Post by Admin on Dec 26, 2020 4:07:44 GMT
Details of emergent variant
VOC 202012/01 is defined by 17 mutations (14 non-synonymous mutations and 3 deletions),
among which eight are located in the spike protein. At least three mutations have a potential
biological significance. Mutation N501Y is one of the key contact residues in the receptor
binding domain and has been shown to enhance binding affinity to human ACE2 (2, 3). The
function of mutation P681H is unclear, but it is located immediately adjacent to the furin
cleavage site in spike, a known region of importance for infection and transmission (4, 5). The
deletion of two amino acids at positions 69-70 in spike has arisen in multiple independent
circulating lineages of SARS-CoV-2, is linked to immune escape in immunocompromised
patients and enhances viral infectivity in vitro (6, 7). This deletion is also responsible for certain
commercial diagnostic assays failing to detect the spike glycoprotein gene (S gene drop-out),
with genomic data confirming these S gene target failures are primarily due to the new variant
(1). Accordingly, molecular evidence is consistent with a potentially altered infectiousness
phenotype for this variant.
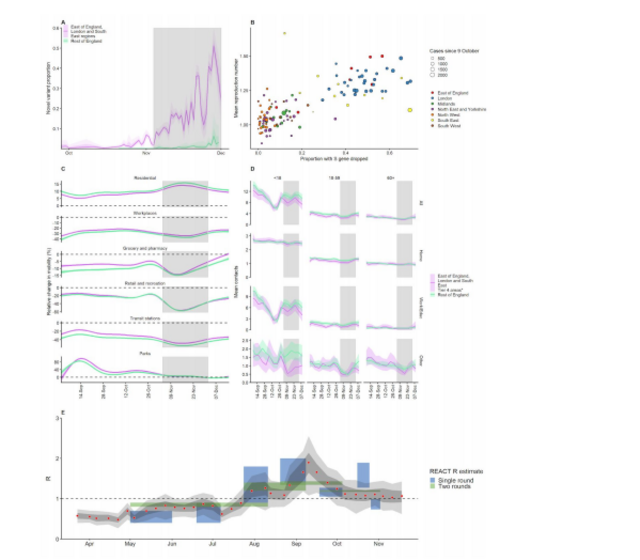
Fig. 1. (A) Proportion of VOC 202012/01 in South East, East of England, and London NHS
England regions versus the rest of England from 28 September – 1 December 2020 (mean and
95% CI). Grey shaded areas (panels A, C, D) reflect the period of time when England was in a
second national lockdown. We cut off the data after 1 December 2020 due to a substantial
decrease in representativeness after this time (Fig. S4). (B) Proportion of S gene drop-outs (5 –
11 December) versus mean reproduction number (27 November – 4 December) by local
authority in England. The one-week lag accounts for delays from infection to test. (C)
Percentage change (95% CI) in Google Mobility indices relative to baseline over time and (D)
setting-specific mean contacts (95% CI) from the CoMix study (9) over time and by age for local
authorities that went into Tier 4 compared to the rest of England. Educ = education setting.
*Some local authorities that were within the South East, East of England, and London NHS
England regions did not go into Tier 4 and were therefore included in the rest of England for
panels C and D. (E) Estimates of R0 (50% and 95% CI) from CoMix social contact survey (9)
compared to Rt estimates from REACT-1 SARS-CoV-2 prevalence survey (13) for England. Rt
estimates based on single and aggregated REACT-1 survey rounds are shown.
The proportion of COVID-19 cases caused by VOC 202012/01 is growing rapidly in the South
East, East of England and London regions (Fig. 1A) and is associated with an increase in the
estimated reproduction number Rt (Fig. 1B). Social contacts and mobility data suggest that the
rise in relative prevalence of VOC 202012/01 within England is unlikely to be caused by founder
effects: that is, if certain regions had higher levels of transmission as a result of more social
interactions, genetic variants that were more prevalent within these regions could become
more common overall. However, we did not find evidence of differences in social interactions
between regions of high and low VOC 202012/01 prevalence, as measured by Google mobility
(8) and social contact survey data (9) (Fig. 1C, D), despite the fact that changes in measured
contact patterns appear to closely correlate with changes in the reproduction number inferred
from community infection prevalence (Fig 1E) and that regionally-differentiated mobility data
have previously informed accurate predictions for COVID-19 dynamics in England (10). This
apparent decoupling between social contacts and transmission during November and
December could therefore indicate a change in characteristics of the virus.
To evaluate this possibility, we extended an age- and regionally-structured mathematical model
of SARS-CoV-2 transmission (10, 11) to consider two co-circulating variants of SARS-CoV-2 (Fig.
S1). The model is fitted across 7 NHS England regions to observed hospital admissions, hospital
and ICU bed occupancy, deaths within 28 days of a positive SARS-CoV-2 test, PCR prevalence,
seroprevalence, and, specifically for the East of England, London and South East NHS England
regions, to the relative frequency of VOC 202012/01 identified by genomic sequencing
surveillance from the COVID-19 Genomics UK Consortium (12). We use this model to capture
the emergence of VOC 202012/01 and consider its potential impact.
VOC 202012/01 is defined by 17 mutations (14 non-synonymous mutations and 3 deletions),
among which eight are located in the spike protein. At least three mutations have a potential
biological significance. Mutation N501Y is one of the key contact residues in the receptor
binding domain and has been shown to enhance binding affinity to human ACE2 (2, 3). The
function of mutation P681H is unclear, but it is located immediately adjacent to the furin
cleavage site in spike, a known region of importance for infection and transmission (4, 5). The
deletion of two amino acids at positions 69-70 in spike has arisen in multiple independent
circulating lineages of SARS-CoV-2, is linked to immune escape in immunocompromised
patients and enhances viral infectivity in vitro (6, 7). This deletion is also responsible for certain
commercial diagnostic assays failing to detect the spike glycoprotein gene (S gene drop-out),
with genomic data confirming these S gene target failures are primarily due to the new variant
(1). Accordingly, molecular evidence is consistent with a potentially altered infectiousness
phenotype for this variant.

Fig. 1. (A) Proportion of VOC 202012/01 in South East, East of England, and London NHS
England regions versus the rest of England from 28 September – 1 December 2020 (mean and
95% CI). Grey shaded areas (panels A, C, D) reflect the period of time when England was in a
second national lockdown. We cut off the data after 1 December 2020 due to a substantial
decrease in representativeness after this time (Fig. S4). (B) Proportion of S gene drop-outs (5 –
11 December) versus mean reproduction number (27 November – 4 December) by local
authority in England. The one-week lag accounts for delays from infection to test. (C)
Percentage change (95% CI) in Google Mobility indices relative to baseline over time and (D)
setting-specific mean contacts (95% CI) from the CoMix study (9) over time and by age for local
authorities that went into Tier 4 compared to the rest of England. Educ = education setting.
*Some local authorities that were within the South East, East of England, and London NHS
England regions did not go into Tier 4 and were therefore included in the rest of England for
panels C and D. (E) Estimates of R0 (50% and 95% CI) from CoMix social contact survey (9)
compared to Rt estimates from REACT-1 SARS-CoV-2 prevalence survey (13) for England. Rt
estimates based on single and aggregated REACT-1 survey rounds are shown.
The proportion of COVID-19 cases caused by VOC 202012/01 is growing rapidly in the South
East, East of England and London regions (Fig. 1A) and is associated with an increase in the
estimated reproduction number Rt (Fig. 1B). Social contacts and mobility data suggest that the
rise in relative prevalence of VOC 202012/01 within England is unlikely to be caused by founder
effects: that is, if certain regions had higher levels of transmission as a result of more social
interactions, genetic variants that were more prevalent within these regions could become
more common overall. However, we did not find evidence of differences in social interactions
between regions of high and low VOC 202012/01 prevalence, as measured by Google mobility
(8) and social contact survey data (9) (Fig. 1C, D), despite the fact that changes in measured
contact patterns appear to closely correlate with changes in the reproduction number inferred
from community infection prevalence (Fig 1E) and that regionally-differentiated mobility data
have previously informed accurate predictions for COVID-19 dynamics in England (10). This
apparent decoupling between social contacts and transmission during November and
December could therefore indicate a change in characteristics of the virus.
To evaluate this possibility, we extended an age- and regionally-structured mathematical model
of SARS-CoV-2 transmission (10, 11) to consider two co-circulating variants of SARS-CoV-2 (Fig.
S1). The model is fitted across 7 NHS England regions to observed hospital admissions, hospital
and ICU bed occupancy, deaths within 28 days of a positive SARS-CoV-2 test, PCR prevalence,
seroprevalence, and, specifically for the East of England, London and South East NHS England
regions, to the relative frequency of VOC 202012/01 identified by genomic sequencing
surveillance from the COVID-19 Genomics UK Consortium (12). We use this model to capture
the emergence of VOC 202012/01 and consider its potential impact.