|
|
Post by Admin on Feb 5, 2020 20:47:18 GMT
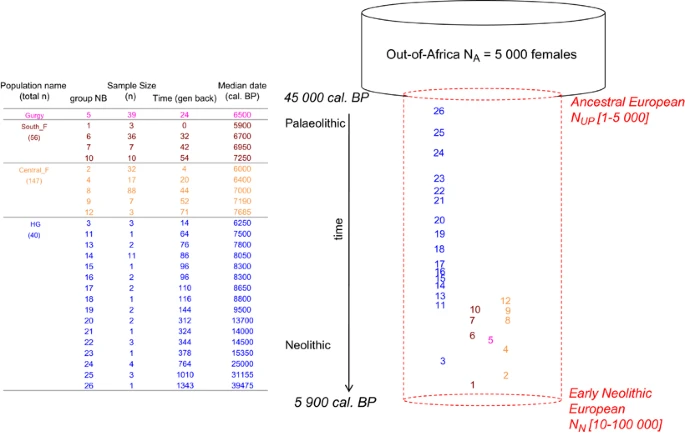 Figure 2 Demographic model simulated with the serial coalescent. Time is the median calibrated C14 years before present (cal BP) backward in time from ‘t0’ and expressed in generations. ‘t0’ refers to 5900 years cal. BP, the median C14 date of the youngest ancient mtDNA sample. Groups are numbered backward in time from the most recent to the most ancient. The dashed cylinder shows constant population size between NUP and NN, but the simulated population can undergo expansion or decline depending on the combinations of these parameter values.  Figure 3 Probability of obtaining simulated FST value greater than that observed for the six pairwise population groups compared (see text for details). Corresponding observed pairwise FST are shown in the top left corner of each grid. The 50 × 50 grids show values of assumed effective population size NN on the x-axis and values of parameterized NUP on the y-axis (note that 25 values are shown on each axis for clarity, see Supplementary Table S2). The top right area delimited by vertical and horizontal black lines outline NN and NUP ranges, respectively, used in comparable studies.2, 11 Gray shows proportions of observed FST greater than observed (proportion >0.05), for which panmixia cannot be rejected. Color-scale represents significance level from blue (proportion lower or equal to 0.05) to red (proportion close to 0). Proportions were obtained over 50 000 simulated pairwise FST per combination of NN and NUP value. A full color version of this figure is available at the European Journal of Human Genetics journal online. Following previous studies,2, 11 we performed serial coalescent simulations under a single panmictic population model with two demographic events: an initial colonization of Europe 45 000 years ago of female effective population size NUP, followed by exponential growth or decline to the Neolithic transition in Western Europe 5900 years cal. BP of female effective population size NN. Before NUP, we assume an ancestral female effective population size NA of 5000, derived from the commonly used long-term effective human population size of 10 000 individuals outside Africa13 and assuming a 1:1 female to male ratio. We explored 50 values for NUP ranging from 1 to 5000 and 50 values for NN ranging from 10 to 100 000 (Supplementary Table S2). We generated 50 000 mitochondrial genealogies of ancient HG and farmer sequences using fastsimcoal version 2.5.1 (ref. 12) under each of the 2500 NUP–NN combinations (Supplementary Table S2). We used a fixed mutation rate of 5 × 10-6/bp/generation,15 assuming a 25 years generation time. These simulated genealogies were used to compute expected pairwise FST values for the six sample comparisons (Figure 2). We recorded the proportion of simulated FST values that were greater than those observed per FST and parameter combination (Figure 3). We also tested if the six observed pairwise FST values as well as eight within sample group statistic values (number of segregating sites and of pairwise differences) could be recovered from simulations under this simple model by performing an approximate Bayesian computation (ABC)-related approach16 (see details in SI). We used the rejection algorithm of the ‘abc’ package17 available in R to retain the parameter combinations that generated simulated pairwise FST the closest to the six observed values. Even though we provide some effective population size estimates, we caution against over-interpretation since there is likely insufficient information in the data to make precise estimates. |
|
|
|
Post by Admin on Feb 6, 2020 20:37:00 GMT
Results and discussion
Analyses indicate that for the six pairwise population group comparisons, some NUP–NN combinations can result in simulated differentiation greater than the one observed (gray area on Figure 3). Notably, results show that we cannot reject the possibility that European HG, South-F, Central-F and Gurgy were sampled from a single panmictic population. Whereas these results may appear to contrast with previous studies that have used serial coalescent simulations to address local mtDNA population continuity between diachronic HG and farmers samples,2, 11 we highlight that our analyses do not address ‘population continuity’ as defined in these studies. The grouping of diachronic samples may artificially reduce the level of differentiation that would be observed in case of significant mtDNA population structure. This grouping none-the-less allows us to investigate the genetic relationships between set of lineage samples associated with specific archeological Neolithic contexts.
We confirmed that our panmictic population model generated simulated between and within population group diversity values close to the observed using an ABC-rejection algorithm (see SI and Supplementary Figure S1). The 95% credible intervals estimated from the retained simulations are (5–3500) NUP females and (200–7750) NN females. These estimates concur with the observation that the parameter space for which a panmictic population model may hold is rather narrow (Figure 3). Most NN values tested and compatible with the level of mtDNA differentiation observed are relatively low (10–200 females for the South-F and Central-F comparison, Figure 3). Noteworthy, some NUP–NN combinations imply a population decline that clearly contrasts with previous studies based on modern DNA data, which have inferred female effective population size growth in Europe during the Holocene.18 However, we were not constrained to simulate population expansion, since we did not consider modern DNA data in our analyses. Moreover, a Holocene population decline in Europe corroborates recent Y chromosome data18 and various archeological evidence support demographic fluctuation of Neolithic populations.19, 20
Our results indicate that a simple panmictic population model can account for the mtDNA differentiation observed between European HG and Early/Middle Neolithic farmers; a larger proportion of the HG–Gurgy explored parameter space failed to reject panmixia. This result suggests increasing HG admixture into farmers’ group migrating farther west in Europe. Similarly, we note that a larger proportion of the explored parameter space fails to reject panmixia when comparing Gurgy and South-F than when comparing Gurgy and Central-F. Thus, our results seem to support Gurgy as the most ancient Neolithic sample studied so far with appreciable admixture between pre-Neolithic HG and Early/Middle Neolithic farmers from both streams of Neolithization in Europe (with a suspected higher participation of Mediterranean farmers).
As with any model, the one we test here has a few assumptions that may not hold, for example, NA of female to male ratio of 1 (ref. 18) and no population structure in any of the four groups.5 Moreover, the panmictic population model proposed would need to be compared against alternatives (eg, ref. 11). Such a simple panmictic population model nevertheless lays the ground for building more complex ones.17 Notably, a serial coalescent approach coupled with ABC would allow estimation of the possible contribution of each of the three population groups (HG, Mediterranean and Central Europe farmers) in shaping Gurgy mitochondrial diversity.
European Journal of Human Genetics volume 25, 388–392(2017)
|
|
|
|
Post by Admin on Feb 7, 2020 19:58:55 GMT
OVER 5,000 YEARS AGO IN what is today Slovakia, a Neolithic community erected a new building. It wasn’t the first “longhouse” in Vráble, an early town comprising about 100 buildings in all. But like those that came before it, the new construction sat a little leeward. So did the one after that. And the one after that. Over time, the entire village slowly turned counterclockwise—and the Stone Age inhabitants of Slovakia likely had no idea it was happening. They weren’t alone. “We find [these longhouses] from the Paris Basin to Ukraine,” says Nils Müller-Scheeßel, an archaeologist at Kiel University and lead author of a recent paper, published last month in the journal PLOS One. “And what we find archaeologically is almost indistinguishably similar. They basically use the same building technique.” These buildings were put up roughly once every 30 or 40 years, and each time the skew was counterclockwise—a pattern that occurred consistently over the course of 300 years.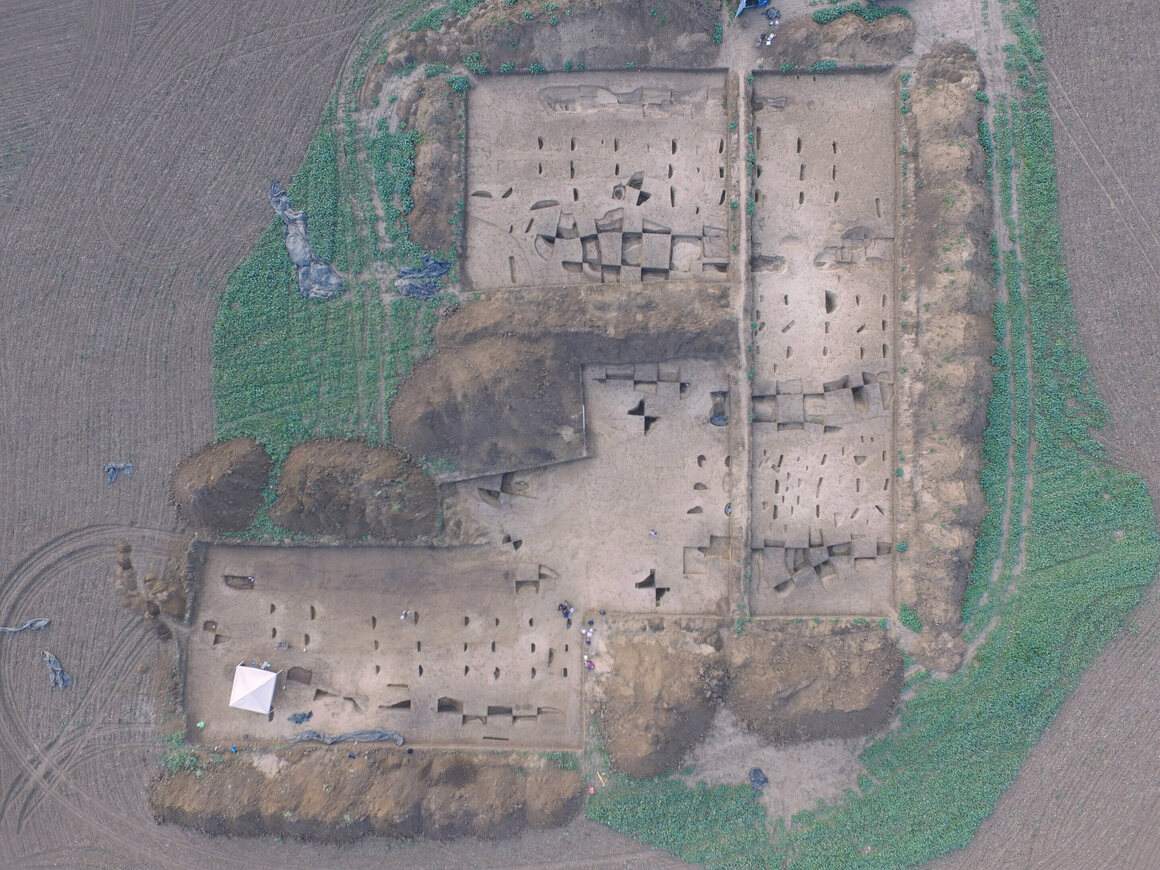  The sides of our brains have different responsibilities: The left hemisphere, for example, is in charge of producing speech, just as the right manages spatial attention. And those splits can sometimes manifest themselves in our interactions with the material world. A portmanteau of “neglect,” pseudoneglect is a natural psychological phenomenon that causes individuals to pay more attention to the left side of their world. It has been observed in species other than humans, and in humans has been shown to affect senses other than sight, such as hearing. But it is most obviously apparent visually, and in the way that individuals will give weight to the left side of their field of view. “It refers to a surplus of attention that is deployed into the left hemispace,” says Mark McCourt, a psychologist at North Dakota State University who is unaffiliated with the recent paper. “The commonly held idea is that it is a byproduct—an epiphenomenon—of the fact that the right hemisphere is specialized for the deployment of spatial attention.”  Twenty years ago, McCourt asked some test subjects to do a number of laboratory tests, all which split a line down the middle. (Some of the tests involved physical things like paper or a rod, others a cursor on a computer screen.) Few subjects split the line just so; most got it a little wrong, loitering on either side of the line’s true center. The majority of subjects erred to the left—a corroboration, McCourt says, of pseudoneglect’s imperceptible lefty bias in our brains. Now, Müller-Scheeßel is arguing that the same process is responsible for the subtle rotation of Slovakia’s Neolithic towns. His hope is that his new research will prove the existence of the phenomenon on a larger, more manifest scale than McCourt’s experiment did. In the Early Neolithic, much of Europe was dominated by the Corded Ware Culture, so named for the population’s proclivity for making and using banded ceramics, which litter the sites of their settlements across the continent. These sites are also known for their longhouses, which have a particularly significant presence in southwestern Slovakia.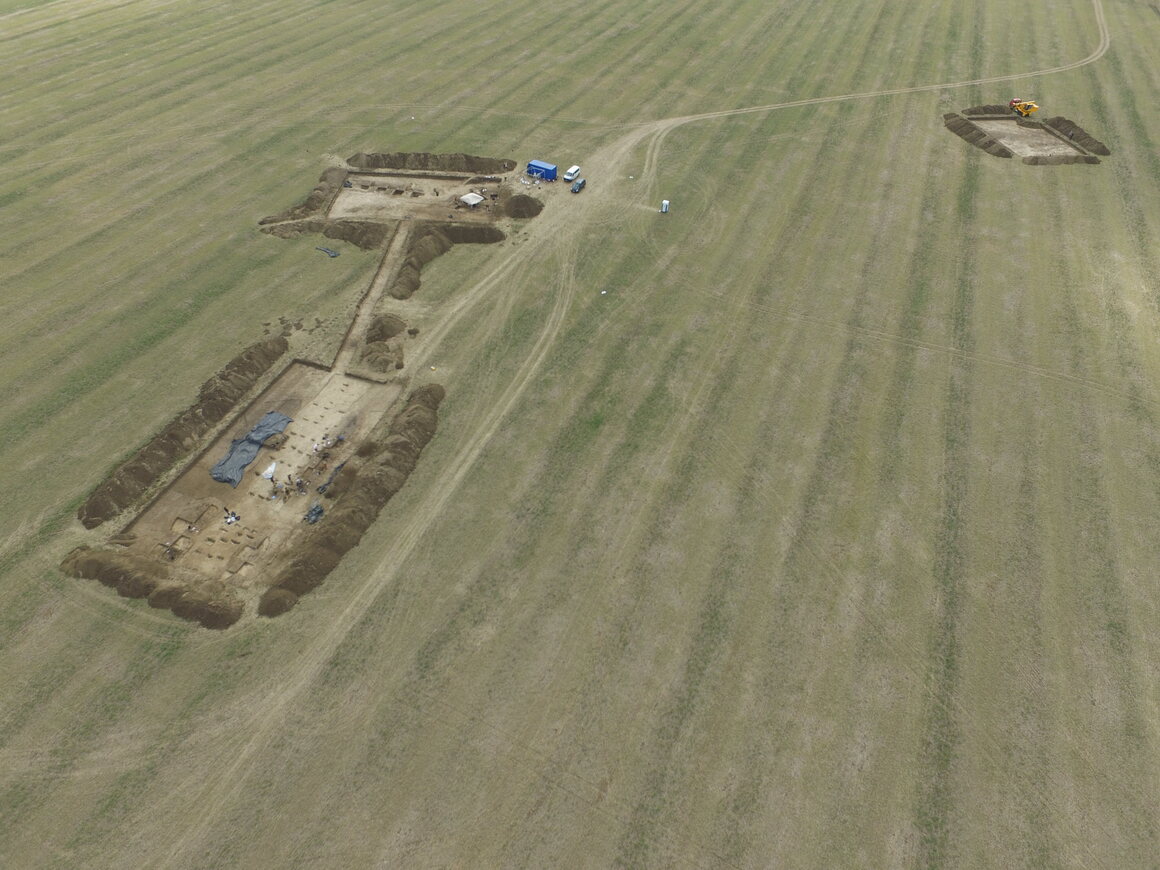  Though the buildings are long gone, Müller-Scheeßel’s team was able to use magnetic surveys to detect the parallel lines of timbers on which the houses were constructed. Despite the size of the settlements and the ample room the various communities had to work with, their urban planning—likely done by mere eyeballing—was pretty consistent. “[They] could [have built] their houses in every direction,” Müller-Scheeßel says. “But there’s a certain tendency for alignment.” Neolithic house rotation is evident in other sites across Europe, and a number of theories have been put forth about why the Corded Ware Culture’s longhouses were oriented the way they were. Some argue that they were built at angles that would best withstand the wind. Others think that celestial bodies like the sun played a role. Still others suggest that longhouses were oriented toward each community’s ancestral homeland. The matter is far from settled, but pseudoneglect could have played a role regardless of the primary reason for how the sites were laid out.  Pseudoneglect could also be connected to the environment of a given population. Recent research in Namibia showed that when rural populations moved to urban settings, they developed a greater left-leaning bias in their perception than they’d previously had. Of course, those are modern populations. These are Neolithic ones. Which makes it a difficult thing to test. Studies of past groups force us to look at what they left behind. Without any ancient brains to study, it’s hard to say whether pseudoneglect operated the same way then as it does today. “We can’t know how Neolithic brains differed from our own,” says McCourt. “It’s puzzling.” There’s an old brainy joke about left-handed people being the only ones in their right minds. The rotating constructions of five millennia ago are a good way to remember that at the end of the day, we all bear left. |
|
|
|
Post by Admin on Mar 11, 2020 20:36:40 GMT
Interactions between earliest Linearbandkeramik farmers and central European hunter gatherers at the dawn of European Neolithization Alexey G. Nikitin, Peter Stadler, Nadezhda Kotova, Maria Teschler-Nicola, T. Douglas Price, Jessica Hoover, Douglas J. Kennett, Iosif Lazaridis, Nadin Rohland, Mark Lipson & David Reich Abstract Archaeogenetic research over the last decade has demonstrated that European Neolithic farmers (ENFs) were descended primarily from Anatolian Neolithic farmers (ANFs). ENFs, including early Neolithic central European Linearbandkeramik (LBK) farming communities, also harbored ancestry from European Mesolithic hunter gatherers (WHGs) to varying extents, reflecting admixture between ENFs and WHGs. However, the timing and other details of this process are still imperfectly understood. In this report, we provide a bioarchaeological analysis of three individuals interred at the Brunn 2 site of the Brunn am Gebirge-Wolfholz archeological complex, one of the oldest LBK sites in central Europe. Two of the individuals had a mixture of WHG-related and ANF-related ancestry, one of them with approximately 50% of each, while the third individual had approximately all ANF-related ancestry. Stable carbon and nitrogen isotope ratios for all three individuals were within the range of variation reflecting diets of other Neolithic agrarian populations. Strontium isotope analysis revealed that the ~50% WHG-ANF individual was non-local to the Brunn 2 area. Overall, our data indicate interbreeding between incoming farmers, whose ancestors ultimately came from western Anatolia, and local HGs, starting within the first few generations of the arrival of the former in central Europe, as well as highlighting the integrative nature and composition of the early LBK communities. Introduction The Linearbandkeramik or Linear Pottery culture (LBK) played a key role in the Neolithization of central Europe. Culturally, economically, and genetically, the LBK had its ultimate roots in western Anatolia, but it also displayed distinct features of autochthonous European Mesolithic hunter-gatherer societies. Several models for the origins of the LBK culture have been proposed over the years1.  Figure 1 The Indigenist model suggests the LBK was founded through the adaptation of elements of the West Asian Neolithic Package by indigenous Mesolithic populations exclusively through frontier contact and cultural diffusion. The Integrationist model views the formation of LBK as the integration of Mesolithic hunter-gatherers into an agro-pastoral lifeway through mechanisms such as leapfrog colonization, frontier mobility and contact. According to this model, small groups associated with the Starčevo-Körös-Criş (SKC) culture, the likely LBK predecessors in Europe, left their homelands in the Balkans (where most of their own ancestors had arrived earlier from Anatolia), and settled new areas to the northwest. Contacts with local Mesolithic groups and exchange of products would have resulted in the co-optation of hunter-gatherers into farming communities, where they would have adopted farming practices1. Evidence of such interactions exists at the Tiszaszőlős-Domaháza site in northeastern Hungary, containing interments of individuals of mostly hunter gatherer genetic ancestry buried in a clearly SKC context2,3. The Migrationist model suggests that a sparsely populated territory of Mesolithic central Europe was taken over by pioneering agro-pastoral groups associated with the SKC culture, which gradually displaced indigenous hunting-gathering populations, who did not significantly influence the arriving Starčevo colonizers. According to this model, newcomers would have replicated their ancestral material culture in the newly settled territory without incorporating the material culture features of the local indigenous populations. Some variation, due to innovation and adaptation to the new environment and sources, would have involved changes in technology such as pottery and building material as well as lithic tool sources. At the same time, symbolic systems, such as decorative designs and cultural objects, would have remained unchanged. This model appeared at the end of the 1950s4 and gained wide support in the second half of 20th century5,6,7,8. To date, ancient DNA (aDNA) studies have convincingly shown that Neolithic European farming populations were primarily genetic descendants of central and western Anatolian Neolithic farmers (ANFs)9,10,11. Their genetic signature is clearly distinct from autochthonous Mesolithic European hunter gatherers (HGs) of central Europe (WHGs) at the level of uniparental markers such as mitochondrial DNA (mtDNA) and Y chromosome as well as genome-wide. Nevertheless, the extent to which the newcomers interacted both culturally and genetically with local hunter gatherers remains unclear; that is, it remains unclear to what extent an Integrationist or Migrationist model is accurate. Genetically, Neolithic central European farmers carried a minor proportion of genetic ancestry characteristic to WHG populations, but the extent and the timing of the WHG admixture in the gene pool of the European Neolithic descendants of Anatolian farmers varies across central Europe3. While the amount of WHG ancestry in European Neolithic farmers had been observed to increase throughout the Neolithic in the present-day territories of Hungary, Germany and other regions of Europe3,9,12,13,14,15, the initial degree of exchange remains unresolved, in part due to a scarcity of human remains contemporaneous with the earliest stages of the Neolithic farming migration. The Brunn 2 archaeological site, part of the Brunn am Gebirge, Wolfholz archaeological complex south of Vienna, Austria (Fig. 1), is the oldest Neolithic site known in Austria and one of the oldest in all of central Europe. It belongs to the earliest stage of the development of LBK, called the Formative phase. Radiocarbon dates obtained for Brunn 2 time the site to about 5670–5350 cal BCE16,17,18,19,20. The main characteristic of the settlements of the Formative phase is the absence of fine pottery and the use of coarse pottery with clear Starčevo features. The leading role of Anatolian migrants in the formation of cultural attributes of the earliest farmers of Europe is evident through the comparative typological analysis of material culture artifacts from the Brunn 2 site20. In addition to rich trove of culture artifacts, Brunn 2 yielded four human burials. The initial radiocarbon dating of the remains confirmed these to be contemporaneous with the earliest phase of the Brunn am Gebirge complex20 and, thus, to represent some of the earliest central European Neolithic farmers. We set out to perform a bioarchaeological analysis of these individuals to examine genetic ancestry as well as diet and mobility at the dawn of the European Neolithization, in an effort to refine the model of the establishment of farming in the Neolithic central Europe. |
|
|
|
Post by Admin on Mar 11, 2020 22:26:23 GMT
Results Genetic analyses We obtained genetic data passing quality control for three out of the four individuals interred at Brunn 2. No usable genetic data was obtained from Individual 4. Individuals 1–3 were males by genetic typing. The mitochondrial lineages of Individuals 1–3 were J1, U5a1, and K1b1a (Table 1), while their Y chromosomal lineages were BT, CT, and G2a2a1a, respectively. For Individual 1, we note that we ostensibly observed derived alleles at the diagnostic haplogroup P sites CTS3446 and F212, the R1 site CTS997, and the R1b1a1a2 sites PF6444 and L749 (nomenclature from the International Society of Genetic Genealogy, , but these were mostly carried on long sequencing reads (41, 96, 74, 131, and 96 bases, respectively), none of which had evidence of ancient DNA damage, so we believe some or all of them to be due to low levels of contamination. We also observed an ancestral allele at the haplogroup R site L1225 (read length 45, likewise not damaged). SNP data obtained from whole genome sequencing were used as the basis for a PCA plot comparing to Neolithic Anatolian and early Neolithic European individuals (Fig. 2). Individual 2 (I6913) fell closest to WHGs but shifted toward EEFs/ANFs, while Individuals 1 (I6912) and 3 (I6914) grouped with Anatolian Neolithic farmers and closely related central European farming groups. In the zoomed-in plot (Supplementary Fig. S1), I6914 appears to be borderline between ANFs and ENFs, while I6912 is on the high end of WHG relatedness for early European farmers. We note though that I6912 and especially I6913 have relatively low sequencing coverage, so their exact positions in PCA should be interpreted with caution. 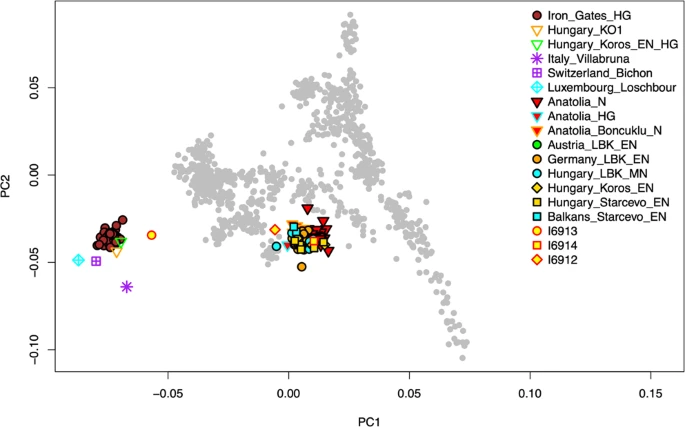 Figure 2 To formalize these observations, we used f-statistics to measure asymmetries in allele sharing (see Materials and Methods). For Individuals 1–3, we estimated genome-wide WHG-related ancestry proportions of 12 ± 3%, 57 ± 8%, and <1%, respectively, consistent with the PCA results (Table 1). For Individual 1, we determined his WHG ancestry to be more closely related to western and central European hunter-gatherers than to southeastern European hunter-gatherers (f4(#1, Anatolia N; W/C WHG, SE WHG) = −2.0*10−3, |Z| = 2.4; cf. ref. 3); for Individual 2, this distinction could not be made with precision (|Z| < 1). Finally, we tested for unequal relatedness of Individual 3 (who yielded the highest sequencing coverage) to various published Neolithic farmers and found that while he is approximately symmetrically related to Neolithic Anatolians and Starčevo-associated individuals from southeastern Europe, he does share excess alleles with other LBK groups from central Europe (|Z| > 3 for Germany, |Z| > 5 for Austria). We note that this signal is not caused by WHG-related ancestry in the LBK groups, as the statistic value for WHG has the opposite sign (f4(#3, Mbuti; WHG, Anatolia N) ≪ 0, Z < −17). Stable isotope (δ13C and δ15N) data Two of the bone collagen samples (Individuals 1 and 2) failed the quality control parameters, with C:N ratios outside the accepted range of 2.9–3.6 (see40), likely indicating collagen depletion (Table 2). Therefore, the data from those two bone samples should be considered unreliable. The δ13C and δ15N values for the bone collagen sample which passed quality control (Individual 3), are consistent with the isotope values observed in the dentine samples (see below). Ideally, a number of wild and domestic animal bone samples from the site would have been analyzed, to form an isotopic baseline to compare and interpret the human samples against27,31. In their absence, carbon and nitrogen isotope analyses of domestic fauna from the approximately contemporary, and environmentally similar LBK site of Vedrovice, in the Czech Republic63, will be utilized. The domestic cattle samples at Vedrovice produced δ13C values of −20.2‰ ± 0.3 and δ15N values of 6.2‰ ± 1, the sheep/goat samples produced values of −19.8‰ ± 0.2 and 5.9‰ ± 0.4 for δ13C and δ15N respectively, and finally the domestic pig samples −20.4‰ ± 0.4 (δ13C) and 8.2‰ ± 0.9 (δ15N). The faunal δ13C and δ15N values from Vedrovice reflect a C3 terrestrial environment, and if we use these values as an imperfect baseline to examine human diet at Brunn, we could tentatively suggest that cattle and sheep/goat (or similar herbivores) formed the mainstay of diet. If we used the δ15N values of pig samples from Vedrovice, we would suggest the values are too enriched to contribute significantly to human diet, as we would anticipate human δ15N values of 11‰ or above. However, as the fauna do not originate from the same site, it is difficult to produce a definite conclusion. Despite the caveats associated with using bulk dentine values, the isotope values for the dentine samples of Individuals 1, 2 and 3 (Table 2) indicate dietary pathways reliant on C3 terrestrial proteins (e.g. C3 plant species and/or the herbivores which feed on them). It is not possible to elucidate whether a nursing signal is present in the isotopic values of the dentine. When the δ13C and δ15N values from the Brunn 2 individuals are plotted against the average δ13C and δ15N values from European LBK sites30, European Inland Neolithic farming communities (EIN) and European Inland Mesolithic HGs (EIM)64, as well as Anatolian Neolithic farmers (AN)9, all three Brunn 2 individuals place within the range of Anatolian and European Neolithic farmers (Fig. 3). The values for Individual 2 from dentin and bone collagen overlap with the lower margin for δ15N values of EIM, while the δ15N values for Individuals 1 and 2 fall outside of the EIM margin and within the EIN margin (Fig. 3). 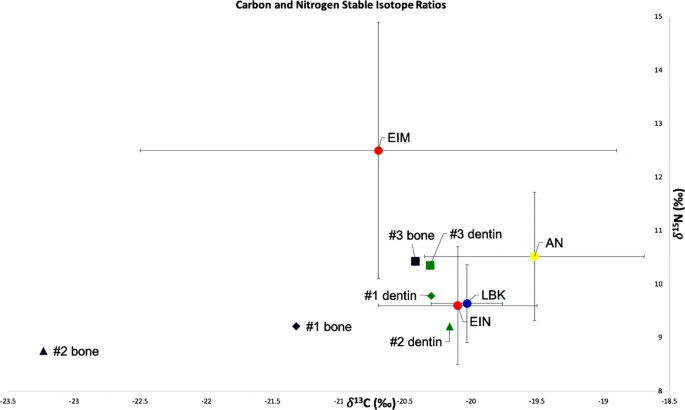 Figure 3 Strontium isotope analysis A small-scale strontium isotope analysis of two of the burials (Individuals 1 and 2) from Brunn 2 was undertaken, limited by the amount of enamel that had survived. Strontium isotope ratios (86Sr/87Sr) in human tooth enamel reflect the nutrients consumed during tissue formation in early childhood, assumed to represent the place of birth. Local or baseline ratios at the place of burial should reflect the place of death. Difference between enamel and baseline should identify individuals of non-local birth. Baseline values were not directly available from the site of Brunn 2 but had been measured at a nearby Longobard cemetery less than 100 m from Brunn 2 (Peter Stadler, personal communication, 2019). Measurement of modern vegetation suggests a local range of 86Sr/87Sr values between 0.7085 and 0.7098. The results of 86Sr/87Sr measurements are presented in Table 2. If we use the Longobard cemetery values in consideration of the two human burials from Brunn 2 then Individual 1 is local and Individual 2 is non-local, falling outside the local range based on the value from the nearby cemetery. |
|









