|
|
Post by Admin on Aug 19, 2019 18:47:45 GMT
The Merovingian period refers to the dynasty of a family known as the "long-haired kings" who ruled over Germanic people called the Franks from the middle of the 5th century until 751AD.  At the height of the dynasty, most of western Europe was under Merovingian rule and the realm is often regarded as the most powerful of the western European states following the fall of the Western Roman Empire.  Since the discovery of the mysterious skeleton, more excavations have unearthed pieces of Merovingian pottery and an area that is thought to have been an old kitchen.  All these finds will be examined by experts from the French National Institute for Preventative Archaeological Research. This institute is the Henri-Martin museum in Cahors, where the finds will also be kept from now on. The Merovingians were a powerful dynasty that ruled over the confederation of Germanic people known as the Franks for nearly 300 years, from the 5thto 8thcenturies. They exercised control over much of modern-day France, Germany, Switzerland, Austria and the Low Countries. 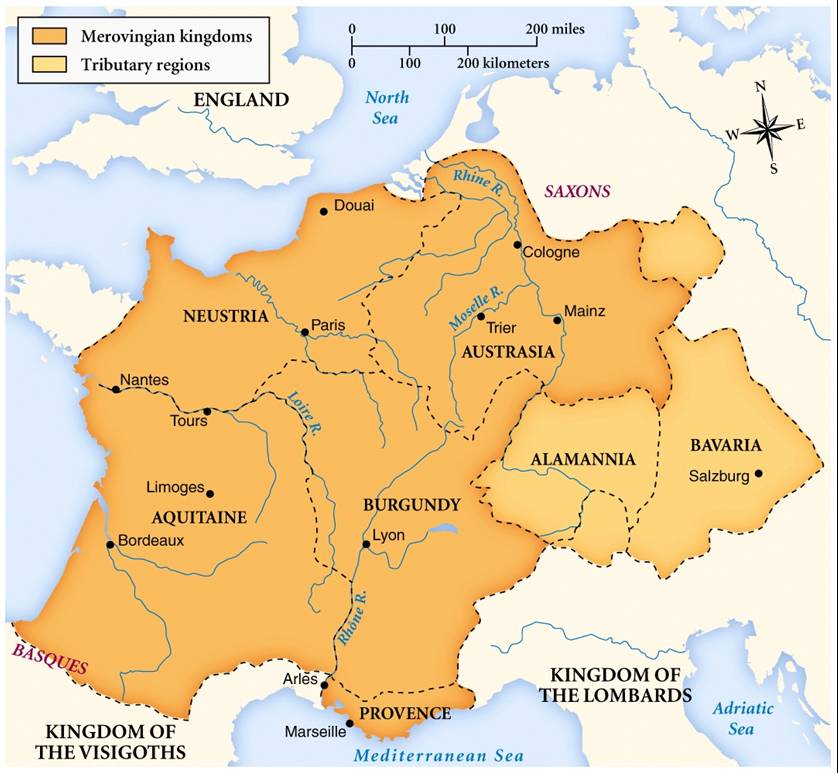 The Merovingian kingdoms were one of the most powerful political powers to rise after the collapse of the Western Roman Empire. Their kings as sometimes referred to as the “long-haired kings” as they kept long hair to distinguish them among the Franks, who commonly kept their hair short. The analysis of ancient DNA recovered from archaeological remains can be used to reconstruct kinship among the occupants of a necropolis and provide a more detailed portrait of the community considered. Such palaeogenetic analyses have been conducted on sarcophagi excavated from the Merovingian necropolis in Jau-Dignac et Loirac (7th–8th century AD, Aquitaine, southwest France). The genetic study consisted of the analysis of mitochondrial DNA and nuclear STRs (Short Tandem Repeats) from nine skeletons deposited in three grouped sarcophagi. Only data concerning the mitochondrial genomes could be obtained, and six different mitochondrial lineages were retrieved from eight samples. Our analyses permitted a high confidence characterisation of maternal relationships between individuals deposited in the same sepulchre. These results are important and novel for the period and region and argue that individuals were grouped inside sarcophagi according to relationship criteria. The presence of perinatal remains in one sarcophagus was particularly striking because access to this type of funerary structure during this period was generally reserved for older children. Moreover, we demonstrated genetically that the perinatal remains were not related maternally to two women found in the same sarcophagus (whereas the maternal relationship between the two young women could be determined), and we proposed different possible explanations for this unexpected observation. Overall, archaeological, anthropological and genetic data suggest that the Jau-Dignac et Loirac necropolis groups together the closely and distantly related members of a High Middle Ages familia. Our ancient DNA analyses note the important contribution of palaeogenetic analyses to archaeological kinship studies. Journal of Archaeological Science Volume 41, January 2014, Pages 399-405 |
|
|
|
Post by Admin on Aug 19, 2019 19:48:20 GMT
A Merovingian necropolis is an ancient burial ground. Many of these ancient ‘cities of the dead’ infact resemble small cities, containing elaborate tombs with multiple chambers and intricate details. The burial practices of the Merovingians are currently unknown. Much of what we know about them comes from studying their graves. The study highlighted here focused on the Jau Dignac et Loirac necropolis. The primary goal was determining the biological relationship between the remains from a specific sections of the tomb. They opted for genetic analyses, as it is the most conclusive way to determine biological relationships between skeletal remains. 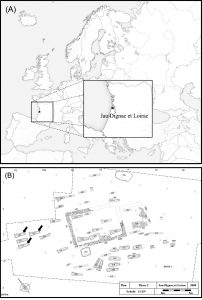 (A) Localisation of the Jau Dignac et Loirac site and (B) localisation of sarcophagi 169, 170 and 171 (Image from Deguilloux MF et al. (2014) J. Archaeol. Sci.) Investigators looked at biological relationships between skeletal remains of 9 individuals using mitochondrial DNA analyses. The high copy number, rapid evolution rate and strict maternal inheritance makes mitochondrial DNA the most suitable and informative DNA type for looking at ancient human remains. There are three regions of the mtDNA that can be analyzed – two hypervariable regions (HVR1 and HVR2) and the coding region. Investigators obtained the full HVR1 sequence from six skeletons, and a partial HVR1 sequence from the three remaining skeletons. 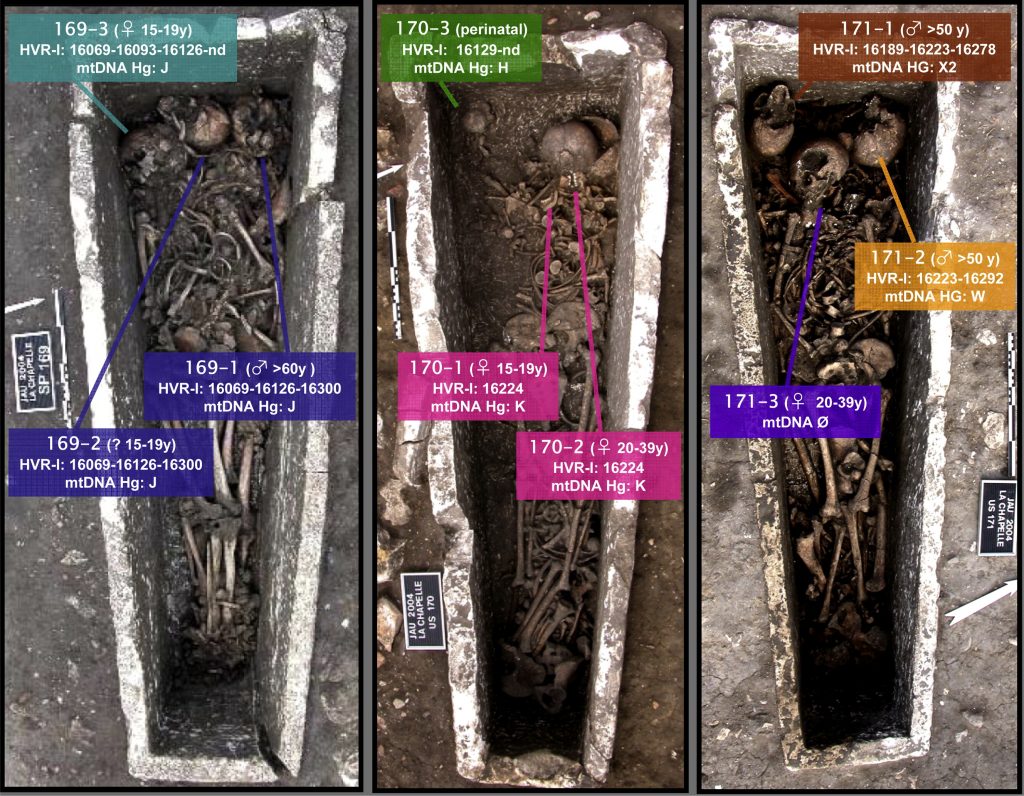 Details of sarcophagi 169, 170 and 171 and mitochondrial data obtained for different human remains Image from Deguilloux MF et al. (2014) J. Archaeol. Sci.) Sarcophagus 169 This sarcophagus contained remains of three people: an older male and two young adults one female and one of undetermined gender. According to mtDNA profiles the older man and the young female were maternally related to each other. But the third individual was not maternally related. Sarcophagus 170 Remains of three individuals were found in this coffin: a young, likely female adult, a young female and an infant. The two women shared a mtDNA profile, however the infant was not maternally related to either of the females. Sarcophagus 171 This coffin also contained remains of three people: two males, and a young female. MtDNA analyses concluded that the two males were not maternally related. However, it’s possible that there’s some familial relationship between the men (e.g. paternal relationship). 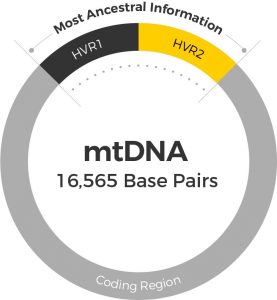 Mitochondrial DNA has numerous advantages over nuclear DNA for the analysis of ancient remains. But, mtDNA can only uncover maternal relationships, a distinct disadvantage of this technique. This study established that maternal relationships existed between some of the skeletons in the shared coffins. But others showed no maternal relationships. It’s possible that paternal relationships exist between the individuals buried at Jau-Dignac et Loirac. This study did not examine paternal relationships. The DNA tests conducted in this study have identified six different mtDNA profiles from the Merovingian rulers of the 7th– 8thcentury AD. If you have taken the DNA Maternal Ancestry Test, you can compare your mtDNA against them to see if you share a similar mtDNA profile. |
|
|
|
Post by Admin on Aug 20, 2019 18:19:39 GMT
Ancient DNA has the potential to identify precise kinship patterns between groups of skeletons. Maternal, paternal and general family relationships can be revealed by mitochondrial DNA typing (mtDNA), the examination of markers on the Y chromosome and autosomal STR (Short Tandem Repeats) genotyping, respectively. Autosomal DNA markers are powerful discriminators in the study of close parental relationships; however, this technique is hampered by the main problems inherent in aDNA research, which are template degradation and contamination with modern exogenous DNA (Hofreiter et al., 2001; Pääbo et al., 2004). Satisfactory results for nuclear DNA are dependent on exceptional DNA conservation in ancient human remains and are most often linked to specific environmental parameters that are favourable to biomolecule conservation, such as dryness (Gamba et al., 2010) or cold (Gill et al., 1994; Keyser-Traqui et al., 2003; Haak et al., 2008). 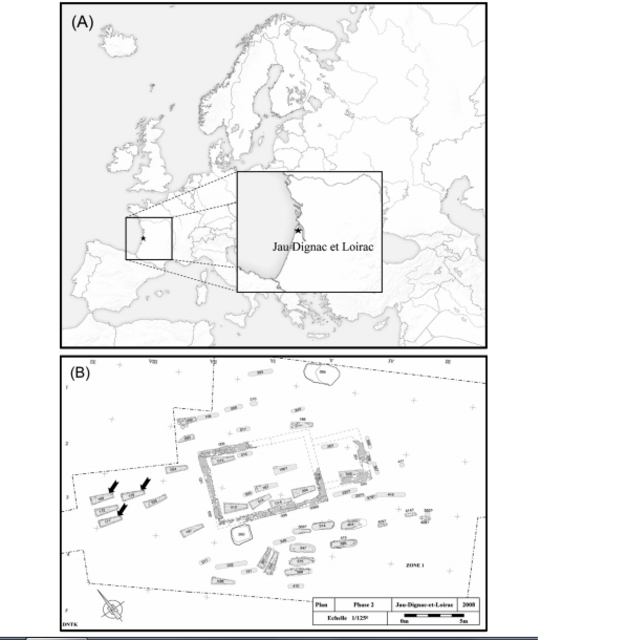 Fig. 1.. (A) Localisation of the Jau Dignac et Loirac site and (B) localisation of sarcophagi 169, 170 and 171 in the Merovingian necropolis (highlighted by black arrows; Cartron and Castex, 2010). Consequently, mtDNA has often been the preferred target in aDNA studies because mtDNA is more likely to be preserved in ancient remains due to its higher copy number than nuclear DNA. The second methodological problem of aDNA studies is the ubiquitous risk of contamination with modern exogenous DNA, and this fact is especially true when working with human remains (Handt et al., 1996). Contaminant DNA can theoretically be introduced at any point during the process, from archaeological excavation to laboratory work (Yang and Watt, 2005). Consequently, the necessity of pre-laboratory contamination controls in contributing to the overall success of ancient DNA studies, in combination with procedures permitting contamination detection and ancient sequences authentification, is regularly confirmed (Cooper and Poinar, 2000; Yang and Watt, 2005; Deguilloux et al., 2011a). Although success has been achieved with material conserved in exceptional conditions, authenticated kinship studies with archaeological specimens remain rare (for example, Gill et al., 1994; Dudar et al., 2003; Keyser-Traqui et al., 2003; Capellini et al., 2004; Mooder et al., 2005; Rudbeck et al., 2005; Bouwman et al., 2008; Haak et al., 2008; Gamba et al., 2010).  The potential of aDNA to identify kinship patterns between groups of skeletons, and thereby organisation of ancient human societies, has been tested on sarcophagi from the Merovingian necropolis in Jau-Dignac et Loirac (7the8th century AD, Aquitaine, southwest France). The excavations conducted by I. Cartron and D.Castex in this Medoc peninsula site (Fig. 1A) have characterised several occupations of the site (Cartron and Castex, 2006). The first occupation consisted of the construction of a Gallo-Roman temple, which was transformed and then slowly abandoned (1st-5th century AD). After two centuries of no occupation, the temple was in ruins and was transformed into a small funerary church associated with a small necropolis that functioned during the High Middle Ages (7the11th century AD). The church was later destroyed, replaced by a chapel (12the15th century AD) associated with a small cemetery and used until the 18th century AD. The long occupation of the site provides important data for understanding the genesis of funerary places and places of worship (Cartron and Castex, 2010).  The High Middle Ages necropolis (7th-11th century AD) brings together 74 sepulchres (including 20 sarcophagi) closely grouped inside and around the church (Cartron and Castex, 2009; Fig. 1B). Nonmetric trait analysis showed a high biological homogeneity of the individuals found in the necropolis (Laforest et al., 2011). Rare discrete trait sharing was particularly observed for a specific group of 7 sarcophagi localised in the western part of the necropolis (Fig. 1B). The grouping and specific alignment of these sarcophagi in the western part of the necropolis, contemporaneous use, discrete trait sharing, sarcophagus type and the homogeneity of funerary practices highlighted the striking singularity of this group (Cartron and Castex, 2010). Moreover, the sarcophagi were reused several times, with two to four individuals buried in the same structure, and the exceptional presence of a baby in one of the sarcophagi was observed. These observations led to the question of kinship between the individuals buried in the same sarcophagus or among sarcophagi. Consequently, we decided to perform a genetic study based on nuclear STRs and mtDNA analysis on 9 skeletons buried in 3 different sarcophagi to uncover the familial relationship and geographic origin of the individuals. The precautions taken during sampling and palaeogenetic analyses enabled us to propose reliable genetic arguments that lead to a better understanding of the funerary function of the necropolis. |
|
|
|
Post by Admin on Aug 21, 2019 8:05:48 GMT
The skeletal material under study comes from recent excavations directed by I. Cartron and D. Castex in Jau-Dignac et Loirac (Aquitaine; Cartron and Castex, 2010). The 9 individuals buried in sarcophagi 169, 170 and 171 were analysed in the present study (Fig. 2). Sarcophagus 169 groups 3 successively buried individuals: one male (older than 60 years old, individual 169-1) and two young adults (15e19 years old, individuals 169-2 and 169-3), one female and one undetermined (Fig. 2). The different burials were successive and did not cause many disturbances. Only the craniofacial bloc, pelvis and right femora of the first individual were moved along the sarcophagus’ left side before the burial of the second individual. The burial of the third individual did not involve manipulations of the individuals previously present in the structure. Sarcophagus 170 contained the remains of 3 successively buried individuals: a young, likely female adult (15e19 years old, individual 170-1), a young female (20e29 years old, individual 170-2) and an infant (still-born or deceased just after birth, individual 170-3) (Fig. 2). When the sarcophagus was reused for the burial of the second female, a majority of the bones from the inferior part of the first female’s body was removed and then placed over the second individual.  Fig. 2.. Details of sarcophagi 169, 170 and 171 and mitochondrial data obtained for different human remains (Ø: not determined; images from Cartron and Castex, 2010). A strikingly original feature of this sarcophagus is the presence of infant remains because young children were often subject to distinct funeral practices in this period. Indeed, young children were generally buried individually in containers such as wooden coffins. The off-centre position of the second female and her proximity to the infant suggested that both individuals were buried simultaneously Finally, sarcophagus 171 also contained 3 successively buried individuals: two males (older than 50 years old, individuals 170-1and 171-2) and a young female (20e39 years old, individual 171-3) (Fig. 2). The skeletons of both males were initially laid on top on one another in the sarcophagus and were manipulated before the placement of the female. The superior parts of the male skeletons were moved to the right superior corner of the sarcophagus, whereas the inferior parts of the skeletons were grouped in the inferior part of the structure. The absence of superimposition between different sepulchres encountered in the western part of the necropolis suggested surface signalling or the preserved memory of their localisation (potentially for successive burials).  Datations obtained for individuals in sarcophagus 171 (171-1: 659-773 AD; and 171-3: 879-992 AD) indicated that successive burials in the same structure occurred over relatively short period. Table 1 summarises the sample information. For each individual, a complete mandible was collected in situ during the excavation while using all precautions against contamination (including the wearing of mask and gloves) for sarcophagus 169. Mandibles from sarcophagi 170 and 171 were collected without a mask and gloves, but the mandibles were not washed or manipulated. All the samples were directly deposited into hermetically sealed sterile bags and immediately stored at 20 C. For adult skeletons, entire teeth in an excellent state of preservation without external fissures or cracks and still fixed to the jaw were selected for genetic analyses. For the immature individual, one hemi-mandible was used.  |
|
|
|
Post by Admin on Aug 21, 2019 18:16:49 GMT
We were able to retrieve complete HVR-I sequences for six out of nine individuals. SNPs from the coding region of mitochondrial DNA were obtained for 6 of the 7 samples analysed with the iPlex technique. Only one sample (female 171-3) did not produce reproducible data for the HVR-I sequence amplification (this sample could not be tested for SNP typing). Only partial HVR-I results were obtained from infant 170-3 and individual 169-3; however, satisfactory results were obtained for SNP typing.  Table 1 Table 1 presents the consensus mitochondrial HVR-I sequences and haplogroups retrieved from the human remains. The sequences were authenticated according to several criteria: (i) the consensus sequences were always deduced from multiple clones originating from different isolations and PCRs and consensus SNP typing was deduced from two replicates; (ii) despite low number of PCR negative controls revealed positive (5%), authenticated sequences were never found in negative controls and never matched the sequences of the people who handled the remains; (iii) the consensus HVR-I sequence determined for each segment showed clear patterns of deamination, a characteristic finding for aDNA that is presumed to be a result of post-mortem miscoding lesions; and (iv) different HVR-I sequences were obtained for individuals originating from the same sarcophagus or from different burials.  Among the eight individuals with complete or partial HVR-I sequences, six different HVR-I sequences were identified that corresponded to five different mtDNA haplogroups (J, H, K, X2 and W). A maternal genetic relationship was proposed for sarcophagi 169 and 170. The man (169-1) and young female (169-2) from sarcophagus 169 share the same mtDNA mutations (16069e16126e16300) that are characteristic of haplogroup J (confirmed by SNP typing). The specific haplotype J encountered in these individuals (characterised by mutation 16300) is rare because it has been encountered in only one individual from northwest Europe (in the database of Richards et al., 2000) and in nine French individuals from the Atlantic Pyrenean region (Richard et al., 2007). While the young individual 169- 3 can be assigned to the same haplogroup J (confirmed through SNP typing), he presented a distinct haplotype (mutations 16069e16093e16126) that negates a maternal relationship with other individuals from the same sarcophagus. We then conclude that both individuals first deposited in sarcophagus 169 (a man older than 60 years old and a young woman age 15e19 years old) were potentially maternally related. We can dismiss a maternal relationship for the third individual buried in the sarcophagus, although another type of familial relationship is possible.  Both women buried in sarcophagus 170 share the same HVR-I sequence (mutation 16224) and can be assigned to haplogroup H or K (because coding region polymorphisms are lacking). SNP typing could only be conducted on female 170-2 (no extra DNA extract was available for female 170-1), and she was characterised as belonging to haplogroup K. A possible hypothesis would be that both females that have the same HVR-I sequence are maternally related and consequently belong to haplogroup K. Haplogroup K, characterised by this specific and unique mutation 16224, is relatively rare in France because only three identical HVR-I sequences have been determined by Richard et al., (2007) in a sample of 868 French individuals (Vendée, Atlantic Pyrenees and Morbihan, i.e., western regions of France). The young women successively buried in sarcophagus 170 were then potentially maternally related. The partial HVR-I sequence obtained for the infant deposited in the same sarcophagus indicated a haplotype distinct from either female’s haplotype, which was characterised by mutation 16129. Mitochondrial SNPs clearly attributed the infant to haplogroup H, confirming the absence of a maternal relationship between the infant and both females encountered in sarcophagus 170.  No mitochondrial lineage sharing was observed for sarcophagus 171 because one man (171-1) had specific mutations of haplogroup X2 (16189e16223e16278), and the other man (171-2) had mutations characteristic of haplogroup W (16223e16292). Both haplogroups have been confirmed by SNP typing. No reproducible HVR-I sequence or SNP typing could be obtained for the woman (171-3) because of a lack of DNA extract. The X2 sequence determined in man 171-1 is rare in France because this sequence has been detected only once in a sample (from the Mediterranean region) reported by Richard et al. (2007). The W sequence found in man 171-2 has been described in 11 individuals originating from various French regions (Richard et al., 2007). Interestingly, both sequences have been described in ancient human samples from southwest France; the X2 sequence has been described for a Middle Neolithic individual from the Prissé-La-Charrière tumulus (Deguilloux et al., 2011b), and the W sequence has been described in a Merovingian funerary site from Dordogne (Peyrat site, 7th-9th century AD; unpublished data). We conclude that the two men (171-1 and 171-2) buried successively in sarcophagus 171 are not maternally related; however, we cannot exclude other types of familial relationship between the men. |
|




















