|
|
Post by Admin on Feb 10, 2020 21:01:49 GMT
 Figure 1 European and African Ancestry-Associated Differences in Immune Response We first aimed to characterize European versus African ancestry-related transcriptional differences in non-infected and infected macrophages. Because self-identified ethnicity is an imprecise proxy for the actual genetic ancestry of an individual, we used the genotype data to estimate genome-wide levels of European and African ancestry in each sample using the program ADMIXTURE (Alexander et al., 2009). Consistent with previous reports (Bryc et al., 2010, Tishkoff et al., 2009), we found that many self-identified AA individuals have a high proportion of European ancestry (mean = 30%, range 0.9%–100%; Figure S1B). In contrast, self-identified EA showed more limited levels of African admixture (mean = 0.4%, range 0%–18%; Figure S1B). Thus, we used these continuous estimates (as opposed to a binary classification of individuals into African or European ancestry) to identify ancestry-associated differentially expressed genes (i.e., pop-DE genes: genes for which gene expression levels are linearly correlated with ancestry levels; see the STAR Methods for details on the nested linear model used for this analysis). Of the 11,914 genes we tested, we identified 3,563 pop-DE genes (30%) in at least one of the experimental conditions, explaining a mean 8.2% of expression variance (range 1.8%–44%) (FDR < 0.05: 1,745 in non-infected :NI, 1,336 in Listeria-infected : L, and 2,417 in Salmonella-infected :s macrophages) (Figures 1B and 1C; Table S2B). These differences primarily influence mean gene expression levels across transcript isoforms, as opposed to the proportion of isoform usage within genes. Specifically, among genes with at least two annotated isoforms (n = 10,223), only 62, 39, and 48 genes exhibited evidence for ancestry-associated differential isoform usage, in the non-infected, Listeria-infected, and Salmonella-infected conditions, respectively (multivariate generalization of the Welch’s t test; FDR < 0.05) (Figures 1D and S2A; Table S2D). These results were unaltered by using an alternative identification approach (Wilcoxon rank sum test, as in Lappalainen et al., 2013; see the STAR Methods for details) or when relaxing the FDR threshold used to define significance (Figure S2B). Despite the low number of genes showing ancestry-associated differences in isoform usage, many of these genes are key regulators of innate immunity, including OAS1 that encodes isoforms with varying enzymatic activity against viral infections (Bonnevie-Nielsen et al., 2005). Next we sought to identify genes for which the response to infection (i.e., fold change in gene expression in infected versus non-infected macrophages, cultured in parallel) significantly correlates with ancestry (see the STAR Methods). We term these genes “population differentially responsive” (pop-DR) genes. We detected 1,005 and 206 pop-DR genes (FDR < 0.05) in response to Salmonella and Listeria, respectively (Figure 1E; Table S2C) (the increased power for Salmonella likely results from the stronger transcriptional response induced by Salmonella relative to Listeria, see Figure 1A). These genes explain a mean 7.4% (range 2.6%–24%) of variance in transcriptional response to infection. Overall, we found that macrophages from individuals of African ancestry produced a markedly stronger transcriptional response to both bacterial infections (Mann-Whitney test, p < 1 × 10−15, Figure 1F). GO term enrichment analyses further revealed that genes related to inflammatory processes were the most enriched among pop-DR genes showing a stronger response to infection in African-descent individuals (Figures 1G and S2C). Together, these results indicate that increased African ancestry predicts a stronger inflammatory response to infection. We hypothesized that ancestry-associated differences in the transcriptional response to infection could translate into ancestry-associated differences in the ability of macrophages to clear the infection. We tested this hypothesis in a subset of 89 individuals by quantifying the number of bacteria remaining inside the macrophages right after the infection step (T0), 2 hr (T2), and 24 hr (T24) post-infection. For both bacteria, increased African ancestry predicted improved control of intracellular bacterial growth. This effect was particularly noticeable in our infection experiments with Listeria. Despite no significant difference in the initial number of bacteria infecting macrophages (Figure 2A, p = 0.95), the number of bacteria inside the macrophages of individuals with high levels of African ancestry at T24 was 3.2-fold lower than that of Europeans (Figure 2A, p = 2.0 × 10−4). Finally, we tested if pop-DE genes were enriched among GWAS-associated genes. We found seven diseases for which susceptibility genes reported by GWAS were significantly enriched among genes classified as pop-DE, in at least one experimental condition (Figure 2B). Contributing to these enrichments are several HLA genes (HLA-DQA1, HLA-DPA1, HLA-DRB1, HLA-DPB1, HLA-DRA), known to be the main genetic risk factors for several immune disorders. Strikingly, six of these seven diseases (all but Parkinson’s disease) are immune-related and tightly connected to a dysregulated inflammatory response. Further, among the diseases most significantly enriched for pop-DE genes were rheumatoid arthritis, systemic sclerosis, and ulcerative colitis, all of which have been reported to differ in incidence or disease severity between AA and EA individuals (Brinkworth and Barreiro, 2014, Pennington et al., 2009). Thus, ancestry-associated gene regulatory differences likely contribute to known ethnic disparities in inflammatory and autoimmune disease susceptibility, in part through affecting the ability of macrophages to control bacterial infections. |
|
|
|
Post by Admin on Feb 11, 2020 21:36:10 GMT
 Figure 2 Increased African Ancestry Predicts Improved Control of Bacterial Growth inside Macrophages To identify whether pop-DE and pop-DR genes are explained by genetic differences between European and African populations, we first mapped genetic variants that are associated with gene expression levels (i.e., eQTL) or transcript isoform usage (alternative splicing QTL [asQTL]) in the complete sample. To do so, we used a linear regression model that accounts for population structure and principal components of the expression data, thus limiting the effect of unknown confounding factors (see the STAR Methods for details). Given that our sample size is too small to robustly detect trans-acting eQTL, we focused our analyses on local associations that, for simplicity, we refer to as cis-eQTL. We define cis-eQTL and cis-asQTL here as SNPs located in the gene body or in the 100 kb flanking the gene of interest. We identified cis-eQTL for 1,647 genes (14% of all genes tested; FDR < 0.01) in at least one of the experimental conditions (875 in non-infected macrophages, 1,087 in the Listeria-infected condition, and 983 in the Salmonella-infected condition; Figure 3A; Table S4A; Figure S3A for number of eQTL found at more relaxed cutoffs). Similarly, we detected a large number of cis-asQTL affecting the ratio of alternative isoforms used for the same gene (1,120 genes, 10% of all genes tested; FDR < 0.01 [Figure 3A; Table S4C]: 886 in non-infected macrophages, 746 in Listeria-infected samples, and 615 in Salmonella-infected samples). Out of all genes with cis-eQTL, a large fraction (21.8%) were associated with an eQTL only in infected macrophages. In contrast, only 7.3% of genes showed an infection-specific cis-asQTL (Figures 3A and 3B). Infection-specific cis-eQTL were further supported by analysis of allele-specific expression (ASE) levels, which provides independent but complementary evidence for functional cis-regulatory variation. As expected, genes with cis-eQTL were significantly enriched for genes with ASE, compared to the background of all 9,588 genes tested (Figure S3B, Fisher’s exact test, p < 1 × 10−15 for all conditions). Further, genes harboring infection-specific eQTL also tended to exhibit infection-specific ASE in the same condition (Listeria or Salmonella) in which the eQTL was identified (Figure 3C, ∼27 fold-enrichment of infection-specific ASE among infection-specific eQTL, relative to shared eQTL; p < 1 × 10−15). Thus, in agreement with previous studies (Fairfax et al., 2014, Lee et al., 2014), genotype-environment (G × E) interactions are common in the context of immune responses to infection, albeit more so for mean expression levels than alternative isoform usage. A complementary approach to identifying G × E interactions for expression levels is to directly map response eQTL (reQTL): QTL associated with the magnitude of change in expression levels after infection (Barreiro et al., 2012, Çalışkan et al., 2015, Lee et al., 2014). In contrast to condition-specific eQTL (an extreme case of G × E interaction), reQTL can capture more subtle interaction effects: eQTL can be shared across conditions as long as their effect size differs between infected and non-infected samples. We detected 244 and 503 genes with a cis-reQTL (FDR < 0.01, Table S4B) for the response to Listeria and Salmonella, respectively. Interestingly, among genes associated with a cis-reQTL, we found several key regulators of the immune response, including the transcription factors STAT4 and IRF2 (Figure 3D). We also found cis-reQTL for known susceptibility loci for ulcerative colitis (e.g., HLA-A, HLA-DQA2, PMPCA), systemic lupus erythematosus (ITGAX, HLA-DQA1), and the infectious diseases hepatitis B and leprosy (e.g., HLA-C, NOD2). To investigate the regulatory mechanisms that account for immune reQTL, we next profiled the genome-wide chromatin accessibility landscape of non-infected and Listeria and Salmonella-infected cells using assay for transposase-accessible chromatin using sequencing (ATAC-seq) (Buenrostro et al., 2013). This approach allowed us to identify transcription factor (TF) binding motifs likely to be occupied by their respective TFs, in both conditions (see the STAR Methods). We found that SNPs within accessible TF binding sites were greater than four times more likely to be identified as reQTL (Figure 3E). Further, reQTL in our analyses were strongly enriched (>20-fold) for PU.1 binding sites (a pioneer TF involved in regulating enhancer activity in macrophages) (Garber et al., 2012) and for virtually all TFs that orchestrate innate immune responses to infection (Figure 3E) (e.g., nuclear factor κB [NF-κB]: >50-fold; AP1: >55-fold; and IRFs: 14-fold for Salmonella only). In striking contrast, we found no such enrichment for eQTL identified in non-infected macrophages (p > 0.05 for NF-κB, AP1, and IRFs) (Figure S3D). These results show that reQTL variants are often conditionally silent in resting macrophages but become functionally relevant post-infection, and this transition is explained by disruption of binding sites for immune response-activated TFs.  Figure 3 eQTL and ASE Analyses Reveal Extensive cis-Regulation of Gene Expression Responses to Pathogens in Macrophages We hypothesized that differences in allele frequencies for some of the eQTL identified above could explain the observed ancestry-associated differences in the transcriptional response to infection. In support of this hypothesis, we found that pop-DE genes were enriched up to 3.3-fold for genes with cis-eQTL (p < 1 × 10−10), and pop-DR genes were enriched up to 5.8-fold for genes with cis-reQTL (p < 1 × 10−10) (Figures 4A and S4A). Additionally, ∼60% of genes that exhibited ancestry-associated isoform usage were associated with an asQTL (up to 24-fold enrichment, p < 1 × 10−10). Thus, although rare, ancestry-associated changes in isoform usage are largely genetically driven. To explicitly quantify the contribution of our eQTL set to transcriptional differences detected between populations, we devised a new score based on PST estimates (Leinonen et al., 2013, Pujol et al., 2008). PST is the phenotypic analog of the population genetic parameter FST, providing a measure of the proportion of overall gene expression variance explained by between-population phenotypic divergence (as opposed to within-population diversity). PST values range from 0 to 1, with values close to 1 implying that the majority of a gene’s expression variance is due to differences between populations. Our score, deltaPST (ΔPST), is defined as the difference between PST values before and after regressing out the effect of the cis-SNP that was most strongly associated with the target gene’s expression level (regardless of significance level), divided by the PST value observed before removing the genotype effect. ΔPST therefore quantifies the proportion of ancestry-associated expression level differences that stem from the strongest cis-associated variant. Among all pop-DE genes, we found that cis-regulatory variants explained an average of 31%, 31%, and 26% of ancestry-related differences in expression observed in non-infected, Listeria-infected and Salmonella-infected samples, respectively (Figure 4B). Further, the larger the effect of ancestry in the original pop-DE analysis, the larger the contribution of cis-regulatory variation to these differences: for pop-DE genes identified at a stringent FDR of 1 × 10−4, cis-regulatory variation explained close to 50% (on average) of ancestry effects (Figure 4B). We observed a similar pattern for pop-DR genes after regressing out the genotype effect of the lead cis-reQTL SNP (Figure 4B). In support of the substantial role of cis-regulatory variation in explaining pop-DE and pop-DR genes, gene expression values for 30% and 45% of pop-DE and pop-DR genes, respectively, were no longer significantly associated with ancestry once we regressed out cis-genetic effects (Figure 4C). Importantly, ΔPST values never exceeded 5% when we regressed out either (1) the genotype effect of randomly selected SNPs matched for the allele frequency of the lead cis-SNP, or (2) lead cis-SNPs identified after permuting the genotype data. Thus, our results cannot be simply explained by population structure (Figure 4B). Based on their known importance in the genetic control of gene regulation and because of power limitations, our main analysis of ancestry-associated gene expression patterns focused on the role of cis-eQTL. However, in a separate analysis, we recalculated ΔPST using the lead trans-SNP for each gene in place of the lead cis-SNP (although only 51, 21, and 22 trans-eQTL genes survived genome-wide multiple testing correction (FDR < 0.1) in non-infected, Listeria-infected and Salmonella-infected samples, respectively). Intriguingly, we found that lead trans-SNPs accounted for an average of ∼23% and ∼20% of ancestry effects on gene expression levels for pop-DE and pop-DR genes, respectively (Figure 4B; at least 2-fold more than estimates based on permuted data, p < 1 × 10−10). These results suggest that lead trans-SNPs, although difficult to detect at a genome-wide significance level, are enriched for true trans-associations that could be resolved with larger sample sizes. Together, a single cis- or trans-acting variant was sufficient to explain almost all ancestry effects (ΔPST > 75%) on gene expression levels for 804 pop-DE genes and pop-DR genes (Figure 4D), including for master regulators of the immune response such as CASP1, STAT4, and MICA. Our results thus provide a comprehensive genome-wide map of cis- and trans-genetic variants associated with African and European ancestry-related differences in the immune response to infection. |
|
|
|
Post by Admin on Feb 12, 2020 20:51:01 GMT
 Figure 4 Contribution of cis and trans Genetic Variation to pop-DE and pop-DR Genes Finally, we sought to determine the impact of recent local positive selection in either African or European populations on ancestry-related divergence in gene expression levels. To do so, we first calculated FST values between the Yoruba African (YRI) and the western European population (CEU) in Phase 3 data from the 1000 Genomes Project (Auton et al., 2015). To generate gene-specific estimates, we averaged FST values for variants within a window of 10 kb around the transcription start site (TSS) of each gene we analyzed (11,914 genes). As a complementary approach, we also calculated integrated haplotype scores (iHS) for all SNPs with a minor allele frequency (MAF) >5% in the CEU and YRI samples. In contrast to FST, iHS is a within-population measure of recent positive selection that is not affected by the levels of population differentiation (Voight et al., 2006). Our analyses identified significantly higher mean FST values among genes that were pop-DE, pop-DR, or showing differences in isoform usage between populations (p ≤ 1 × 10−3; Figures 5A and S5A for similar results when using alternative window sizes). Further, variants identified as cis-eQTL were significantly enriched (∼2-fold) for high iHS values (i.e., iHS > 99th percentile of genome-wide distribution, Figure 5B, p < 1 × 10−8), consistent with the importance of regulatory genetic variation in recent human evolution (Fraser, 2013). cis-reQTL and cis-asQTL were even more strongly enriched among high iHS values (up to 3.6-fold; Figure 5B, p < 1 × 10−5). Overall, within the set of cis-eQTL-, cis-reQTL-, or cis-asQTL-associated genes, 258 carried a signature of recent positive selection in either CEU or YRI samples (|iHS| ≥ 99th percentile of the genome-wide distribution) (Figure 5C; Table S5A). These variants were also significantly enriched for high XP-EHH values (Sabeti et al., 2007) (∼6-fold, p < 1 × 10−10, Figure S5C), further supporting that these variants have been important in recent, population-specific human adaptation. However, because outlier methods for detecting selection can be susceptible to false positives (Kelley et al., 2006), we complemented our iHS analysis with a model-based approach. Specifically, we compared the observed iHS value for each putatively selected allele to those observed under neutral coalescent simulations matched to the known demographic history of African and European populations (Gutenkunst et al., 2009), the candidate allele’s observed frequency, and the local recombination rate. The vast majority (92%) of all sites tested exhibited significantly larger observed iHS statistics than expected under a neutral model (p < 0.01, Table S5B), providing strong convergent support for recent positive selection at these loci. Far more of these genes are pop-DE or pop-DR than expected by chance (47% and 23%, respectively: Figure 5D, p < 0.001), showing that natural selection has contributed to present-day inter-population differences in innate immune responses to infection. Neanderthal ancestry makes up ∼2% of the ancestry of living humans found outside of Africa (Kelso and Prüfer, 2014). It is therefore plausible that interbreeding between Neanderthal and modern human populations could also contribute to some of the ancestry-related differences in gene expression we observed, especially if it enabled the ancestors of modern Europeans to more rapidly adapt to a new pathogen environment (Ségurel and Quintana-Murci, 2014). To test this hypothesis, we identified sites where the derived allele is shared between Neanderthals and non-African populations, but is absent in sub-Saharan Africans samples considered. This class of sites, which we call “Neanderthal-like sites” (NLS), is a conservative indicator of Neanderthal introgression (Sankararaman et al., 2014). Among the 18,862 NLS tested in our cis-QTL analyses, 297 were significantly associated with transcriptional variation of 145 genes (NLS-QTL). Among these 145 genes, 46% (FDR < 0.05) were differentially regulated in at least one experimental condition (non-infected, Listeria-infected, Salmonella-infected, or in the response to either type of infection) between Europeans and Africans (63% at a more relaxed FDR < 0.1). Thus, a non-negligible proportion of ancestry-related gene expression divergence probably results from introgression of functional Neanderthal variants into the ancestors of modern Europeans. Interestingly, some of these variants (n = 16) also have elevated iHS values (|iHS| ≥ 2) (Figure 5C; Table S5A) and therefore represent new candidates for adaptive introgression in humans. |
|
|
|
Post by Admin on Feb 13, 2020 21:14:13 GMT
 Figure 5 Natural Selection on eQTL and Its Contribution to Ancestry-Associated Regulatory Differences Together, our results provide a comprehensive characterization of genes for which the transcriptional responses of primary cells to live pathogenic bacteria differs depending on European versus African ancestry. We show that 34% of genes expressed in macrophages show at least one type of ancestry-related transcriptional divergence, whether in the form of differences in gene expression (30%), the transcriptional response to infection (9.3%), or, less commonly, differences in isoform usage (1%). Notably, the modest contribution of differences in isoform usage to ancestry-related expression levels differs from previous results in lymphoblastoid cell lines (LCLs), where they were found to be quite common (Lappalainen et al., 2013). The discrepancy between our results and those reported for LCLs may be related to differences in the experimental procedures used to produce the two sets of LCL lines, which were generated more than 20 years apart (Dausset et al., 1990). One of the most striking observations from our study was the markedly stronger response to infection induced in macrophages from individuals of African descent, particularly among inflammatory response genes. This result agrees with previous reports showing that AAs have higher frequencies of alleles associated with an increased pro-inflammatory response (Ness et al., 2004), increased levels of circulating C-reactive protein (Kelley-Hedgepeth et al., 2008), and a much higher rate of inflammatory diseases than EA individuals (Pennington et al., 2009). Although the exact causal link between ancestry and the pro-inflammatory response has yet to be established, we speculate that the stronger inflammatory response associated with African ancestry accounts for the increased ability of macrophages in African ancestry individuals to control bacterial growth post-infection. Nevertheless, the evolutionary pressures that explain these differences remain an open question. One possibility is that, after human populations migrated out of Africa, they were exposed to lower pathogen levels (Guernier et al., 2004), which reduced the need for strong, costly pro-inflammatory signals. Change in this direction may have been favored due to the detrimental consequences of acute or chronic inflammation, which are key contributors to the development of autoinflammatory and autoimmune diseases (Okin and Medzhitov, 2012). This hypothesis is consistent with previous reports showing a signature of positive selection in Europeans on a high-frequency non-synonymous variant in the Toll-like receptor 1 gene, which is also associated with impaired NF-κB-mediated signaling (Barreiro et al., 2009). Alternatively, the weaker inflammatory response detected in Europeans could have resulted from relaxation of selective constraint in an environment where the pathogen burden was reduced, or at least different in nature, from that found in Africa.  Figure S1 Study Design and Evaluation of Technical Confounders, Related to STAR Methods Because our samples were derived from individuals with their own unknown life histories and environmental exposures, the ancestry-related differences we observed could be explained by both environmental and genetic factors. However, our eQTL analyses suggest that genetic contributions are probably substantial. We estimate that, on average, ∼30% and 20% of ancestry-associated expression differences in pop-DE genes are accounted for by cis- and trans-regulatory variants, respectively. Further, among the genes with the most robust association with genetic ancestry (pop-DE genes with FDR < 1 × 10−4), putatively cis-acting variants explain up to ∼50% of ancestry effects. Notably, these numbers probably underestimate the true genetic contribution to ancestry-related differences in gene expression, given our low power to detect trans associations, our exclusion of non-SNP regulatory variants, which may also influence gene expression (Gymrek et al., 2016), our conservative assumption that genes have only one major cis-eQTL (many genes have at least two independent cis-eQTL) (Lappalainen et al., 2013); and the fact that we limited our cis-eQTL mapping to a 100-kb window around the targeted gene. The extent to which positive selection has contributed to recent human evolution remains a matter of intense debate (Enard et al., 2014, Fagny et al., 2014, Hernandez et al., 2011). Here, we show that variants associated with regulatory QTL are strongly enriched for signatures of recent selection, supporting an important role of adaptive regulatory variation in recent human evolution. More specifically, our results suggest that a significant fraction of population differences in transcriptional responses to infection are a direct consequence of local adaptation driven by regulatory variants. Notably, several positively selected regulatory QTL (or SNPs in strong LD with them [r2 > 0.8]) have been associated with common diseases by GWAS, further reinforcing the link between past selection and present-day susceptibility to disease (Barreiro and Quintana-Murci, 2010, Brinkworth and Barreiro, 2014). Some examples include positively selected variants affecting the expression of HLA-DQA1, the major genetic susceptibility factor for celiac disease (Abadie et al., 2011), ERAP2, a susceptibility factors for Crohn’s disease (Jostins et al., 2012), and the transcription factor IRF5, which is associated with systemic lupus erythematosus, rheumatoid arthritis, ulcerative colitis, and systemic sclerosis (reviewed in Eames et al., 2016). Finally, our results provide new insight into the contribution of adaptive introgression from admixture with Neanderthals to the diversification of the immune system among modern human populations. We found 17 positively selected NLS regulatory-QTL (associated with 16 genes) that are candidates for adaptive introgression in humans. These genes include previously identified candidates such as TLR1 (Dannemann et al., 2016, Deschamps et al., 2016) but also a large set of loci that have not previously been associated with adaptive introgression. For example, one of the strongest signatures of selection was found for eQTL for DARS, a gene associated with neuroinflammatory and white matter disorders (Wolf et al., 2015). However, in agreement with evidence that most introgressed variation from Neanderthals was probably deleterious (Sankararaman et al., 2014, Vernot and Akey, 2014), as putative cases of adaptive introgression remain relatively rare. DOI: doi.org/10.1016/j.cell.2016.09.025 | |
|
|
|
Post by Admin on Feb 21, 2020 1:12:56 GMT
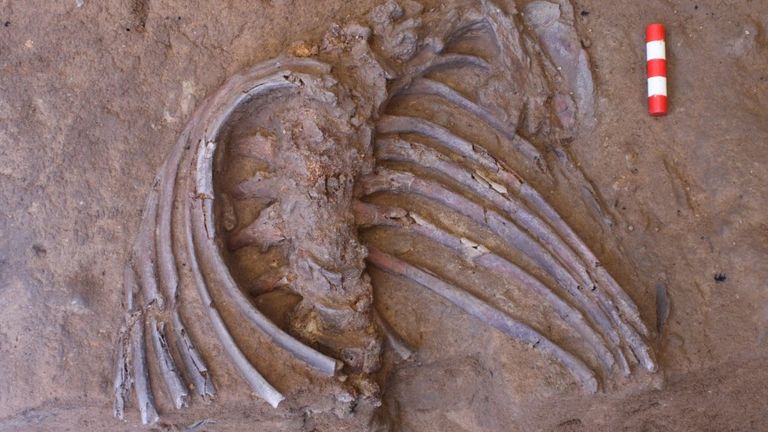 Researchers have described the first "articulated" remains of a Neanderthal to be discovered in a decade. An articulated skeleton is one where the bones are still arranged in their original positions. The new specimen was uncovered at Shanidar Cave in Iraq and consists of the upper torso and crushed skull of a middle-aged to older adult. Excavations at Shanidar in the 1950s and 60s unearthed partial remains of 10 Neanderthal men, women and children. During these earlier excavations, archaeologists found that some of the burials were clustered together, with clumps of pollen surrounding one of the skeletons.  The researcher who led those original investigations, Ralph Solecki from Columbia University in New York, claimed it was evidence that Neanderthals had buried their dead with flowers. This "flower burial" captured the imagination of the public and kicked off a decades-long controversy. The floral interpretation suggested our evolutionary relatives were capable of cultural sophistication, challenging the view - prevalent at the time - that Neanderthals were unintelligent and animalistic. Before the most recent specimen uncovered in Iraq, the last articulated Neanderthal remains were unearthed at Sima de las Palomas in 2006-7 and at Cova Forada in 2010. Both sites are located in south-east Spain. But Dr Emma Pomeroy, from the University of Cambridge, said the new skeleton - dubbed Shanidar Z - is more substantial and more completely articulated than those previous finds.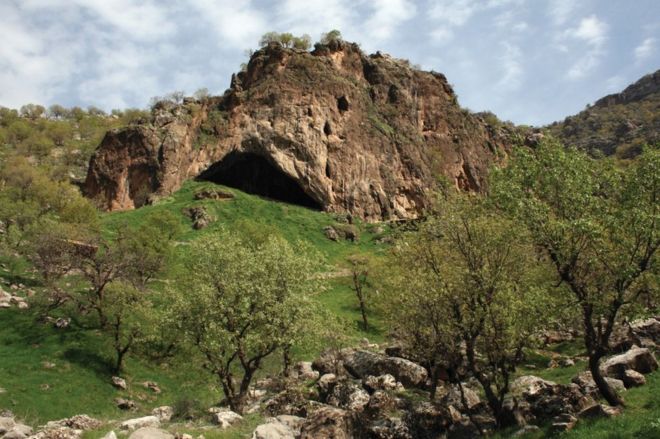  |
|









