Post by Admin on Dec 12, 2019 7:44:04 GMT
Plague DNA, Epidemiology, and Ecology
One line of evidence that has, despite its novelty, received significant attention in histories of the JP and the First Pandemic is the isolation of Y. pestis DNA from the skeletal remains of late antique plague victims. Successful captures of the pathogen have, for most scholars, resolved a controversial debate about the plague diagnosis of the JP and the First Pandemic (e.g., refs. 21, 54, and 55). They are also employed to uphold the claim that the JP and late antique plague recurrences killed millions.
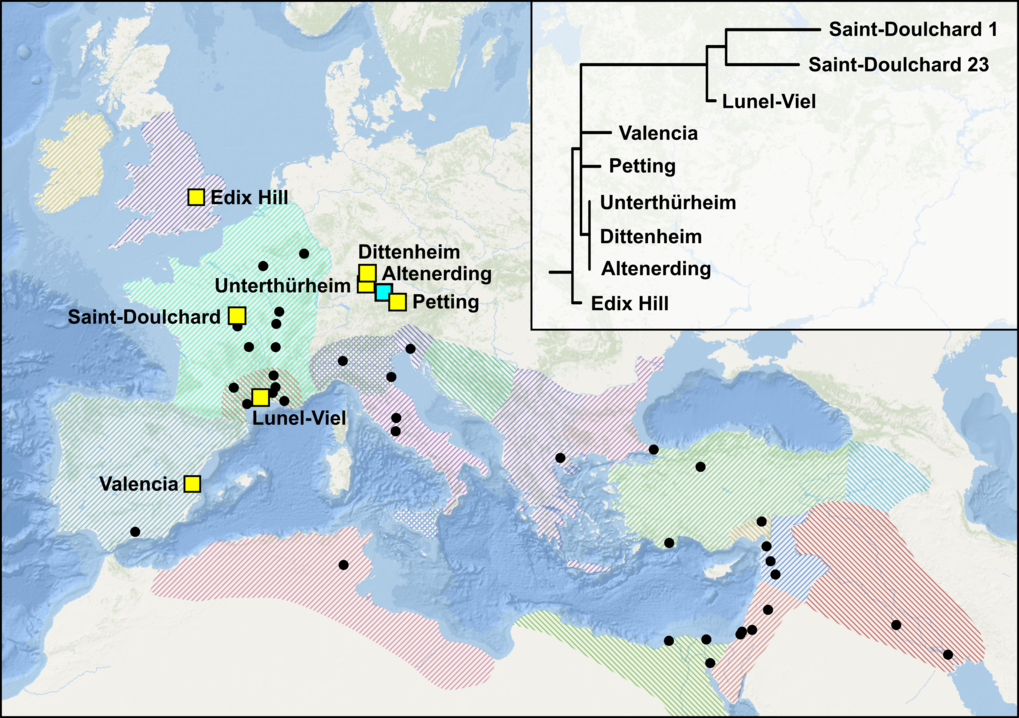
To date, Y. pestis DNA has been isolated from ∼45 individuals in central and western Europe dating to late antiquity (SI Appendix, Table S1). The identification of the plague pathogen in late antique skeletal assemblages is often construed to suggest that the JP and late antique plague recurrences killed a very large number of people (1, 6, 17, 22, 40), but this is merely a conjecture. It cannot be assumed that Y. pestis killed thousands or millions everywhere it is found. In the absence of written or serial evidence for demographic trends, such as that available for the Second Pandemic, the evidence presented above is among the most representative of population trends during the First Pandemic and it fails to support the claim that the JP significantly impacted population levels.
False-negatives are a reality for several reasons (SI Appendix), but few Y. pestis DNA finds correlate spatiotemporally (roughly or with precision) to documented outbreaks of late antique plague, such as Procopius’ account of the JP at Constantinople. Furthermore, few Y. pestis isolations are associated with mass graves (SI Appendix, Table S1). While such burials are not necessarily markers of plague outbreaks, as discussed below, they have received much attention in relation to the JP (1, 2). Roughly 40 mass graves (5 or more simultaneous inhumations) in the Mediterranean region and Europe are known for the 6th and 7th centuries, which surpasses the number known for the 5th and 8th centuries (1, 2), but this is nonetheless a minuscule tally. Several factors account for the failure to isolate Y. pestis DNA consistently or at all from mass graves and areas where large plague outbreaks are reported (SI Appendix), but the current concentration of positive detections in other contexts does not support maximalist accounts.

Some of the early attempts to isolate Y. pestis DNA remain problematic and incomplete (55, 56). For example, PCR-amplified Y. pestis DNA, identified in mass graves from Sens and Vienne (SI Appendix, Table S1), could not have been present in those remains or was mistyped, as it was identified as a branch 1 strain (Orientalis) and, likely evolved less than 250 y ago (15, 16).
The more recent single-nucleotide polymorphism (SNP)-typing and genomic work that has now identified Y. pestis in England, France, Germany, and Spain (4, 16, 17) provides several important insights on First Pandemic plague. It is now clear, for example, that the JP and the Black Death were separate emergences of a very similar yet distinct Y. pestis strain from what are likely to have been wild (but as of yet unidentified) rodent reservoirs, that plague was present in England in the mid-6th century, that Y. pestis diversified and was sustained in Europe during the 6th and 7th centuries, and that late antique plague strains interleave Y. pestis strains isolated from wild rodents and their fleas in Xinjiang, China, and neighboring Kyrgyzstan. Available data also suggest that the JP resulted from a singular emergence with what appears presently to be low genetic diversity, although this is a common characteristic of this clonal pathogen and the available sample size is small.
As valuable as these insights are, they do not lend credence to maximalist interpretations of the JP or the First Pandemic as some suppose. In fact, based on current genetic evidence, it could be argued that plague was less prevalent in the First Pandemic than the second. Y. pestis genomes isolated from late antique remains have no known descendants, meaning either that they have yet to be sampled in an existing plague reservoir or that they went extinct as many Y. pestis strains have over the bacterium’s evolutionary history (4, 16, 17, 57). This stands in current contrast with Second Pandemic strains, the most basal of which sits very close to the base of a polytomy of multiple branches on the Y. pestis tree, which are in turn very closely related (putatively directly ancestral) to Third Pandemic strains and others disseminated across the globe today (58). That currently sequenced Y. pestis strains associated with the JP and the First Pandemic are not directly ancestral to later strains could mean that late antique plague was not established in multiple, long-lived, or highly active reservoirs in any region. This includes western Eurasia and North Africa, where the First Pandemic is known to have spread, central Asia, where it may have emerged and where other plague strains have long persisted, and East Africa, from where some scholars hypothesize the JP reached the Mediterranean (4, 16, 58, 59). Notably, the fewer, shorter lived, and less active the reservoirs, the less human plague there will be.
Furthermore, the recent detection of virulence-linked gene loss in some JP-era Y. pestis genomes, one of which dates to within 80 y of the onset of the First Pandemic, may be important (16). This gene decay may have “facilitated immune evasion,” as previously suggested (16), or it may indicate that the pathogen adjusted to European reservoir species or that it went through attenuation not long after the First Pandemic commenced. This attenuation could have minimized the number of successful transmissions (i.e., virulence) and, therefore, the number of potentially infected individuals. Thus, while the isolation of Y. pestis from 7th and possibly 8th century contexts (SI Appendix, Table S1) indicates late antique Mediterranean or European plague reservoirs maintained the pathogen for decades or longer, and while plague has been isolated from late antique individuals from multiple European regions, available genomic data leave open the possibility that First Pandemic plague was less prevalent than Second Pandemic plague.
Many questions remain unanswered, and more genetic data are certain to emerge, but in the available DNA evidence itself one cannot find confirmation or support of maximalist readings of the JP or First Pandemic as previously claimed. It is also critical to emphasize that our understanding of JP-era Y. pestis is incomplete. Most reconstructed First and Second Pandemic genomes are draft genomes; they have been, as future reconstructed genomes will also be, enriched via the use of baits (small RNA molecules), designed on the basis of existing genomic diversity of Y. pestis from publicly available databases (60). Truly “novel” sequence, in late antique Y. pestis genomes, which might have enabled transmission and thus wide dissemination for instance, will not, under the current methods, be detected, unless enough Y. pestis DNA preserves in an individual from which the genome can be de novo reconstructed.
To understand the scope and probable demographic toll of the JP and the First Pandemic, an idea of how the disease spread within and between settlements would be very valuable. The debate about the mechanics of the Black Death’s dissemination within Mediterranean and European spheres, and beyond, continues, as it does for later Second Pandemic outbreaks (9, 54, 61). With far less evidence for the JP and First Pandemic plague, we are unlikely to solve this problem soon, if at all. Plague is an ecologically complex disease and a number of variables must coalesce for it to spread widely and kill many. The well-documented Third Pandemic demonstrates how, in a more recent context, mortality and prevalence were considerably lower than the Second Pandemic and highly variable on regional and local scales.
That transmission mechanics of Y. pestis during late antiquity remain unknown must further caution us against claiming that the JP or late antique plague in general was nearly omnipresent. Indeed, for over a century scholars have debated how the JP diffused within and between settlements. Current evidence and models implicate sylvatic and commensal rodents, rodent fleas, human ectoparasites (fleas and lice), and respiratory transmission (pneumonic plague). Gastrointestinal plague has hitherto received less attention. Although transmission mechanics greatly determine probable disease incidence, prevalence, and mortality, there is currently no way to discern how the JP or First Pandemic recurrences spread. Therefore, mortality estimates for settlements or regions where the disease was present remain wholly speculative. That potential for Y. pestis dispersal and transmission mechanics would have differed greatly between settlements and regions in late antiquity, as they have in other periods, complicates this matter considerably.
One line of evidence that has, despite its novelty, received significant attention in histories of the JP and the First Pandemic is the isolation of Y. pestis DNA from the skeletal remains of late antique plague victims. Successful captures of the pathogen have, for most scholars, resolved a controversial debate about the plague diagnosis of the JP and the First Pandemic (e.g., refs. 21, 54, and 55). They are also employed to uphold the claim that the JP and late antique plague recurrences killed millions.

To date, Y. pestis DNA has been isolated from ∼45 individuals in central and western Europe dating to late antiquity (SI Appendix, Table S1). The identification of the plague pathogen in late antique skeletal assemblages is often construed to suggest that the JP and late antique plague recurrences killed a very large number of people (1, 6, 17, 22, 40), but this is merely a conjecture. It cannot be assumed that Y. pestis killed thousands or millions everywhere it is found. In the absence of written or serial evidence for demographic trends, such as that available for the Second Pandemic, the evidence presented above is among the most representative of population trends during the First Pandemic and it fails to support the claim that the JP significantly impacted population levels.
False-negatives are a reality for several reasons (SI Appendix), but few Y. pestis DNA finds correlate spatiotemporally (roughly or with precision) to documented outbreaks of late antique plague, such as Procopius’ account of the JP at Constantinople. Furthermore, few Y. pestis isolations are associated with mass graves (SI Appendix, Table S1). While such burials are not necessarily markers of plague outbreaks, as discussed below, they have received much attention in relation to the JP (1, 2). Roughly 40 mass graves (5 or more simultaneous inhumations) in the Mediterranean region and Europe are known for the 6th and 7th centuries, which surpasses the number known for the 5th and 8th centuries (1, 2), but this is nonetheless a minuscule tally. Several factors account for the failure to isolate Y. pestis DNA consistently or at all from mass graves and areas where large plague outbreaks are reported (SI Appendix), but the current concentration of positive detections in other contexts does not support maximalist accounts.
Some of the early attempts to isolate Y. pestis DNA remain problematic and incomplete (55, 56). For example, PCR-amplified Y. pestis DNA, identified in mass graves from Sens and Vienne (SI Appendix, Table S1), could not have been present in those remains or was mistyped, as it was identified as a branch 1 strain (Orientalis) and, likely evolved less than 250 y ago (15, 16).
The more recent single-nucleotide polymorphism (SNP)-typing and genomic work that has now identified Y. pestis in England, France, Germany, and Spain (4, 16, 17) provides several important insights on First Pandemic plague. It is now clear, for example, that the JP and the Black Death were separate emergences of a very similar yet distinct Y. pestis strain from what are likely to have been wild (but as of yet unidentified) rodent reservoirs, that plague was present in England in the mid-6th century, that Y. pestis diversified and was sustained in Europe during the 6th and 7th centuries, and that late antique plague strains interleave Y. pestis strains isolated from wild rodents and their fleas in Xinjiang, China, and neighboring Kyrgyzstan. Available data also suggest that the JP resulted from a singular emergence with what appears presently to be low genetic diversity, although this is a common characteristic of this clonal pathogen and the available sample size is small.
As valuable as these insights are, they do not lend credence to maximalist interpretations of the JP or the First Pandemic as some suppose. In fact, based on current genetic evidence, it could be argued that plague was less prevalent in the First Pandemic than the second. Y. pestis genomes isolated from late antique remains have no known descendants, meaning either that they have yet to be sampled in an existing plague reservoir or that they went extinct as many Y. pestis strains have over the bacterium’s evolutionary history (4, 16, 17, 57). This stands in current contrast with Second Pandemic strains, the most basal of which sits very close to the base of a polytomy of multiple branches on the Y. pestis tree, which are in turn very closely related (putatively directly ancestral) to Third Pandemic strains and others disseminated across the globe today (58). That currently sequenced Y. pestis strains associated with the JP and the First Pandemic are not directly ancestral to later strains could mean that late antique plague was not established in multiple, long-lived, or highly active reservoirs in any region. This includes western Eurasia and North Africa, where the First Pandemic is known to have spread, central Asia, where it may have emerged and where other plague strains have long persisted, and East Africa, from where some scholars hypothesize the JP reached the Mediterranean (4, 16, 58, 59). Notably, the fewer, shorter lived, and less active the reservoirs, the less human plague there will be.
Furthermore, the recent detection of virulence-linked gene loss in some JP-era Y. pestis genomes, one of which dates to within 80 y of the onset of the First Pandemic, may be important (16). This gene decay may have “facilitated immune evasion,” as previously suggested (16), or it may indicate that the pathogen adjusted to European reservoir species or that it went through attenuation not long after the First Pandemic commenced. This attenuation could have minimized the number of successful transmissions (i.e., virulence) and, therefore, the number of potentially infected individuals. Thus, while the isolation of Y. pestis from 7th and possibly 8th century contexts (SI Appendix, Table S1) indicates late antique Mediterranean or European plague reservoirs maintained the pathogen for decades or longer, and while plague has been isolated from late antique individuals from multiple European regions, available genomic data leave open the possibility that First Pandemic plague was less prevalent than Second Pandemic plague.
Many questions remain unanswered, and more genetic data are certain to emerge, but in the available DNA evidence itself one cannot find confirmation or support of maximalist readings of the JP or First Pandemic as previously claimed. It is also critical to emphasize that our understanding of JP-era Y. pestis is incomplete. Most reconstructed First and Second Pandemic genomes are draft genomes; they have been, as future reconstructed genomes will also be, enriched via the use of baits (small RNA molecules), designed on the basis of existing genomic diversity of Y. pestis from publicly available databases (60). Truly “novel” sequence, in late antique Y. pestis genomes, which might have enabled transmission and thus wide dissemination for instance, will not, under the current methods, be detected, unless enough Y. pestis DNA preserves in an individual from which the genome can be de novo reconstructed.
To understand the scope and probable demographic toll of the JP and the First Pandemic, an idea of how the disease spread within and between settlements would be very valuable. The debate about the mechanics of the Black Death’s dissemination within Mediterranean and European spheres, and beyond, continues, as it does for later Second Pandemic outbreaks (9, 54, 61). With far less evidence for the JP and First Pandemic plague, we are unlikely to solve this problem soon, if at all. Plague is an ecologically complex disease and a number of variables must coalesce for it to spread widely and kill many. The well-documented Third Pandemic demonstrates how, in a more recent context, mortality and prevalence were considerably lower than the Second Pandemic and highly variable on regional and local scales.
That transmission mechanics of Y. pestis during late antiquity remain unknown must further caution us against claiming that the JP or late antique plague in general was nearly omnipresent. Indeed, for over a century scholars have debated how the JP diffused within and between settlements. Current evidence and models implicate sylvatic and commensal rodents, rodent fleas, human ectoparasites (fleas and lice), and respiratory transmission (pneumonic plague). Gastrointestinal plague has hitherto received less attention. Although transmission mechanics greatly determine probable disease incidence, prevalence, and mortality, there is currently no way to discern how the JP or First Pandemic recurrences spread. Therefore, mortality estimates for settlements or regions where the disease was present remain wholly speculative. That potential for Y. pestis dispersal and transmission mechanics would have differed greatly between settlements and regions in late antiquity, as they have in other periods, complicates this matter considerably.