|
|
Post by Admin on Sept 2, 2020 1:41:07 GMT
The story in Africa remains murky, however, as researchers have not been able to reconstruct human history in vivid detail, in part because hominin fossils informative about our species’ emergence and coexistence with other species are rare in Africa. As a result, finds such as the Kabwe skull continue to raise more questions than answers. If Homo heidelbergensis wasn’t one of our recent ancestors, then who was? If our species really did overlap in time with Homo heidelbergensis, what role did they play in our evolutionary history? In recent years, a field that has traditionally relied on fossil discoveries has acquired helpful new tools: genomics and ancient DNA techniques. Armed with this combination of approaches, researchers have begun to excavate our species’ early evolution, hinting at a far more complex past than was previously appreciated—one rich in diversity, migration, and possibly even interbreeding with other hominin species in Africa. “To piece together that story, we need information from multiple different fields of study,” remarks Eleanor Scerri, an archaeologist at the Max Planck Institute for the Science of Human History in Jena, Germany. “No single one is really going to have all the answers—not genetics, not archaeology, not the fossils, because all of these areas have challenges and limitations.”  A sparse fossil record Bones easily disintegrate in many parts of Africa, in acidic forest soils or dry, sun-exposed areas. Moreover, the continent is largely unexplored by archaeologists. While northwestern Africa and former British territories in eastern and southern Africa have a long tradition of professional archaeological research, few researchers have looked for fossils anywhere else, notes archaeologist Khady Niang of Cheikh Anta Diop University in Senegal. That’s especially the case for the western and central parts of the continent, where preservation conditions are also poor and excavations difficult at times due to political instability. “We might be missing some really, really important parts of the story,” adds Yale University anthropologist Jessica Thompson. What African hominin fossils do make clear is the depth of humanity’s roots on that continent. Researchers have found some of the most abundant fossils in sediments between 3.5 million and 3.2 million years old. That appeared to be the heyday of the australopiths (including the genus Australopithecus), apes that walked upright and are believed to have used stone tools, but still climbed trees and had relatively small brains. It’s thought that somehow our own genus, Homo, emerged from transitional ape species some 2.8 million years ago as a clan of hominins with distinctive teeth, probably adapted to an eclectic diet that allowed them to thrive in a wide range of habitats. But there are few sediments, let alone fossils, left behind from that time, making the birth of our genus one of the most poorly understood periods in our evolution, Thompson notes. The fossil record yields more secrets about the time shortly after the emergence of Homo, revealing a diversity of different Homo species in Africa, of which Homo erectus seems to persist the longest. Homo erectus crops up in Africa’s limited fossil record around 2 million years ago and hangs around on the continent until roughly a million years ago. It was the first hominin that shows evidence of having lived in human-like social groupings and used fire, and it is thought to be a human ancestor. When and how Homo sapiens emerged isn’t at all clear, but what is apparent is that we weren’t alone; fossils suggest that several other hominin species, such as that represented by the Kabwe skull, inhabited the continent at the time our species appeared. Another relatively small-brained hominin, Homo naledi, is also thought to have lived in southern Africa around 300,000 years ago. And inside a Moroccan cave called Jebel Irhoud, 300,000-year-old skeletons were found that carry very early features of Homo sapiens. It’s not yet known how long those different hominin species existed, however, or whether they physically overlapped and perhaps even shared genes with one another, Stringer notes, or whether there were others. By around 160,000 years ago, the constellation of physical features that defines us today—such as a globular braincase and a pointed chin—had begun to emerge in ancient hominin groups represented by fossils found across Africa. Later, some of these anatomically modern humans crossed the thin spit of land that connects Africa to Eurasia, probably on several occasions. On that new continent, they eventually met Neanderthals and Denisovans, which, like two hobbit-size Homo species found on southeast Asian islands, are thought to be the evolutionary products of earlier hominin migrations out of the continent. “Africa was this sort of leaky faucet, and hominins were just dribbling out of it all the time,” Thompson says.  Fossil finds over the years have steadily bolstered a long-held idea that anatomically modern humans first emerged in Africa. This “Out of Africa” model, proposed by anthropologists in the late 20th century, posited that all humans of Eurasian ancestry descended from a single ancestral African population, which then spread throughout the world and displaced all other hominins. The opposing “multi-regionalism” model, by contrast, conceived that multiple human subpopulations—which stemmed from regional lineages of an ancestral species such as Homo erectus—existed across Europe, Asia, and Africa, and through continuous mixing evolved together to form the present human population. While fossils supported the former theory, it was the advent of genetic research that showed unequivocally that populations outside of Africa descended from a single population in Africa. But the story had a twist: in two groundbreaking studies published in 2014, researchers compared ancient DNA extracted from Neanderthal bones and compared it with modern-day people, and found that 2 percent of the average European genome is Neanderthal in origin. Our species originated in Africa, but interbred with hominins outside of it. These findings, and many since, have highlighted the power of genetics in resolving questions about human ancestry that fossils alone cannot. Investigations of the genomes of living Africans are now underway to help fill in the gaps of Africa’s fossil record. “[Such studies] are really providing important insights into our population history and African origins,” says Yale University evolutionary biologist Serena Tucci. “We are getting to know and understand processes that happened very early on in our evolutionary history.” |
|
|
|
Post by Admin on Sept 3, 2020 4:27:05 GMT
Our History in Africa Hominin fossils that reveal clues to the emergence of Homo sapiens are rare in Africa, but in combination with studies of modern human genomes, researchers are piecing together an ever more complex timeline of human history. 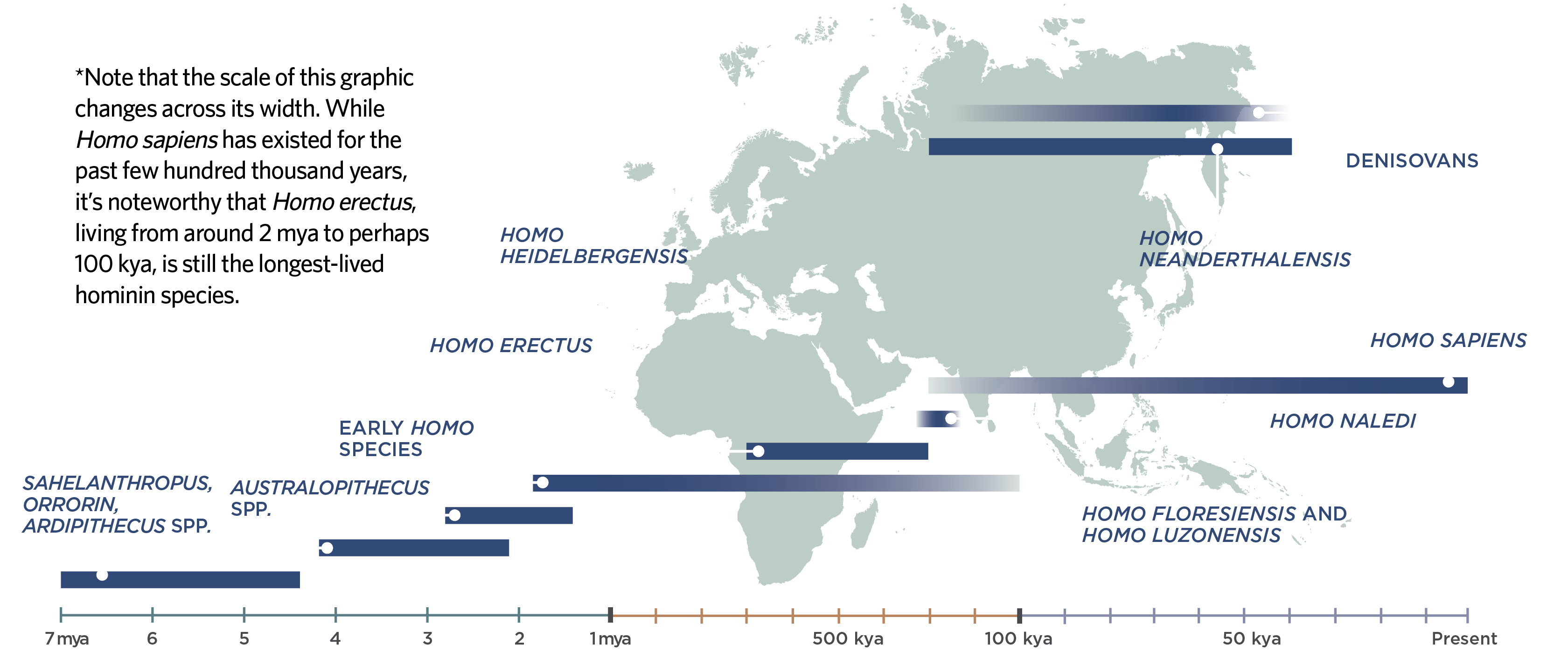 Ghost hominins Even the very first investigations of our genetic ancestry, gleaned from small, bite-size chunks of genetic material, positioned Africa as the cradle of humanity. One widely publicized 1987 study compared mitochondrial gene snippets from 147 people across the world, and concluded that Africans have the highest mitochondrial diversity, suggesting that our species originated and spent most of its evolutionary history there. Specifically, the authors traced all human mitochondrial diversity back to a single theoretical woman who lived in East Africa hundreds of thousands of years ago, whom the media popularized as “mitochondrial Eve.” Later studies estimated that the most recent common ancestor of modern Y chromosome variation (dubbed “Y chromosome Adam”) could also be traced back to Africa1. Subsequent studies of nuclear DNA have validated our African birthplace and refined our knowledge of the human genetic landscape. Several studies of genetic variation among modern-day Khoe and San individuals, two groups of indigenous people in southern Africa known for their click language, have suggested they represent our species’ most genetically diverse lineage. Collectively known as Khoe-San, this group is thought to have split from other populations between 200,000 and 350,000 years ago, making them the most ancient population of modern humans to diverge. Non-Africans, meanwhile, represent a reduced subset of the diversity in Africa, and likely trace most of their ancestry back to just one small population—probably no more than a few thousand individuals—who ventured out of the continent between 60,000 and 70,000 years ago2. Some scientists see the extraordinary diversity in modern Khoe-San people as evidence that our species arose in southern Africa. Along with some archaeological evidence from the region3, that challenges the long-held idea of an East African origin, which was based on the fact that many early hominin fossils were found there. However, trying to pinpoint the precise location of our species’ origins from DNA is often criticized for the simple reason that people move around—it’s not known if the populations living in one place today were there hundreds or thousands of millennia ago. In fact, some researchers, including Scerri, Stringer, and Thompson, have recently constructed an entirely new theory of our origins: that anatomically modern humans didn’t arise from a single place, but gradually emerged from a web of interconnected populations sprawled across Africa—a continental gene-sharing bonanza that hominin lineages besides our own may have participated in. “It’s a good way to interpret the data we have right now,” says Niang. In addition to where we evolved, researchers are interested in how: which genes gave us a selective advantage to survive in particular environments, and which ancestors contributed to our genomes? Unfortunately, modern African DNA is severely underrepresented in genetic research, making these questions particularly challenging to answer. Most sequenced genomes are of European origin, with fewer than 2 percent coming from Africans. This dearth of African genomes is compounded by the fact that the genetic scaffold underlying some frequently studied traits such as skin pigmentation appear to be far more complex in Africans than in other populations, notes Brenna Henn, a population geneticist at the University of California, Davis. “The twelve to fifteen genes [for skin pigmentation] that people cite in Eurasian populations explain less than 25 percent of the variation in Africans.” African population history complicates matters further. Large-scale migrations pulled people back and forth across the continent for thousands of years. People from Eurasia also migrated back to Africa. Where people moved, they swapped their genes with local populations, shuffling patterns of ancestry across African genomes. This upheaval of ancient population structures creates one of the biggest challenges in teasing out archaic history from modern genomes, notes University of Pennsylvania geneticist Sarah Tishkoff. “It can make it very tricky to distinguish that older history when there’s been this newer wave of gene flow messing with your modeling.” Still, geneticists have been able to tease out some signals from our distant past, using computational models that ask what kind of evolutionary processes—such as mutation, selection, and interbreeding with other groups—best explain the pattern of variation across modern genomes. One intriguing finding of such studies is possible evidence of mixture with now-extinct, unknown groups of modern humans and other hominins: “ghost” populations that, like Neanderthals, left traces in modern genomes. In one analysis of 15 sequenced genomes, Tishkoff’s group investigated the sources of genetic variation in three different modern African hunter-gatherer groups4. The team’s models suggested that interbreeding with an archaic hominin species—which seemed as different from modern humans as are Neanderthals—was the most likely origin for a set of unusual sequences they found. “The model that includes a ghost population is always better [to fit the data], basically,” Tishkoff says. A handful of similar studies have also revealed traces of ghost hominins in modern African genomes, sometimes accounting for up to 10 to 20 percent of the genetic variation. Some research suggests that mixing took place after the ancestors of modern Eurasians left Africa, hinting that other kinds of hominins could have existed alongside Homo sapiens in Africa until very recently. “It’s actually pretty convincing,” says Henn, who wasn’t involved in these studies. “Ten percent of the genome—I’m going to have a hard time invoking one single other process that can explain a signal like that.” Ultimately, researchers need samples of DNA from ancient hominins to prove whether archaic African species did in fact contribute to modern genetic variation. While scientists have managed to overcome some of the technical hurdles of sequencing highly degraded ancient DNA from human fossils in Africa, the oldest human DNA found on the continent is just 15,000 years old, an age that pales in comparison to some 400,000-year-old hominin DNA found in a cave in Spain with relatively cool, stable temperatures. Archaeologists can only dream of finding intact DNA that old on the African continent, notes Tessa Campbell, an ancient DNA specialist at Iziko Museums of South Africa. “No one wants to say never . . . but it’s very unlikely.” Because DNA is unlikely to survive very long in the African heat, researchers have largely refrained from drilling into the fossils they’ve found of other hominins in Africa for fear of destroying them. But efforts are underway to study ancient DNA from younger fossils of Homo sapiens to crack other mysteries about human history on the continent, Tucci notes. “This is definitely a new era for African genomics.” 1. science.sciencemag.org/content/278/5339/804.summary2. academic.oup.com/mbe/advance-article/doi/10.1093/molbev/msaa140/58749453. www.pnas.org/content/108/13/51544. www.cell.com/cell/fulltext/S0092-8674(12)00831-8?_returnURL=https://linkinghub.elsevier.com/retrieve/pii/S0092867412008318?showall=true |
|
|
|
Post by Admin on Sept 3, 2020 7:04:53 GMT
Excavating a Continent A number of researchers suspect that Homo sapiens arose not in a single place in Africa, but across the entire continent, emerging from a network of interconnected hominin populations. But for decades, archaeologists positioned East and South Africa as important places for hominin evolution and the putative birthplace of our species. That’s likely because most fossils, including groundbreaking findings that have transformed our understanding of human evolution, have been found in those regions.  KABWE, ZAMBIA, 1921 “Kabwe skull,” 300,000 years ago Also called “Broken Hill skull,” the specimen is considered a representative of Homo heidelbergensis. NEAR SAFI, MOROCCO, 1961 Human remains at Jebel Irhoud, 315,000 years ago Flint blades and Homo sapiens–like skeletons in a Moroccan cave known as Jebel Irhoud may represent the oldest Homo sapiens artifacts. The skeletons have modern features such as round skulls and modern-human–like teeth and faces. OMO NATIONAL PARK, ETHIOPIA, 1967-1974 Omo Kibish remains, 195,000 years ago Fragments from two skulls, four jaws, a legbone, a few hundred teeth, and some other bones were found at a site in Ethiopia, and are classified as anatomically modern Homo sapiens. AFAR REGION, ETHIOPIA, 1974 “Lucy,” 3.2 million years ago Lucy—the skeletal remains of an Australopithecus afarensis female—is one of the best-known hominin fossils. Studies suggest that she was both tree-dwelling and capable of an upright gait, providing an important evolutionary stepping stone from more primitive ape species to modern humans. LAKE TURKANA, KENYA, 1984 “Turkana Boy,” 2 million years ago A nearly complete skeleton of an ancient Homo erectus child found near Kenya’s Lake Turkana provides a rare glimpse into how quickly this species reached adulthood and how similar their skeletons were to ours. RISING STAR CAVE, SOUTH AFRICA, 2013 Homo naledi, 236,000–335,000 years ago In 2013 and 2014, cavers found skeletons of two adults and one juvenile of what is believed to be a new species: Homo naledi. Its tiny brain and ape-like shoulders—indicating it was a good climber—suggest it may be an evolutionary off-shoot lineage that went extinct. AFAR REGION, ETHIOPIA, 2013 Adult jawbone, 2.8 million years ago A mandible fragment is the earliest known trace of the genus Homo, although the species it belongs to is a mystery. In 2015, an international team of researchers managed to harvest the first ancient DNA1 in Africa—the genome of Mota, a man who left behind 4,500-year-old remains in an Ethiopian cave. In the five years since that publication, researchers have published nearly 100 other full and partial ancient human sequences from Africa2. These genomes have helped scientists better understand the messy signatures from recent migration events that make studies of modern genomes so difficult. For instance, mitochondrial DNA from the skulls of seven people who lived some 15,000 years ago in modern-day Morocco revealed that they were closely related to Natufians, hunter-gatherers who dwelled in the Near East, as well as people living south of the Sahara desert. This finding suggested that there were far-flung connections between North Africa, the Near East, and sub-Saharan Africa before the dawn of agriculture3. Analyses of ancient DNA have also helped researchers understand how ancient migrations affected the genomes of people alive today. One such migration is the Bantu expansion, which gradually spread West African farming practices across the continent between roughly 5,000 and 1,000 years ago. By comparing DNA from ancient hunter-gatherer remains in southern Africa with modern-day Khoe-San people, evolutionary biologist Carina Schlebusch of Uppsala University in Sweden and her colleagues found that some Khoe-San groups carry DNA that ancient farmers brought with them. They also carry mixed Eurasian ancestry that had been introduced to North Africa with earlier back-migrations into the continent and eventually carried to the southernmost tip of Africa as other migrating human populations moved southward, the researchers found4. Such studies have also provided insight into deep divergences that occurred in human populations long before migrations of farmers and herders. Mary Prendergast, an anthropologist at Saint Louis University in Madrid, and her colleagues recently sequenced the first ancient DNA from West Africa, material extracted from the remains of children buried inside a rock shelter in Cameroon5. Comparing the 3,000- and 8,000-year-old DNA with ancient genomes collected elsewhere and with genomes of modern people allowed the researchers to reconstruct some of the earliest branches of our species’ evolutionary tree. In addition to the deep split between Khoe-San groups and other African populations—from which non-Africans also descend—their model suggested that two other major lineages split just as deeply, diverging from one another more than 200,000 years ago. One lineage is ancestral to central African hunter-gatherers known as Aka and Mbuti, and the second is a previously unknown “ghost” lineage whose fate is uncertain. “There’s all this deep, deep population structure with various differentiated branches of the human tree throughout the Pleistocene in Africa that we haven’t really appreciated very much yet,” Prendergast says. Only time will tell whether researchers’ current arsenal of technologies is enough to untangle the complete story of human evolution. Perhaps novel technologies—such as paleoproteomics, a nascent field that aims to reconstruct ancestry from fossilized proteins, which are more durable than DNA—will help researchers “push further back in time,” notes biological anthropologist Rebecca Ackermann of the University of Cape Town. What is already abundantly clear is that human evolution was far more complex than previously appreciated by anthropologists. It was not a streamlined process of australopiths steadily evolving into modern humans, but a messy and haphazard journey that includes interwoven ancestries of many groups, some of which have never been discovered other than through the genetic traces they left in ancient and modern genomes. “We have a long history. A lot of things happened, and a lot of ancestors contributed to our genomes today,” Schlebusch says. “It’s not going to be a simple story.” Decolonizing Studies of Human Evolution The San people of southern Africa are one of the most intensively studied indigenous groups in the world. Their click language and traditional hunter-gatherer lifestyles have long fascinated anthropologists. And the antiquity of their genetic lineage makes them a treasure trove for geneticists studying human evolutionary history. However, studies on San lifestyles and genomes have not always been conducted ethically. For instance, scientists have sometimes referred to the San as “bushmen,” a derogatory term associated with colonial-era researchers using modern indigenous groups as models of primitive human ancestors, and have taken photographs of children and breastfeeding mothers without permission. “We’re not saying that everybody is bad. But you get those few individuals who don’t respect the San,” Leana Snyders, head of the South African San Council in Upington, South Africa, told Science in 2017. Ethical conduct in genomic research came to the foreground in 2010 following a high-profile analysis of San genomes in Nature in which the authors had, among other transgressions, not asked San leaders for permission to conduct the study. All disciplines that study human evolution in Africa have at times been criticized for their extractive nature. Archaeological research—a field pioneered by European colonial nations—has long been driven by Western researchers digging up fossils from Africa to study them, sometimes taking them elsewhere to do so. Some hominin fossils are still displaced, such as the Kabwe skull, a famous Homo heidelbergensis specimen that remains in London’s Museum of Natural History, despite Zambia’s multiple requests to repatriate the skull. According to an April press release, the museum has approached Zambian authorities to begin discussing the possible return of the skull following a 2018 agreement between the UK and Zambia to find a solution to the issue. Some scientists have called for regulations to protect fossil collections from ancient DNA research, whereby African hominin fossils undergo the damaging process of extracting DNA. Now, “African museums are taking a leading role to make sure this [research] happens through collaboration and regulation,” notes anthropologist Mary Prendergast of Saint Louis University’s Madrid campus, as geneticists are working to develop new, less destructive techniques for ancient DNA analysis.  The San, for their part, created a code of research conduct in 2017 that, for example, requires researchers to respect their communities and to allow them to comment on findings prior to publication to avoid derogatory interpretations. Researchers are also required to compensate the community for their cooperation, through financial support, knowledge, or job opportunities, for instance. A number of scientists have called for a greater role of African scientists in human evolutionary research. To make that possible, Western funding agencies and institutions have an obligation to support African efforts to improve their countries’ antiquities infrastructure, so that “the next generation of African scholars [can] take control of the research in their areas,” notes anthropologist Eleanor Scerri of the Max Planck Institute in Germany. Foreign research teams should also foster stronger collaboration with African researchers, rather than simply seeking their help with fossil excavations, which has sometimes been the case, notes University of Cape Town biological anthropologist Rebecca Ackermann. Research groups have become more diverse, she notes, but the transition is slow. “I do see a change. It’s just not as fast as I would like.” 1. science.sciencemag.org/content/350/6262/8202. www.sciencedirect.com/science/article/pii/S0959437X203005993. science.sciencemag.org/content/360/6388/5484. science.sciencemag.org/content/358/6363/652.abstract5. www.nature.com/articles/s41586-020-1929-1 |
|
|
|
Post by Admin on Sept 5, 2020 5:44:03 GMT
Ancient Ethiopian genome reveals extensive Eurasian admixture in Eastern Africa M. Gallego Llorente1,*,†, E. R. Jones2,*,†, A. Eriksson1,3, V. Siska1, K. W. Arthur4, J. W. Arthur4, M. C. Curtis5,6, J. T. Stock7, M. Coltorti8, P. Pieruccini8, S. Stretton9, F. Brock10,11, T. Higham10, Y. Park12, M. Hofreiter13,14, D. G. Bradley2, J. Bhak15, R. Pinhasi16,*,‡, A. Manica1,*,‡ Science 13 Nov 2015: Vol. 350, Issue 6262, pp. 820-822 DOI: 10.1126/science.aad2879 Abstract Characterizing genetic diversity in Africa is a crucial step for most analyses reconstructing the evolutionary history of anatomically modern humans. However, historic migrations from Eurasia into Africa have affected many contemporary populations, confounding inferences. Here, we present a 12.5× coverage ancient genome of an Ethiopian male (“Mota”) who lived approximately 4500 years ago. We use this genome to demonstrate that the Eurasian backflow into Africa came from a population closely related to Early Neolithic farmers, who had colonized Europe 4000 years earlier. The ability to sequence ancient genomes has revolutionized our understanding of human evolution. However, genetic analyses of ancient material have focused on individuals from temperate and Arctic regions, where ancient DNA is preserved over longer time frames (1). Africa has so far failed to yield skeletal remains with much ancient DNA, with the exception of a few poorly preserved specimens from which only mitochondrial DNA could be extracted (2). This is particularly unfortunate, as African genetic diversity is crucial to most analyses reconstructing the evolutionary history of anatomically modern humans, by providing the baseline against which other events are defined. In the absence of ancient DNA, geneticists rely on contemporary African populations, but a number of historic events, in particular a genetic backflow from West Eurasia into Eastern Africa (3, 4), act as confounding factors. Here, we present an ancient human genome from Africa and use it to disentangle the effects of recent population movement into Africa. By sampling the petrous bone (5), we sequenced the genome of a male from Mota Cave (herein referred to as “Mota”) in the southern Ethiopian highlands, with a mean coverage of 12.5× (6). Contamination was estimated to be between 0.29 and 1.26% (6). Mota’s remains were dated to ~4500 years ago [direct calibrated radiocarbon date (6)] and thus predate both the Bantu expansion (7) and, more importantly, the 3000-year-old West Eurasian backflow, which has left strong genetic signatures in the whole of Eastern and, to a lesser extent, Southern Africa (3, 4). We compared Mota to contemporary human populations (6). Both principal component analysis (PCA) (Fig. 1A) and outgroup f3 analysis using Ju|’hoansi (Khoisan) from Southern Africa as the outgroup (Fig. 1, B and C) place this ancient individual close to contemporary Ethiopian populations, and more specifically to the Ari, a group of Omotic speakers from southern Ethiopia, to the west of the highland region where Mota lived. Our ancient genome confirms the view that the divergence of this language family results from the relative isolation of its speakers (8), and indicates population continuity over the last ~4500 years in this region of Eastern Africa.  Fig. 1 Mota shows a very high degree of similarity with the highland Ethiopian Ari populations. (A) PCA showing Mota projected onto components loaded on contemporary African and Eurasian populations. The inset magnifies the PCA space occupied by Ethiopian and Eastern African populations. (B) Outgroup f3 quantifying the shared drift between Mota and contemporary African populations, using Ju|’hoansi (Khoisan) as an outgroup; bars represent SE. (C) Map showing the distribution of outgroup f3 values across the African continent. In (A) and (B), populations speaking Nilo-Saharan languages are marked with blue shades, Omotic speakers with red, Cushitic with orange, Semitic with yellow, and Bantu with green. Mota is denoted by a black symbol. The age of Mota means that he should predate the West Eurasian backflow, which has been dated to ~3000 years ago (3, 4). We formally tested this proposition by using an f4 ratio estimating the West Eurasian component (6), following the approach adopted by Pickrell et al. (3). As expected, we failed to find any West Eurasian component in Mota (table S5), thus providing support for previous dating of that event (3, 4). Given that Mota predates the backflow, we searched for its most likely source by modeling the Ari, the contemporary population closest to our ancient genome, as a mixture of Mota and another West Eurasian population (6). We investigated both contemporary sources (3) and other Eurasian ancient genomes (5, 9). In this analysis, contemporary Sardinians and the early Neolithic LBK (Stuttgart) genome stand out (Fig. 2A). Previous analyses have shown Sardinians to be the closest modern representatives of early Neolithic farmers (10, 11), implying that the backflow came from the same genetic source that fueled the Neolithic expansion into Europe from the Near East/Anatolia, before recent historic events changed the genetic makeup of populations living in that region. An analysis with haplotype sharing also identified a connection between contemporary Ethiopians and Anatolia (4, 12). Interestingly, archaeological evidence dates the arrival of Near Eastern domesticates (such as wheat, barley, and lentils) to the same time period (~3000 years ago) (13, 14), suggesting that the direct descendants of the farmers that earlier brought agriculture into Europe may have also played a role in the development of new forms of food production in the Horn of Africa.  Fig. 2 Quantifying the geographic extent and origin of the West Eurasian component in Africa. (A) Admixture f3 identifying likely sources of the West Eurasian component (lowest f3 values). Contemporary populations in blue, ancient genomes in red; bars represent SE. (B) Map showing the proportion of West Eurasian component, λMota,LBK, across the African continent. Using Mota as an unadmixed African reference and the early farmer LBK as the source of the West Eurasian component, it is possible to reassess the magnitude and geographic extent of historical migrations, avoiding the complications of using admixed contemporary populations (6). We estimated a substantially higher Eurasian backflow admixture than previously detected (3), with an additional 4 to 7% of the genome of most African populations tracing back to a Eurasian source. Moreover, we detected a much broader geographical impact of the backflow, going all the way to West and Southern Africa (Fig. 2B). Even though the West Eurasian component in these regions is smaller than in Eastern Africa, it is still sizable, with Yoruba and Mbuti, who are often used as African reference populations (15, 16), showing 7% and 6%, respectively, of their genomes to be of Eurasian origin (table S5). Since Mota predates recent demographic events, his genome can act as an ideal African reference to understand episodes during the out-of-Africa expansion. We used him as the African reference to quantify Neandertal introgression in a number of contemporary genomes (6). Both Yoruba and Mbuti, which are routinely used as African references for this type of analysis (15, 16), show a marginally closer affinity with Neandertal than Mota on the basis of D statistics, and an f4 ratio analysis detected a small Neandertal component in these genomes at around 0.2 to 0.7%—greater than previously suggested (16) and consistent with our estimates of the magnitude of their Western Eurasian ancestry (6). Although the magnitude of Neandertal ancestry in these contemporary African populations is not enough to change conclusions qualitatively (estimates of Neandertal ancestry in French and Han only increased marginally when tested with Mota as a reference), it should be accounted for when looking for specific introgressed haplotypes (17) or searching for unknown ancient hominins who might have hybridized with African populations (18). We also investigated the Mota genome for a number of phenotypes of interest (6). As expected, Mota lacked any of the derived alleles found in Eurasian populations for eye and skin color, suggesting that he had brown eyes and dark skin. Mota lacked any of the currently known alleles that confer lactose tolerance, which may have implications concerning when pastoralism appeared in southwestern Ethiopia. In addition, Mota did possess all three selected alleles that recently have been shown to play a role in the adaptation to altitude in contemporary highland Ethiopian populations (19). The presence of these mutations supports our conclusion that Mota is the descendant of highland dwellers, who have lived in this environment long enough to accumulate adaptations to the altitude (20, 21). Until now, it has been necessary to use contemporary African populations as the baseline against which events during the worldwide expansion of anatomically modern humans are defined (16, 22–24). By obtaining an ancient whole genome from this continent, we have shown that having an unadmixed reference that predates the large number of recent historical migrations can greatly improve our inference. This result stresses the importance of obtaining unadmixed baseline data to reconstruct demographic events, and the limitations of analyses that are solely based on contemporary populations. Even older African genomes will thus be needed to investigate key demographic events that predate Mota, such as earlier instances of backflows into Africa (25). |
|
|
|
Post by Admin on Sept 10, 2020 21:10:06 GMT
Evolutionary History and Adaptation from High-Coverage Whole-Genome Sequences of Diverse African Hunter-Gatherers Joseph Lachance Benjamin Vernot Clara C. Elbers Kun Zhang Joshua M. Akey Sarah A. Tishkoff Open Archive Published:July 26, 2012 DOI:https://doi.org/10.1016/j.cell.2012.07.009 Summary To reconstruct modern human evolutionary history and identify loci that have shaped hunter-gatherer adaptation, we sequenced the whole genomes of five individuals in each of three different hunter-gatherer populations at >60× coverage: Pygmies from Cameroon and Khoesan-speaking Hadza and Sandawe from Tanzania. We identify 13.4 million variants, substantially increasing the set of known human variation. We found evidence of archaic introgression in all three populations, and the distribution of time to most recent common ancestors from these regions is similar to that observed for introgressed regions in Europeans. Additionally, we identify numerous loci that harbor signatures of local adaptation, including genes involved in immunity, metabolism, olfactory and taste perception, reproduction, and wound healing. Within the Pygmy population, we identify multiple highly differentiated loci that play a role in growth and anterior pituitary function and are associated with height. Graphical Abstract  Highlights 13.4 million variants identified in African hunter-gatherers, many of which are novel Evidence of archaic admixture found in the genomes of African hunter-gatherers Selection scans implicate loci involved in taste perception, metabolism, and immunity Genetic associations with height are found for Pygmy variants located on chromosome 3 Introduction Due to recent advances in sequencing technologies, whole-genome sequencing of multiple individuals in multiple populations is now feasible (Henn et al., 2010). This burgeoning field of population genomics allows inference of demographic history and natural selection that is free of SNP ascertainment bias (Luikart et al., 2003; Teo et al., 2010). Initial efforts of the 1000 Genomes Project have already identified millions of variants in diverse populations (1000 Genomes Project Consortium, 2010). However, whole-genome sequencing in the 1000 Genomes Project has generally been at low coverage, and genetic diversity in many ethnically diverse populations is yet to be characterized, particularly with respect to Africa, the ancestral homeland of all modern humans (Campbell and Tishkoff, 2010). Furthermore, differences in diet, climate, and exposure to pathogens among ethnically and geographically diverse African populations are likely to have produced distinct selection pressures, resulting in local genetic adaptations. Despite the important role that African populations have played in human evolutionary history, they remain one of the most understudied groups in human genomics. To date, few high-coverage African genomes have been analyzed in published studies: four Yoruba, one !Xhosa, and one San (1000 Genomes Project Consortium, 2010; Bentley et al., 2008; Drmanac et al., 2010; Schuster et al., 2010). A comprehensive knowledge of levels and patterns of variation in African genomes is critical for a deeper understanding of (1) human genetic diversity, (2) the allelic spectrum of functionally important genetic variation, including disease susceptibility, (3) the genetic basis of adaptation to diverse environments, and (4) the origins of modern humans. Previous analyses of mtDNA, Y chromosome, and autosomal genetic lineages in African hunter-gatherer populations indicate that they have some of the deepest divergence times of our species (Tishkoff et al., 2007, 2009; Veeramah et al., 2012). The forest-dwelling Pygmies of Cameroon include the Baka, Bakola, and Bedzan populations. As a result of the expansion of Bantu-speaking agriculturalists within the last few thousand years into Pygmy territories, Cameroonian Pygmy populations are highly admixed (Jarvis et al., 2012) and the ancestral Pygmy language has been lost (Berniell-Lee et al., 2009; Verdu et al., 2009). The Hadza and Sandawe populations live in Tanzania and speak languages that contain click-consonants and are classified as Khoesan, but these languages are highly divergent from each other and from Southern African San Khoesan-speaking populations (Sands, 1998). At present, little is known about the genetic relationships between click-speaking populations (Henn et al., 2011; Tishkoff et al., 2007, 2009; Veeramah et al., 2012). The Sandawe currently number more than 30,000 individuals, and following the Bantu expansion, many Sandawe have switched from nomadic hunting to an agricultural subsistence pattern (Newman, 1970). By contrast, the Hadza have a current population size of ∼1,000 individuals, the vast majority of whom still practice a hunter-gatherer lifestyle (Blurton Jones et al., 1992; Marlowe, 2010). Data from autosomes, mtDNA, and the Y chromosome indicate that, although the Hadza, Sandawe, Pygmy, and San populations have an ancient divergence, they also share lineages that are rare in other populations (Behar et al., 2008; Henn et al., 2011; Tishkoff et al., 2007, 2009). To extend our knowledge of genomic variation in ethnically diverse Africans, we sequenced the genomes of five males from each of three African hunter-gatherer populations (Western Pygmy, Hadza, and Sandawe) at high coverage (Figure 1A). We then compared these genome sequences to a previously published genome sequence from a San hunter-gatherer (Schuster et al., 2010) and to publicly available whole-genome sequence data from other ethnically, linguistically, and geographically diverse African populations (http://www.completegenomics.com/sequence-data/download-data/).  Figure 1 Genomic Variation in African Hunter-Gatherers and Other Global Populations |
|











