|
|
Post by Admin on Sept 9, 2019 23:24:39 GMT
Genetic Link Between Gender Dysphoria and Sex Hormone Signaling Madeleine Foreman, Lauren Hare, Kate York, Kara Balakrishnan, Francisco J Sánchez, Fintan Harte, Jaco Erasmus, Eric Vilain, Vincent R Harley  Abstract Context There is a likely genetic component to gender dysphoria, but association study data have been equivocal. Objective We explored the specific hypothesis that gender dysphoria in transgender women is associated with variants in sex hormone–signaling genes responsible for undermasculinization and/or feminization. Design Subject-control analysis included 380 transgender women and 344 control male subjects. Associations and interactions were investigated between functional variants in 12 sex hormone–signaling genes and gender dysphoria in transgender women. Setting Patients were recruited from the Monash Gender Clinic, Monash Health, Melbourne, Australia, and the University of California, Los Angeles. Patients Caucasian (non-Latino) transgender women were recruited who received a diagnosis of transsexualism [Diagnostic and Statistical Manual of Mental Disorders (DSM)-IV) or gender dysphoria (DSM-V)] pre- or postoperatively. Most were receiving hormone treatment at the time of recruitment. Main Outcome Measured Genomic DNA was genotyped for repeat length polymorphisms or single nucleotide polymorphisms. 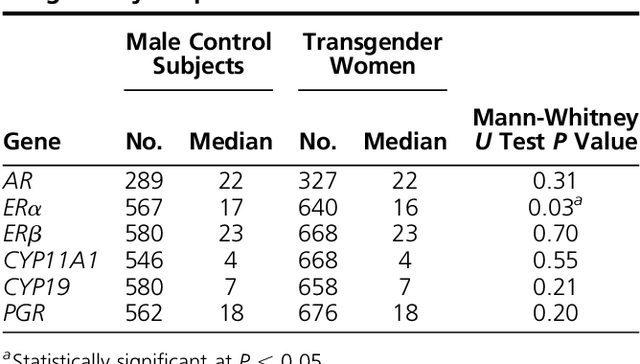 Results A significant association was identified between gender dysphoria and ERα, SRD5A2, and STS alleles, as well as ERα and SULT2A1 genotypes. Several allele combinations were also overrepresented in transgender women, most involving AR (namely, AR-ERβ, AR-PGR, AR-COMT, CYP17-SRD5A2). Overrepresented alleles and genotypes are proposed to undermasculinize/feminize on the basis of their reported effects in other disease contexts. Conclusion Gender dysphoria may have an oligogenic component, with several genes involved in sex hormone–signaling contributing. The Journal of Clinical Endocrinology & Metabolism, Volume 104, Issue 2, February 2019, Pages 390–396. |
|
|
|
Post by Admin on Sept 10, 2019 18:29:45 GMT
Polymorphic fragments lengths were obtained for 320 to 343 transgender women and 269 to 283 male control subjects. The number of repeats identified for each allele is shown in Supplemental Table 3. Median values were calculated from 24 repeat-length distributions. A difference in median repeat length was identified, with transgender women having a significantly shorter median repeat length (16 TA repeats) for ERα when compared with male control subjects (17 TA repeats; P = 0.03; Table 1).
Table 1.
Comparison of Repeat Length Distributions of AR, ERα, ERβ, CYP11A1, CYP1,9 and PGR Repeat Length Polymorphisms
Gene Male Control Subjects
Transgender Women
No. Median No. Median Mann-Whitney U Test P Value
AR 289 22 327 22 0.31
ERα 567 17 640 16 0.03a
ERβ 580 23 668 23 0.70
CYP11A1 546 4 668 4 0.55
CYP19 580 7 658 7 0.21
PGR 562 18 676 18 0.20
a
Statistically significant at P < 0.05.
Allele frequencies were determined for the five SNPs COMT, CYP17, HSD17B6, SULT2A,1 and STS, and the dichotomous SRD5A2 repeat-length polymorphism (Table 2). A difference between transgender women’s and male control subjects’ allele frequencies was identified in two of the six genes. Specifically, in transgender women compared with male control subjects, there was an overrepresentation of the TA (9 repeats) repeat in SRD5A2 (12.3% compared with 8.5%, respectively; P = 0.030) and the G allele in STS (91.2% compared with 85.9%, respectively; P = 0.015).
Table 2.
Allele Frequencies of the COMT, CYP17, HSD17B6, SRD5A2, STS, and SULT2A1 Single Nucleotide or Dichotomous Polymorphisms
Gene Allele Male Control Subjects
Transgender Women
P Valuea
No. % No. %
COMT G 303 58.7 356 54.1 0.11
A 213 41.3 302 45.9
CYP17 T 354 63.7 384 58.9 0.090
C 202 36.3 268 41.1
HSD17B6 T 366 65.6 406 60.2 0.053
C 192 34.3 268 39.8
SRD5A2 TA(0) 516 91.5 598 87.7 0.030b
TA(9) 48 8.5 84 12.3
STS G 238 85.9 302 92.1 0.015b
A 39 14.1 26 7.9
SULT2A1 T 419 80.9 504 77.5 0.16
C 99 19.1 146 22.5
a
The P value was calculated using the χ2 test.
b
Statistically significant at P < 0.05.
To compare genotypes, repeat lengths were assigned to “short” or “long” allele groups on the basis of the median repeat length of the control population for the six repeat-length polymorphism genes (AR, short ≤22, long >22; CYP11A1, short ≤4, long >4; CYP19, short ≤7, long >7; ERα, short, ≤17, long >17; ERβ, short ≤23, long >23; and PGR, short ≤18, long >18). The genotypes of CYP11A1, CYP19, ERα, ERβ, and PGR polymorphisms for all individuals were determined as SS (i.e., two short alleles), SL (i.e., one short and one long allele), or LL (i.e., two long alleles). Because the AR gene is located on the X chromosome, it is hemizygous and the allele and genotype frequencies are equivalent.
Genotype frequencies of the stratified repeat-length polymorphisms (in CYP11A1, CYP19, ERα, ERβ, PGR, and SRD5A2) and of the SNPs (in COMT, CYP17, HSD17B6, STS, and SULT2A1) were analyzed using binary logistic regression. An association was identified between transgender women and the SULT2A1 homozygous TC genotype (P = 0.009) and the ERα SS genotype (P = 0.035) when compared with male control subjects (Table 3). The ORs indicate that within this population, the likelihood of being transgender increases by 1.61 times (95% CI, 1.13 to 2.31) if an individual possesses the SULT2A1 TC genotype and increases by 1.65 times (95% CI, 1.04 to 2.63) if an individual possesses the ERα SS genotype.
Table 3.
Genotype Analysis of the COMT, CYP11A1, CYP17, CYP19, ERα, ERβ, HSD17B6, PGR, SRD5A2, and SULT2A1 Polymorphisms
Gene Genotype Male Control Subjects
Transgender Women
No. % No. % P Value (OR; 95% CI)a
COMT GG 85 32.8 100 30.4 0.06 (1.59; 0.98–2.58)
GA 135 52.1 156 47.4
AA 39 15.1 73 22.2
CYP11A1 SS 89 33.1 124 37.1 0.24 (0.81; 0.57–1.15)
SL 138 51.3 155 46.4
LL 42 15.6 55 16.5
CYP17 TT 112 40.3 109 33.2 0.097 (1.34; 0.95–1.91)
TT 130 46.8 170 51.8
CC 36 12.9 85 14.9
CYP19 SS 86 30.3 83 25.2 0.20 (1.36; 0.85–2.16)
SL 142 50.2 174 52.9
LL 55 19.4 72 21.9
ERα SS 75 27 103 32.2 0.035b (1.65; 1.04–2.63)
SL 138 49.6 163 50.9
LL 65 23.4 54 16.9
ERβ SS 186 65.7 216 64.7 0.81 (1.04; 0.74–1.45)
SL 85 30 103 30.8
LL 12 4.2 15 4.5
HSD17B6 TT 122 43.7 123 36.5 0.091 (1.53; 0.93–2.51)
TC 122 43.7 160 47.5
CC 35 12.5 54 16
PGR SS 81 29.3 106 31.4 0.35 (0.80; 0.50–1.27)
SL 135 48.9 169 50
LL 60 21.7 63 18.6
SRD5A2 TA(0)/(0) 238 84.4 269 78.4 0.12 (1.42; 0.92–2.18)
TA(0)/(9) 40 14.2 64 18.7
TA (9)/(9) 4 1.4 10 2.9
SULT2A1 TT 175 67.6 192 58.9 0.009b (1.61; 1.13–2.31)
TC 69 26.6 122 37.4
CC 15 5.8 12 3.7
Abbreviations: L, long allele; S, short allele.
Of possible two-locus gene interactions modeled using binary logistic regression, four interactions were overrepresented in transgender women when compared with male control subjects: AR- ERβ, AR-PGR, AR-COMT, and CYP17-SRD5A2 (Table 4). Notably, three of the four interactions involve the long CAG repeats of the AR.
Table 4.
ORs and 95% CIs of Statistically Significant Two-Locus Gene Interactions Determined by Binary Logistic Regression Modeling
Two-Locus Gene Interaction OR 95% CI P Value
Lower Upper
AR (L) – ERβ (SL) 2.78 1.15 6.70 0.023
AR (L) – PGR (LL) 5.68 1.63 17.95 0.006
AR (L) – COMT (AA) 3.88 1.2 12.56 0.024
CYP17 (A1A2) – SRD5A2(TA(0)/TA(9) 4.09 1.19 14.03 0.025
|
|
|
|
Post by Admin on Sept 11, 2019 4:22:17 GMT
Discussion
To our knowledge, this is the largest study to date of gender dysphoria conducted; 12 genes were examined. Variants within COMT, CYP11A1, HSD17B6, STS, and SULT2A1 have not previously been studied in transgender men or women, to our knowledge. In addition, to our knowledge, our study is the only one of three studies to identify an association between TA repeats in ERα and gender dysphoria. Our study did not reproduce the independent associations between ERβ and gender dysphoria in transgender women reported by Henningsson et al. (35) or the association with long CAG repeats in AR, previously identified in a subset of our present cohort (21). However, ERβ and AR were identified as overrepresented in transgender women when in combination with other genes, supporting their involvement in the development of gender dysphoria.
Associations were identified among genetic variants in ERα, SRD5A2, STS, and SULT2A1 and this cohort of transgender women. These genetic variants are suspected to be functional, which permits us to examine the predicted functional effects of the specific polymorphism overrepresented in transgender women. In ERα, for example, short TA repeats overrepresented in transgender women are also associated with low bone mineral density in women (36). Therefore, we speculate that estrogen signaling is reduced (37). In SULT2A1, the heterozygous TC genotype is overrepresented in transgender women. The minor, C allele of SULT2A1 is associated with elevated sex hormone–binding globulin (38), a glycoprotein that regulates circulatory sex steroid bioavailability and is present within fetal male blood during early gestation (39). In transgender women with the TC SNP, we speculate that fetal sex hormone–binding globulin levels are increased, which may reduce the effects of circulating hormones. Polymorphisms in two genes were overrepresented in transgender women by allele analysis but not by the (more stringent) genotype analysis. First, TA(9 repeats) of SRD5A2 is associated with reduced prostate cancer risk likely due to reduced DHT (40), suggesting that levels of the potent androgen DHT could be reduced among transgender women. Second, the G allele in STS is associated with reduced enzyme levels; this has been noted mostly in studies of ADHD (41), a condition with fivefold increased incidence of gender dysphoria (42), suggesting a possible overlap in etiology.
Four important, two-locus interactions were identified by binary logistic regression modeling: AR-ERβ, AR-PGR, AR-COMT, and CYP17-SRD5A2. Of these, three involved long CAG repeats of the AR. Although long CAG repeats in AR alone may not have an independent effect on the development of gender dysphoria, this AR polymorphism may interact with other genes to increase the likelihood of being transgender. This is consistent with a previous finding by Hare et al. (21), who identified an increased proportion of long AR repeat lengths within a subset of the population of transgender women in their study. Long CAG repeats reduce AR signaling (43–45). Similarly, long repeats in ERβ have been associated with decreased ERβ signaling (36), potentially reducing the influence of ERβ on the defeminization of the male brain (46). In combination, both genotypes appear to have additive effects on the development of gender dysphoria. In contrast, the interaction of AR and COMT is unclear where the Met158 homozygous genotype is known to reduce COMT activity (47), affecting estrogen catechol metabolism. Also, the functional effect of the PGR polymorphism is unclear in transgender women. Interaction analysis also identified the specific combination of SRD5A2 and CYP17 polymorphism, the former associated with reduced levels of DHT (48), whereas the latter is known to increase sex steroid precursor production (49). It seems plausible that together, these polymorphisms may increase the production of precursor steroids and testosterone, but not of DHT, the more potent androgen form.
A limitation of the current study is that patients and control subjects were obtained from two sites, one each in Australia and the United States, and therefore likely represent genetically different populations. Another limitation of this study is the use of Caucasian surnames as a selection criterion. Future and more detailed genetic analyses such as genome-wide association studies, would give insight into the ethnicity of the cohort, obviating the need for selection criteria based upon ethnicity.
In summary, the results of our study of transgender women support the hypothesis that gender dysphoria has a polygenic basis, involving interactions among multiple genes and polymorphisms that may alter the sexual differentiation of the brain in utero, contributing to the development of gender dysphoria in transgender women. However, although discordance rates for gender dysphoria suggest that genetics plays a role, it is not the sole determinant of gender identity. Genome-wide association studies, and genome and methylome approaches, especially when coupled with neuroimaging or sex steroid measurements, should be undertaken to better understand how genetic variants contribute to gender dysphoria.
Transgender people continue to be subjected to high rates of gender-based discrimination when seeking medical care, employment, and education (50, 51). Although people’s civil rights should not hinge on science to validate their individuality and lived experience, determining what biological factors contribute to gender dysphoria may influence public opinion and public policies related to the transgender community. More importantly, such knowledge can be used to improve diagnosis and treatment of transgender people (e.g., differentiating which children with gender dysphoria will persist into adulthood, vs which will remit). Therefore, there is a clinical need to investigate further the genetic and biological basis of gender dysphoria.
|
|




