|
|
Post by Admin on Feb 24, 2021 5:14:58 GMT
HSE University researchers have become the first in the world to discover genetic predisposition to severe COVID-19. The results of the study were published in the journal Frontiers in Immunology. T-cell immunity is one of the key mechanisms used by the human body to fight virus infections. The staging ground for cell immunity development is the presentation of virus peptides on the surface of infected cells. This is followed by activation of T lymphocytes, which start to kill the infected cells. The ability to successfully present virus peptides is largely determined by genetics. In human cells, human leukocyte antigen class I (HLA-I) molecules are responsible for this presentation. The set of six such molecules is unique in every human and is inherited from an individual's parents. In simple terms, if the set of alleles detects the virus well, then the immune cells will detect and destroy the infected cells fast; if a person has a set that is bad at such detection, a more severe case of disease is more likely to be observed. Researchers from the HSE Faculty of Biology and Biotechnology - Maxim Shkurnikov, Stepan Nersisyan, Alexei Galatenko and Alexander Tonevitsky -together with colleagues from Pirogov Russian National Research Medical University and Filatov City Clinical Hospital (Tatjana Jankevic, Ivan Gordeev, Valery Vechorko) studied the interconnection between HLA-I genotype and the severity of COVID-19.  Using machine learning, they built a model that provides an integral assessment of the possible power of T-cell immune response to COVID-19: if the set of HLA-I alleles allows for effective presentation of the SARS-CoV-2 virus peptides, those individuals received low risk score, while people with lower presentation capability received higher risk scores (in the range from 0 to 100). To validate the model, genotypes of over 100 patients who had suffered from COVID-19 and over 400 healthy people (the control group) were analysed. It turned out that the modelled risk score is highly effective in predicting the severity of COVID-19. In addition to analysing the Moscow population, the researchers used their model on a sample of patients from Madrid, Spain. The high precision of prediction was confirmed on this independent sample as well: the risk score of patients suffering severe COVID-19 was significantly higher than in patients with moderate and mild cases of the disease. 'In addition to the discovered correlations between the genotype and COVID-19 severity, the suggested approach also helps to evaluate how a certain COVID-19 mutation can affect the development of T-cell immunity to the virus. For example, we will be able to detect groups of patients for whom infection with new strains of SARS-CoV-2 can lead to more severe forms of the disease,' Alexander Tonevitsky said. ### Once the paper is published, it will be available to view online at doi.org/10.3389/fimmu.2021.641900/. EMBARGOED until 23 February 2021, 00:00 US Eastern time / 06:00 (CET). |
|
|
|
Post by Admin on Feb 25, 2021 3:00:58 GMT
Association of HLA Class I Genotypes With Severity of Coronavirus Disease-19 Maxim Shkurnikov1†, Stepan Nersisyan1*†, Tatjana Jankevic2, Alexei Galatenko1,3, Ivan Gordeev2,4, Valery Vechorko4 and Alexander Tonevitsky1,5* Human leukocyte antigen (HLA) class I molecules play a crucial role in the development of a specific immune response to viral infections by presenting viral peptides at the cell surface where they will be further recognized by T cells. In the present manuscript, we explored whether HLA class I genotypes can be associated with the critical course of Coronavirus Disease-19 by searching possible connections between genotypes of deceased patients and their age at death. HLA-A, HLA-B, and HLA-C genotypes of n = 111 deceased patients with COVID-19 (Moscow, Russia) and n = 428 volunteers were identified with next-generation sequencing. Deceased patients were split into two groups according to age at the time of death: n = 26 adult patients aged below 60 and n = 85 elderly patients over 60. With the use of HLA class I genotypes, we developed a risk score (RS) which was associated with the ability to present severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) peptides by the HLA class I molecule set of an individual. The resulting RS was significantly higher in the group of deceased adults compared to elderly adults . In particular, presence of HLA-A*01:01 allele was associated with high risk, while HLA-A*02:01 and HLA-A*03:01 mainly contributed to low risk. The analysis of patients with homozygosity strongly highlighted these results: homozygosity by HLA-A*01:01 accompanied early deaths, while only one HLA-A*02:01 homozygote died before 60 years of age. Application of the constructed RS model to an independent Spanish patients cohort (n = 45) revealed that the score was also associated with the severity of the disease. The obtained results suggest the important role of HLA class I peptide presentation in the development of a specific immune response to COVID-19.
1. Introduction
Human leukocyte antigen (HLA) class I molecules are one of the key mediators of the first links in the development of a specific immune response to Coronavirus Disease-19 (COVID-19) infection. Right after entering the cell, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) induces the translation of its proteins. Some of these proteins enter the proteasomes of the infected cell, become cleaved to peptides the length of 8–12 amino acid residues, and bind to HLA class I receptors. After binding, the complex consisting of the HLA class I molecule and the peptide is transferred to the surface of the infected cell, where it can interact with the T cell receptor of CD8+ T lymphocytes. In response to the interaction, the CD8+ T lymphocyte activates and starts to divide; in 5–7 days, a population of virus-specific cytotoxic CD8+ T lymphocytes capable of destroying infected cells using perforins and serine proteases is formed (1). The crucial role of long-term CD8+ T cells activation in the immune response to COVID-19 has been recently studied in a cohort of patients who had mild disease (2, 3).
There are three main types of HLA class I receptors: HLA-A, HLA-B, and HLA-C. Receptors of every type are present in two variants inherited from parents. There exist dozens of variants of each allele of HLA-I receptors; every allele has an individual ability to recognize various foreign proteins. The distribution of alleles is population/country specific (4).
Individual combinations of HLA class I receptors essentially affect the severity of multiple infectious diseases, including malaria (5), tuberculosis (6), HIV (7), and viral hepatitis (4). There are a number of reported interconnections between the HLA genotype and the sensitivity to SARS-CoV-2. For example, the alleles HLA-B*07:03 (8), HLA-B*46:01 (9), and HLA-C*08:01 (10) are factors of predisposition to a severe form of the disease, and the allele HLA-C*15:02 is associated with a mild form (11).
Information on the interconnection of HLA class I genotype and severity of the course of COVID-19 caused by SARS-CoV-2 is sparse. A sample of 45 patients with varying severity of COVID-19 was used to confirm the results of the theoretical modeling of interaction of SARS-CoV-2 peptides with various HLA-I alleles (12). It was demonstrated that the number of peptides with a high interaction constant are connected with individual HLA genotypes: the more viral peptides with high affinity bind to HLA class I, the easier the course of the disease. It was also shown that the frequency of the occurrence of HLA-A*01:01 and HLA-A*02:01 alleles is related to the number of infections and mortality rate in different regions of Italy (13).
In the present study, we explored whether HLA class I genotypes can be a factor contributing to the critical course of COVID-19. For that, we performed HLA genotyping for n = 111 deceased patients with COVID-19 and the control group (n = 428), and searched for putative associations between genotypes and age at death. Since the total number of distinct HLA class I genotypes is too high to perform frequency-based analysis, we assigned scores to each allele based on capability of presenting SARS-CoV-2 peptides. The obtained scores allowed us to make a valid statistical comparison between HLA genotypes in groups of deceased adults (completed age at death not >60 years, n = 26), elderly adults (age at death over 60, n = 85), and the control. Special attention was paid to “extreme” cases formed by homozygous individuals by some HLA genes. Additionally, we assessed the contribution of each viral protein to the constructed risk model. |
|
|
|
Post by Admin on Feb 25, 2021 4:25:21 GMT
3. Results 3.1. Distribution of HLA Class I Gene Alleles in the Cohort of Deceased COVID-19 Patients and the Control Group We performed HLA class I genotyping for n = 111 deceased patients with confirmed COVID-19 (Moscow, Russia) and the control group consisting of volunteers (n = 428). Deceased patients were divided into two groups: adults (age at death ≤ 60 years) and elderly adults (age at death over 60 years). Demographic and clinical data of these cohorts are summarized in Table 1. Although patients with severe comorbidities were excluded from the study, 76.6% of deceased patients had at least one underlying disease. Only cerebrovascular disease had a statistically significant odds ratio when comparing groups of adults and elderly adults (3.8 vs. 34.1%, Fisher's exact test p = 1.89 × 10−3). Other cardiovascular diseases like coronary artery disease and heart failure were also more frequent in the group of elderly individuals which, however, was not statistically significant. Interestingly, arterial hypertension was diagnosed in 11.5% of adult patients and 24.7% of older adults, which was generally less than populational level in Russia (about 50%) (23). Percentage of diabetes cases was about 3.5% in both analyzed groups, which is a typical value for the current population of Russia (24). Also, frequencies of chronic kidney disease (stages 4–5) in both groups (23.1% for adults and 16.5% for elderly individuals) was significantly higher compared to the background populational value (about 0.05%) (25). TABLE 1 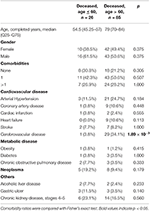 Demographic and clinical data in the cohort of deceased patients with COVID-19. First, we tested whether frequency of a single allele can differentiate individuals from three groups: adult patients who died from COVID-19, elderly patients who died from COVID-19, and the control group. Distribution of major HLA-A, HLA-B, and HLA-C alleles in these three groups is summarized in Figure 1. Fisher's exact test was used to make formal statistical comparisons. As a result, we found that for all possible group comparisons not a single allele had an odds ratio, which can be considered statistically significant after multiple testing correction (all corrected p-values were equal to 1). However, few of them were differentially enriched if no multiple testing corrections were applied (Supplementary Table 4). FIGURE 1 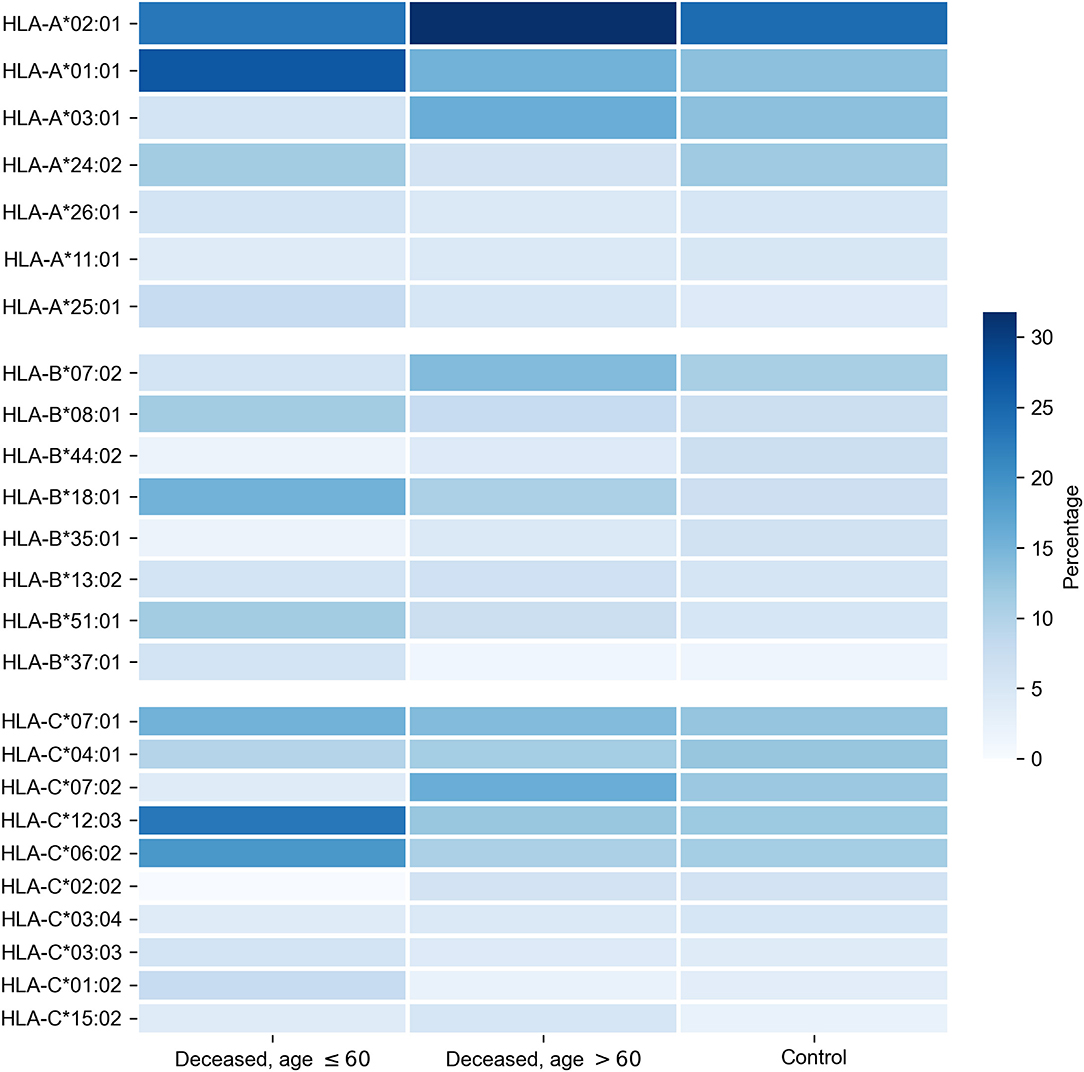 Distribution of HLA-A, HLA-B, HLA-C alleles in cohorts of deceased COVID-19 patients and the control group. Alleles with frequency over 5% in at least one of three considered groups are presented. |
|
|
|
Post by Admin on Feb 25, 2021 20:27:13 GMT
3.2. Binding Affinities of Viral Peptides to HLA Class I Molecules Since sizes of considered cohorts were insufficient for performing frequency analysis at the level of full HLA class I genotypes, we transformed patient genotypes from discrete space to numerical units associated with the potential of interactions with SARS-CoV-2 peptides. To implement this idea, we first constructed a matrix of binding affinities of viral peptides to HLA-A, HLA-B, and HLA-C alleles. For that, we first made computational predictions of viral peptides derived from SARS-CoV-2 strains isolated in Moscow. Then, binding affinities were calculated for each of the predicted peptides and each allele present in patients from the analyzed cohorts. As a result, we obtained a matrix containing affinity values for 6,548 peptides and 107 alleles of genes from major HLA class I. To establish a positive relationship between values from the matrix and binding potential, all affinities were inverted and scaled by a value of 500 nM (the conventional threshold for binding ability). Simultaneous hierarchical clustering was applied to identify groups of similar peptides and HLA-A, HLA-B, and HLA-C alleles (Figure 2). As can be seen, both alleles and peptides clearly formed several dense clusters. The most presented alleles of HLA-A (HLA-A*01:01, HLA-A*02:01, HLA-A*03:01, and HLA-A*24:02) fell in different clusters, while for HLA-B and HLA-C, some major alleles were grouped together (e.g., HLA-B*07:02, HLA-B*08:01, HLA-C*06:02, HLA-C*07:01, and HLA-C*07:02). FIGURE 2 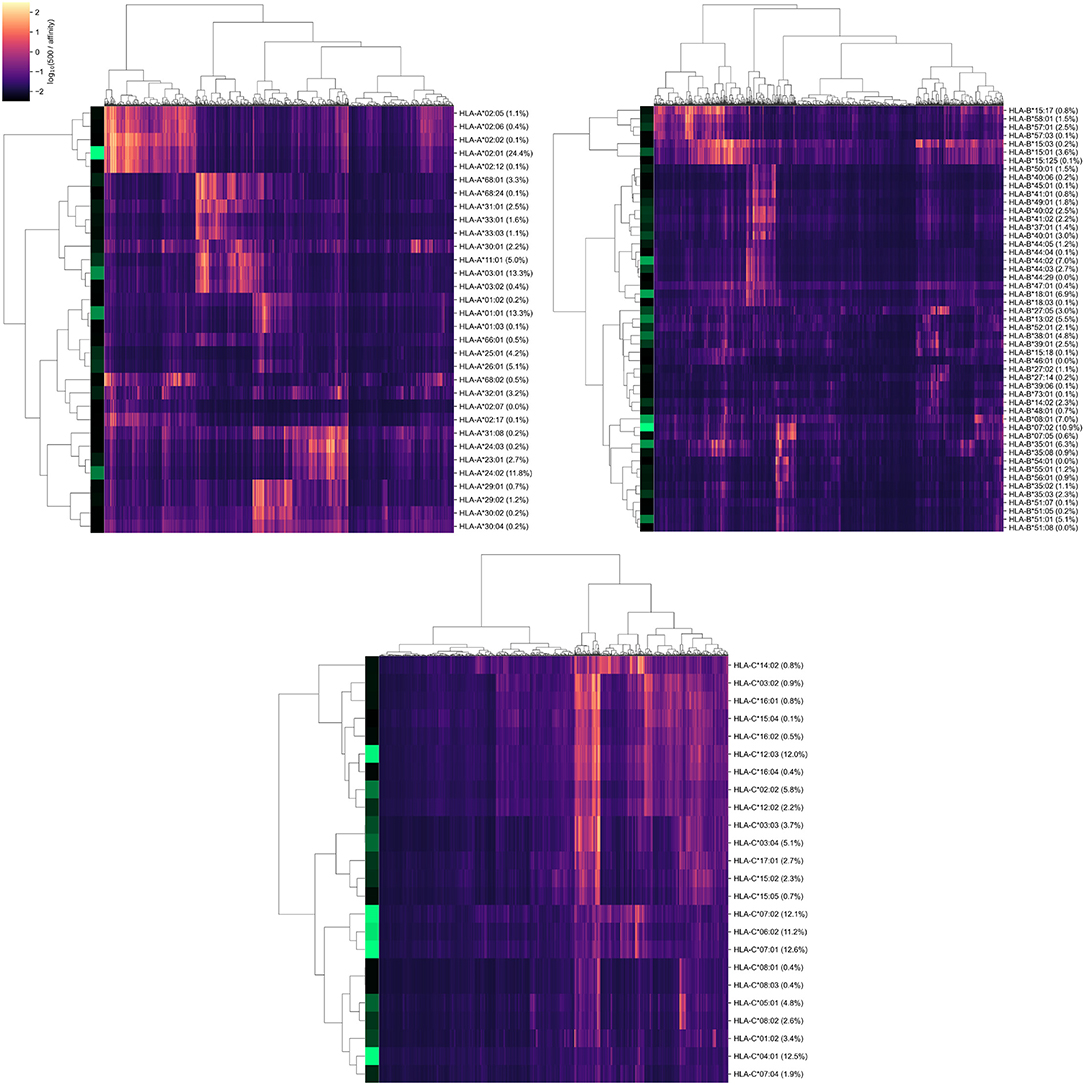 Hierarchical clustering of HLA-A, HLA-B, HLA-C gene alleles and SARS-CoV-2 peptides according to binding affinity matrix. Shades of green in vertical stripes and percents in brackets represent frequency of an allele in the control group. Zero percents refer to rare alleles found only in the group of deceased patients. Note that alleles with similar peptide binding profiles can be linked to different alleles of remaining genes. For example, consider closely clustered alleles HLA-C*06:02, HLA-C*07:01, and HLA-C*07:02. From the analysis of the contingency table of allele pairs in the control group (Figure 3), it follows that each of these alleles has its own spectrum of associated alleles. Specifically, HLA-C*06:02 usually appears with HLA-B*13:02 (Fisher's exact test p = 3.18 × 10−14), HLA-C*07:01 is linked to HLA-A*01:01 (p = 2.97 × 10−7) and HLA-B*08:01 (p = 3.15 × 10−14), while HLA-C*07:02 is coupled with HLA-A*03:01 (p = 9.66 × 10−4) and HLA-B*07:02 (p = 2.72 × 10−26). Interestingly, such linked alleles can have different peptide binding patterns (e.g., see weakly overlapped bars for HLA-A*01:01 and HLA-A*03:01 in Figure 2). FIGURE 3 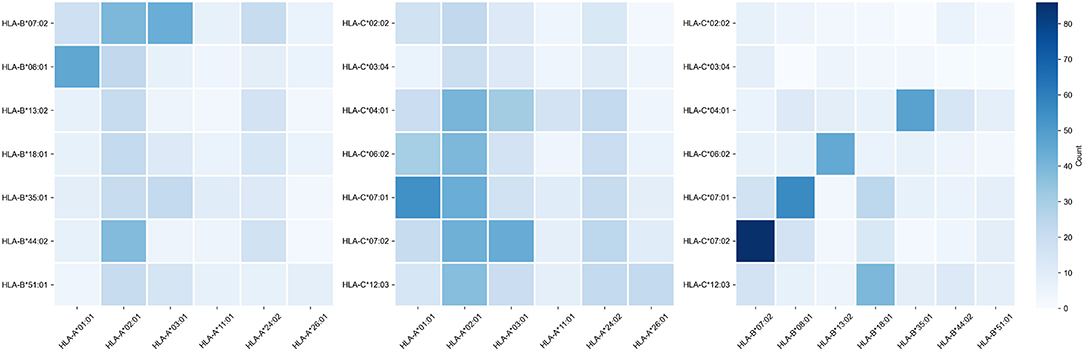 Contingency table of allele counts in the control group. Alleles with frequency over 5% in the control group are presented. |
|
|
|
Post by Admin on Feb 25, 2021 23:21:14 GMT
3.3. Risk Score Based on Peptide-HLA Binding Affinity Is Associated With Early COVID-19 Deaths For each of the considered HLA-A, HLA-B, and HLA-C alleles, we obtained the list of binding affinities to 6,548 unique SARS-CoV-2 peptides. In order to calculate aggregate information on the potential of presenting SARS-CoV-2 peptides by each allele, we used principal component analysis (PCA). In this framework 6,548-element affinity vectors are replaced by the most informative linear combinations of their components. For HLA-A and HLA-C, we found four principal components (PCs), each of which explained at least 5% of data variance, while for HLA-B, the number of essential components was equal to five (Supplementary Table 5). Signs of components were set in the way to achieve positive correlation of component values with age of death of deceased patients. For each individual, HLA class I gene, and PC, we summed PC values associated with two corresponding alleles. After that, we analyzed differences of obtained scores in adult and elderly patients who died from COVID-19. Three of the resulting PCs demonstrated statistically significant differences according to the Mann-Whitney U test. This list included the second and third PCs of HLA-A, and the fourth PC of HLA-C, while no PCs of HLA-B separated the analyzed groups significantly. As an aggregate risk score (RS), we considered the sum of these three components (for convenience, we linearly scaled the range of RS to the [0, 100] interval). The obtained score also significantly separated groups with p = 3.48 × 10−3 (U test) and AUC ROC equal to 0.68 (permutation test p = 3.10 × 10−3), see Figures 4A,B (full information is listed in Supplementary Table 6). Interestingly, the difference of RS distributions in the cohort of adult patients and the control group was also statistically significant (U test p = 3.31 × 10−3), while the difference between the elderly and control groups was not (p = 0.283). FIGURE 4 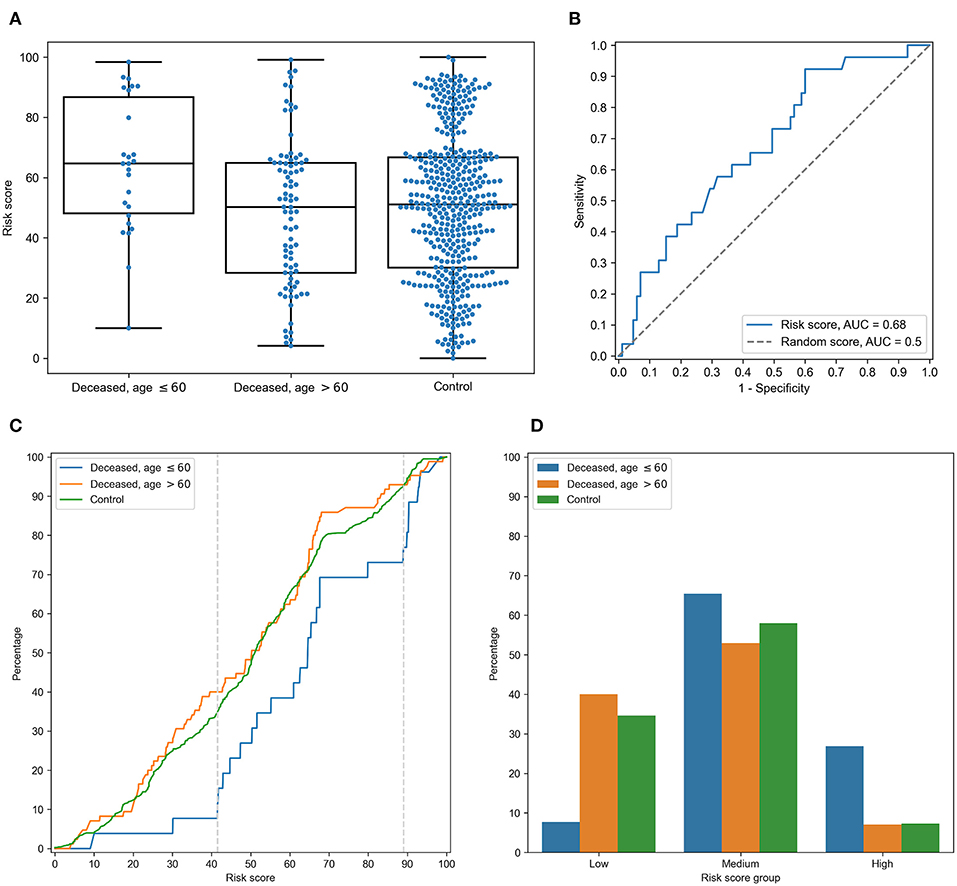 Risk score (RS) separates groups of adult and elderly patients. (A) Distribution of RS in adult, elderly and control patient groups. (B) Receiver operating characteristic curve for RS separating patients from adult and elderly groups. (C) Empirical distribution function of RS in three patient groups. Vertical dotted lines at RS = 41 and RS = 89 define ranges for three RS groups. (D) Distribution of three patient groups over low, medium and high RS. In order to characterize the association between RS and age at death more precisely, we partitioned the range of RS into three groups: low, medium, and high (Figures 4C,D). The lower and higher thresholds were calculated in a way to minimize p-value for Fisher's exact test applied to the number of adult and elderly patients in the whole cohort and in the low/high risk groups, respectively. Interestingly, such partitioning led to significant separation of adult patients both from the elderly and control subjects in the low and high risk groups, while no significance was found within the middle group (Supplementary Table 7). Then, we performed enrichment analysis to identify alleles significantly contributing to each of the RS groups (Table 2). As it can be seen, frequencies of several alleles were dramatically higher in some RS groups. Specifically, HLA-A*02:01 and HLA-A*03:01 were highly overrepresented in the low risk group and completely absent in the high risk group, while the most enriched allele in the high risk group was HLA-A*01:01. Reciprocally to HLA-A*02:01 and HLA-A*03:01 cases, not a single individual in the low risk group carried the HLA-A*01:01 allele. Complete information on allele frequencies in the RS groups is presented in Supplementary Figure 1. Table 2. Enrichment analysis of risk score groups. 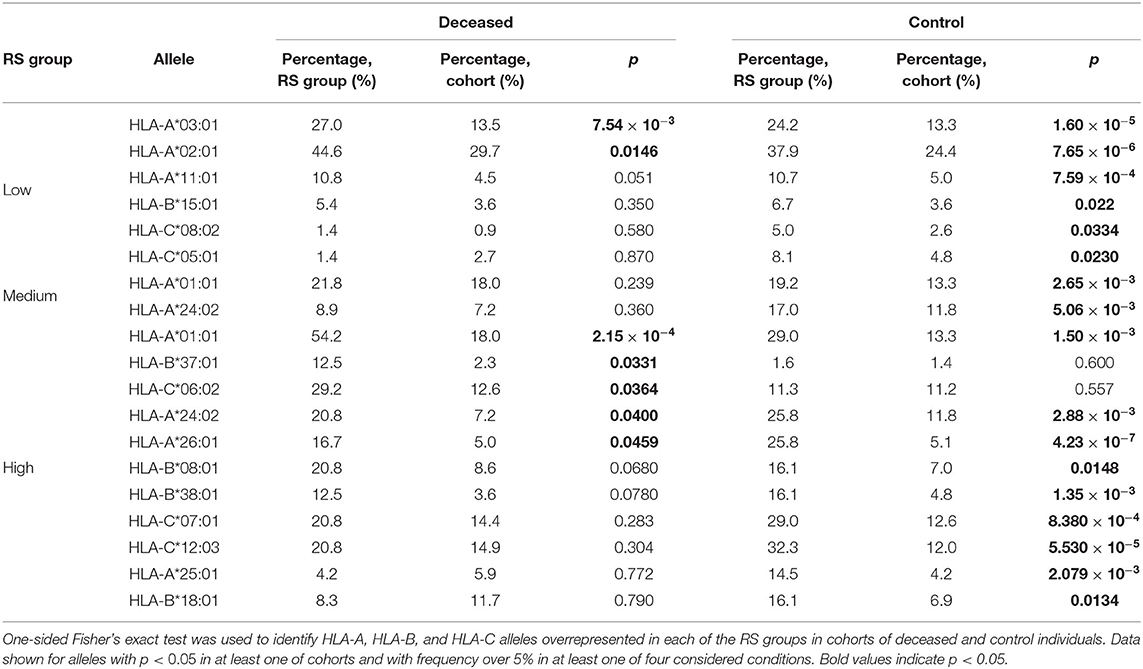 Finally, we assessed the contribution of individual peptides from different viral proteins to the RS. Contribution of a peptide to RS was calculated as an absolute value of the sum of corresponding PC coefficients (PC2 and PC3 for HLA-A, and PC4 for HLA-C). Then, the set of the most RS contributing peptides was composed by taking the top 5% of peptides from the corresponding distribution. The results of the procedure are summarized in Table 3: distribution of peptides with the strongest contributions over SARS-CoV-2 proteins was similar to the one calculated for all peptides after multiple testing correction. Without the correction, only non-structural protein 8 (NSP8) had a statistically significant odds ratio. Thus, considered peptides were spread over proteins without any significant dependence on contribution to the RS. Table 3. Distribution of peptides and viral proteins and their contribution to the RS. 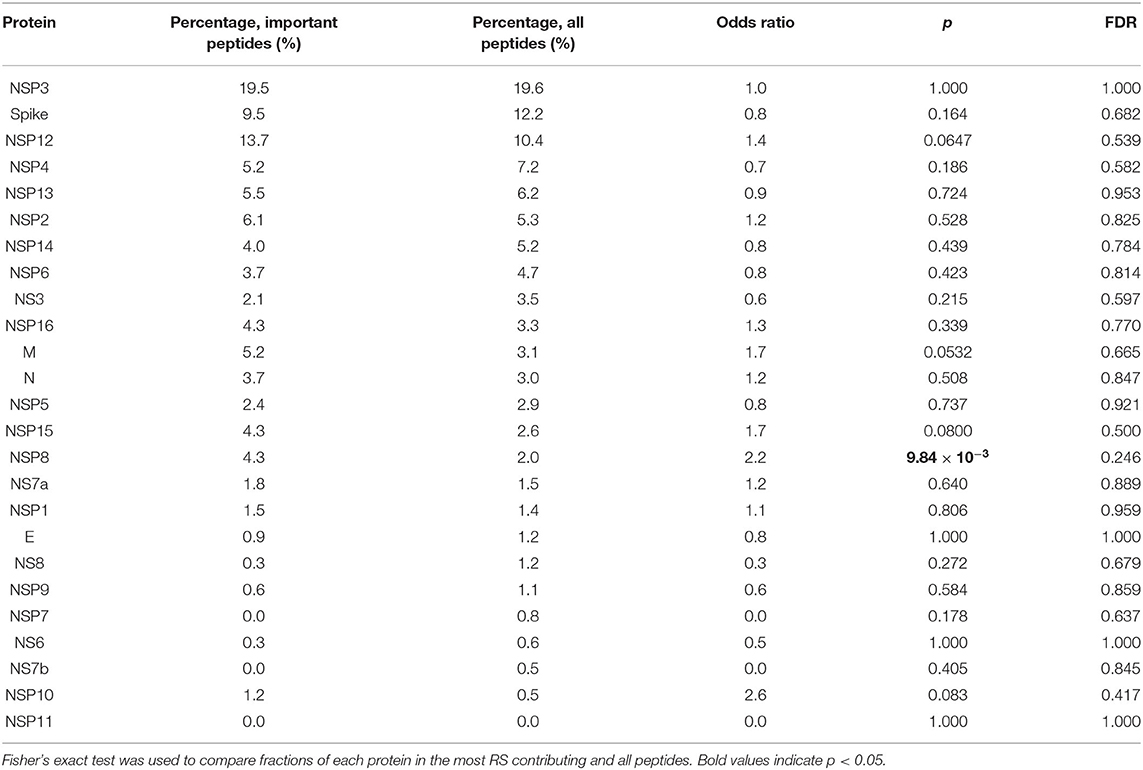 |
|









