linguistics for understanding our ancestry. Reich arrives at Town Hall to enlighten us with provocative research and unparalleled scientific studies that have yielded revolutionary findings—compiled in his book Who We Are and How We Got Here. He reveals the hidden story of our species, offering insight on DNA studies that reveal deep inequalities among different populations, between the sexes, and among individuals. Reich suggests that there might very well be biological differences among human populations—many of which are unlikely to conform to common stereotypes. Join Reich for a captivating glimpse into the origins of humankind, and a chance to apply the genetic findings of the past to our lives today. David Reich is Professor of Genetics at Harvard Medical School and a Howard Hughes Medical Institute Investigator, with a reputation as one of the world’s leading pioneers in analyzing ancient human DNA. In a 2015 article in Nature, he was named one of ten people who matter in all of the sciences for his contribution to transforming ancient DNA data “from niche pursuit to industrial process.” Awards he has received include the Newcomb Cleveland Prize from the American Association for the Advancement of Science, as well as the Dan David Prize in the Archaeological and Natural Sciences for his computational discovery of intermixing between Neanderthals and Homo sapiens. |
|
|
|
Post by Admin on May 26, 2019 17:48:02 GMT
David Reich describes the three biggest discoveries about European pre-history that were revealed through genome-wide analyses of 51 ice-age-era humans. Reich argued that, based on currently available ancient DNA, all of the main Indo-European daughter branches, apart perhaps from Anatolian, may have expanded from the Yamnaya horizon on the Pontic-Caspian Steppe in Eastern Europe via a series of massive migrations soon after the Neolithic. In other words, even Indo-Iranian, including Indo-Aryan, languages might be from the Yamnaya horizon. The team’s findings, published May 2 in Nature, reveal the disappearance and reappearance of a group that formed part of the ancestry of today’s Europeans, describe when and how Europeans acquired DNA from people in the Near East and show that the amount of Neanderthal DNA in modern human genomes has shrunk over the millennia, likely because the DNA was evolutionarily disadvantageous.  “This study raises by about tenfold the number of ice age hunter-gatherers for which there is ancient DNA, and in so doing, it makes it possible to track genetic change over time,” said David Reich, professor of genetics at Harvard Medical School and co-senior author of the paper. “Prior to this work, we had a static view of the first 30,000 years of modern human history in Europe. Now we can begin to see how people moved around and mixed with one another during this period,” said co-senior author Svante Pääbo, director at the Max Planck Institute for Evolutionary Anthropology in Leipzig, Germany. “Archaeologists have gathered a tremendous amount of information about cultural change in ice age Europe,” said co-senior author Johannes Krause, professor of archaeo- and paleogenetics at the University of Tübingen and director at the Max Planck Institute for the Science of Human History in Jena, Germany. “This study connects our knowledge of the material culture of ice age populations—including people who produced elaborate figurines and the world’s first musical instruments—to their biological identities.”  “These bones are very, very old,” said Qiaomei Fu, first author of the paper and a faculty member of the Key Laboratory of Vertebrate Evolution and Human Origins at the Chinese Academy of Sciences in Beijing. “It was an immense privilege to work on them, but their age also made it extremely difficult to extract high-quality genetic information from them.” Fu, who started working on the project as a graduate student in Pääbo’s lab and then continued it as a postdoctoral researcher in Reich’s lab, had to contend not only with the fact that the bones’ original DNA had degraded over time, but also that it had likely been contaminated by the genetic material of microbes and present-day people. She “screened and screened and screened” the bones, applied the best available enrichment and decontamination techniques and discarded all but the best-quality samples. 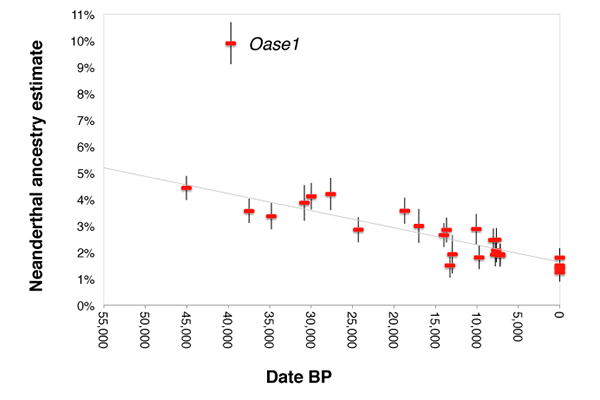 The second major surprise came when the researchers found another previously unknown population turnover. The analysis showed that starting about 14,000 years ago, Europeans started to show a genetic relationship to present-day Near Easterners. “Normally, when we think about genetic connections between these parts of the world, we look at the introduction of agriculture about 8,500 years ago, when farmers from the Near East came into Europe and replaced hunter-gatherers,” said Krause. “But this is a big event that occurred 6,000 years earlier.” Finally, the researchers found that a subset of European hunter-gatherers acquired some East Asian-related DNA after 14,000 years ago, suggesting an additional flow into Europe. “We see that the founding European populations persisted over the last glacial maximum some 25,000 to 14,000 years ago, but afterward there were dramatic re-jiggerings as people moved back from warm-weather refuges in the southwest and the southeast, transforming the human landscape of Europe,” said Pääbo. “It is amazing how ancient DNA now starts to provide us with a detailed account of the earliest history of present-day Europeans.” |
|
|
|
Post by Admin on Apr 12, 2020 19:54:51 GMT
Tales of Human History Told by Neandertal and Denisovan DNA That Persist in Modern Humans Where did we humans come from? When did we become the dominant species on the planet? 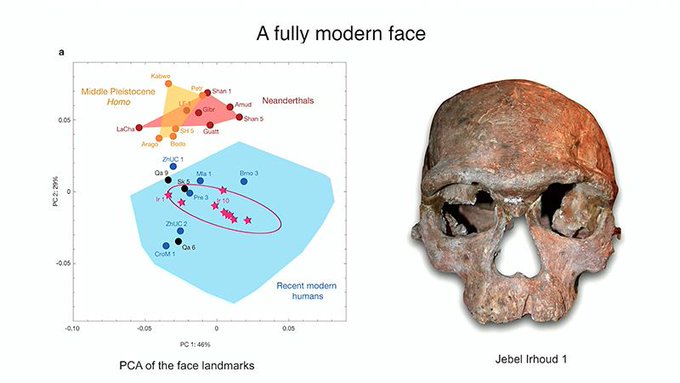 Experts take you on an exploration of the last half-decade of new evidence from ancient DNA, fossils, archaeology and population studies that has updated our knowledge about The Origins of Today's Humans. Recorded on 02/21/2020. [Show ID: 35716] |
|
|
|
Post by Admin on Apr 25, 2021 21:14:58 GMT
Predicting skeletal stature using ancient DNA Samantha L Cox, Hannah Moots, Jay T Stock, Andrej Shbat, Bárbara D Bitarello, Wolfgang Haak, Eva Rosenstock, Christopher B Ruff, Iain Mathieson doi: doi.org/10.1101/2021.03.31.437877Abstract Objectives Ancient DNA provides an opportunity to separate the genetic and environmental bases of complex traits by allowing direct estimation of genetic values in ancient individuals. Here, we test whether genetic scores for height in ancient individuals are predictive of their actual height, as inferred from skeletal remains. We estimate the contributions of genetic and environmental variables to observed phenotypic variation as a first step towards quantifying individual sources of morphological variation. Materials and Methods We collected stature estimates and femur lengths from West Eurasian skeletal remains with published genome-wide ancient DNA data (n=167, dating from 33,000-850 BP). We also recorded genetic sex, genetic ancestry, date and paleoclimate data for each individual, and δ13C and δ15N stable isotope values where available (n=67). Results A polygenic score (PRS) for height predicts 6.8% of the variance in femur length in our data (n=117, SD=0.0068%, p<0.001), controlling for sex, ancestry, and date. This is consistent with the predictive power of height PRS in present-day populations and the low coverage of ancient samples. Comparatively, sex explains about 15% of the variance in femur length in our sample. Environmental effects also likely play a role in variation, independent of genetics, though with considerable uncertainty (longitude: R2=0.0317, SD=0.009, p=0.019). Discussion Polygenic scores explain a small but significant proportion of the variance in height in ancient individuals, though not enough to make useful predictions of individual phenotypes. However, environmental variables also contribute to phenotypic outcomes and understanding their interaction with direct genetic predictions will provide a framework with which to model how plasticity and genetic changes ultimately combine to drive adaptation and evolution. Introduction One of the central goals of biological anthropology is to understand the environmental and genetic contributions to phenotypic change. Skeletal series covering long time periods and diverse environments are informative, but limited by inability to separate these contributions—a limitation that can now be addressed with ancient DNA. The ability to generate genetic data from skeletal remains has had an enormous impact on studies of human history. By identifying genetic links among individuals and populations, ancient DNA allows us to reconstruct demographic histories on both large and small scales (Skoglund & Mathieson, 2018; Racimo et al., 2020), as well as the effects of natural selection (Marciniak & Perry, 2017). To learn about complex trait evolution, ancient DNA can be combined with information from genome-wide association studies (GWAS). These studies, now involving millions of individuals, have identified large numbers of single-nucleotide polymorphisms (SNPs) associated with hundreds of phenotypes (Visscher et al., 2017). Though the effect of any individual variant is typically small, the effects of many SNPs can be summed to produce a polygenic score (PRS) which can be thought of as an estimate of an individual’s genetic potential or risk for a particular phenotype (Torkamani et al., 2018). Height is a polygenic trait which is highly heritable and easy to measure. GWAS have identified thousands of SNPs significantly associated with height (Yengo et al., 2018). Each one has only a tiny effect—±1-2mm per SNP—but combined explain around 25% of the phentoypic variance in present-day populations of European ancestry (Yengo et al., 2018). Contingent upon reasonable preservation of skeletal remains, stature estimation from long bones is relatively straightforward, and there is an excellent record of changes in stature in many parts of the world (Bogin & Keep, 1999; Cohen & Crane-Kramer, 2007; Ruff, 2018; Rosenstock et al., 2019). On a population level, changes in polygenic score in Europe computed from ancient DNA largely track trends in stature in the European skeletal record (Cox et al., 2019). However, environmental effects on stature can still be large, as shown by 20th century secular trends (NCD Risk Factor Collaboration, 2016), and are not confined to recent history. For example, during the Bronze Age, genetic predictions suggest increasing stature, but estimated skeletal heights actually decrease (Cox et al., 2019). Polygenic scores have three main limitations. First, due to incomplete correction of population stratification in GWAS, they can capture environmental variation in present-day populations leading to ancestry-related biases (Sohail et al., 2019; Berg et al., 2019). Second, their accuracy decreases as genetic distance from the present-day European ancestry GWAS populations increases (Martin et al., 2019). Finally, their accuracy can be reduced by environmental differences, even within-population (Mostafavi et al., 2020). We therefore set out to test whether polygenic scores based on present-day GWAS predict height in ancient individuals by collecting data for individuals with both ancient DNA and skeletal measurements. This allows us to assess the validity of complex trait prediction in ancient individuals, and whether we can use this approach to understand the relationship between genetic and environmental components of stature and, perhaps, of other phenotypes. |
|






