|
|
Post by Admin on Feb 15, 2021 2:53:35 GMT
 Darwin’s theory of evolution should be expanded to include consideration of a DNA stability “energy code” – so-called “molecular Darwinism” – to further account for the long-term survival of species’ characteristics on Earth, according to Rutgers scientists. The iconic genetic code can be viewed as an “energy code” that evolved by following the laws of thermodynamics (flow of energy), causing its evolution to culminate in a nearly singular code for all living species, according to the Rutgers co-authored study in the journal Quarterly Reviews of Biophysics. “These revelations matter because they provide entirely new ways of analyzing the human genome and the genome of any living species, the blueprints of life,” said senior author Kenneth J. Breslauer, Linus C. Pauling Distinguished University Professor in the Department of Chemistry and Chemical Biology in the School of Arts and Sciences at Rutgers University–New Brunswick. He is also affiliated with the Rutgers Cancer Institute of New Jersey. “The origins of the evolution of the DNA genetic code and the evolution of all living species are embedded in the different energy profiles of their molecular DNA blueprints. Under the influence of the laws of thermodynamics, this energy code evolved, out of an astronomical number of alternative possibilities, into a nearly singular code across all living species.” Scientists investigated this so-called “universal enigma,” probing the origins of the astounding observation that the genetic code evolved into a nearly uniform blueprint that arose from trillions of possibilities. The scientists expanded the underpinnings of the landmark “survival of the fittest” Darwinian evolutionary theory to include “molecular Darwinism.” Darwin’s revolutionary theory is based on the generational persistence of a species’ physical features that allow it to survive in a given environment through “natural selection.” Molecular Darwinism refers to physical characteristics that persist through generations because the regions of the molecular DNA that code for those traits are unusually stable. Different DNA regions can exhibit differential energy signatures that may favor physical structures in organisms that enable specific biological functions, Breslauer said. Next steps include recasting and mapping the human genome chemical sequence into an “energy genome,” so DNA regions with different energy stabilities can be correlated with physical structures and biological functions. That would enable better selection of DNA targets for molecular-based therapeutics. Energy mapping of the genetic code and genomic domains: implications for code evolution and molecular Darwinism Published online by Cambridge University Press: 04 November 2020 Abstract When the iconic DNA genetic code is expressed in terms of energy differentials, one observes that information embedded in chemical sequences, including some biological outcomes, correlate with distinctive free energy profiles. Specifically, we find correlations between codon usage and codon free energy, suggestive of a thermodynamic selection for codon usage. We also find correlations between what are considered ancient amino acids and high codon free energy values. Such correlations may be reflective of the sequence-based genetic code fundamentally mapping as an energy code. In such a perspective, one can envision the genetic code as composed of interlocking thermodynamic cycles that allow codons to ‘evolve’ from each other through a series of sequential transitions and transversions, which are influenced by an energy landscape modulated by both thermodynamic and kinetic factors. As such, early evolution of the genetic code may have been driven, in part, by differential energetics, as opposed exclusively by the functionality of any gene product. In such a scenario, evolutionary pressures can, in part, derive from the optimization of biophysical properties (e.g. relative stabilities and relative rates), in addition to the classic perspective of being driven by a phenotypical adaptive advantage (natural selection). Such differential energy mapping of the genetic code, as well as larger genomic domains, may reflect an energetically resolved and evolved genomic landscape, consistent with a type of differential, energy-driven ‘molecular Darwinism’. It should not be surprising that evolution of the code was influenced by differential energetics, as thermodynamics is the most general and universal branch of science that operates over all time and length scales. Reference: “Energy mapping of the genetic code and genomic domains: implications for code evolution and molecular Darwinism” by Horst H. Klump, Jens Völker and Kenneth J. Breslauer, 4 November 2020, Quarterly Reviews of Biophysics. DOI: 10.1017/S0033583520000098 |
|
|
|
Post by Admin on Feb 15, 2021 4:48:20 GMT
Introduction
Once the genetic code was deciphered, it was quickly recognized that the code matrix was decidedly nonrandom. Elucidation of the underlying causes of this surprising regularity has been described as ‘The Universal Enigma’. Thought leaders such as Francis Crick, Manfred Eigen, Ed Trifonov, and others (see reviews/overviews and references cited therein) (Crick, 1968; Eigen and Winkler-Oswatitsch, 1992; Trifonov, 2000, 2004; Koonin and Novozhilov, 2009), proposed fundamental, physio-chemical frameworks to explain the origin and evolution of the code, including error minimization (Freeland et al., 2003; Novozhilov and Koonin, 2009), stereochemical (Yarus et al., 2005; Polyansky and Zagrovic, 2013; de Ruiter and Zagrovic, 2015), and coevolution theories (Di Giulio, 2004; Wong, 2005). They sought to explain the near singularity of the code across most living organisms, out of 1084 alternative possibilities (Koonin and Novozhilov, 2009). To this end, a thermodynamic perspective/framework helps rationalize the origins and evolution of the genomic energy landscape, in which differential energy profiles correlate with differential biological outcomes.
Biophysical origins and evolution of the genetic code
Within the context of an energy-based perspective for the origin and evolution of the genetic code, one can hypothesize that the original ancestral codons were comprised of a family of ‘prebiotic’ duplexes of sufficient stability to avoid dissociation from their antiparallel, complementary codons. One might reasonably envision that such codon couplets would code for the most ancient amino acids (Miller, 1953; Miller and Urey, 1959; Miller et al., 1976; Trifonov and Bettecken, 1997; Trifonov, 2000, 2004) if given the proper translational machinery. This core of ‘prebiotic’ ‘codon duplexes’ could have produced the rest of the code through a sequential series of transition and transversion mutations, the order of which is controlled/regulated/influenced via families of interlocking energy cycles. In this way, one can converge to a nearly singular genetic code out of the potential of more than 1084 alternative code tables (Novozhilov et al., 2007; Koonin and Novozhilov, 2009). Such energy-driven convergence of an astronomical number of potential alternative ‘states/codes’ into a single, unique ensemble ‘state/code’ is reminiscent of the energy landscape funnel associated with the protein folding phenomenon (Dill and Chan, 1997). Alternatively, one could argue that evolution was ‘stuck with’ whatever code developed by chance in the primordial environment. From that code, sequential mutations may have worked to drive the code toward evolutionarily traceable changes. Thus, once organisms started using such codes to connect replication (and transcription) to translation, they evolved more rapidly toward a thermodynamically driven code. We prefer the former perspective of an early stage, energy-driven, nonrandom evolutionary shaping of the genetic code through a series of mutations within particularly stable, prebiotic ‘codon duplexes’, all controlled via families of interlocking energy cycles.
Toward a thermodynamic contribution to the genetic code and to molecular Darwinism
In the classic view of Darwinian evolution (Darwin, 1859), a phenotypical characteristic of a species that imparts a survival advantage in a given environment persists through generations. By contrast, the population of those species variants that lack such an advantageous characteristic disappears over time. This ‘survival of the fittest’ phenomenon is what gives rise to what Darwin referred to as ‘natural selection’.
A necessary correlate to such a view of evolution is that the selective survival of the phenotypically advantaged form of a species yields enrichment within the species of genotypic signatures that correspond to production of (coding for) the advantageous phenotypical characteristics. Within such a framework, one could envision the generational persistence of certain species' characteristics that not only provide selective advantage for survival within a given environment, but which also are associated with particularly stable genetic signatures (codes) that resist alterations and thus enable generational persistence. As such, evolution may result from a mixture of contributions from classic, natural selection, Darwinian theory, as well as from what can be called ‘molecular Darwinism’ (Eigen, 1976), or ‘Watson–Crick Darwinism’. In the latter context, some phenotypical characteristics might persist since they are coded for by more stable domains in the genome, even if such characteristics do not maximize species survival. In the classic evolutionary context, characteristics that provide a survival advantage will generationally persist, with no consideration for the stability of the genotypical signature. The net outcome of species evolution may well reflect contributions from both phenotypical (classic Darwinism) and genotypical (molecular Darwinism) influences.
To gain insight into such a DNA-based perspective, one can map the iconic chemical genetic code in terms of an energy code. Perhaps early evolution of the genetic code can be viewed in terms of the differential stabilities of codon/complementary codon couplets that form antiparallel trimeric duplexes, as opposed to exclusively in terms of the functionality of any gene product.
|
|
|
|
Post by Admin on Feb 15, 2021 19:22:31 GMT
The genetic code as an energy code In Table 1, the classic Crick genetic code matrix (Crick, 1968) is elaborated by also listing the stabilizing free energy values for each fully paired codon/complementary codon antiparallel trimeric duplex. These stability parameters were calculated using calorimetrically-derived free energy values previously reported by our labs (Breslauer et al., 1986). The decision to map the genetic code in terms of codon/antiparallel complementary codon energetics is justified on multiple physio-chemical levels. First, trimeric duplexes formed from association of fully complementary, antiparallel, codon pairs, using up to four ‘letters’, reflect the minimum molecular information units required to code for the diversity of amino acids. Second, as demonstrated by Porschke and Eigen (1971), codon/complementary codon, antiparallel, trimeric duplexes possess the minimum stability required to form a complex of sufficient strength such that the associated species do not spontaneously dissociate into their constituent single-stranded components; a circumstance that makes them susceptible to degradation and thereby loss of coding information. On the other hand, making these interactions still more stable, as in a tetramer or higher chain length code, may well work against optimal rates of translation (see Greive and von Hippel, 2005). In other words, in the current context, increased thermodynamic stability might work against the use of more stable (e.g. longer) codon–complementary codon interactions in ‘molecular evolution’. Third, it has been suggested that in primitive primordial molecular machines, translocation events in steps of 3 monomer units (i.e. the size of the codon) correspond to local energy minima that are favored over other translocation step sizes (Aldana et al., 1998; Martinez-Mekler et al., 1999; Aldana-González et al., 2003). It is within this context, that stability data for each antiparallel, codon/complementary codon interaction are shown in Table 1. In this format, one can explore relationships between the differential trimeric duplex stabilities formed from codons and their corresponding complementary codons and biological observations/outcomes. 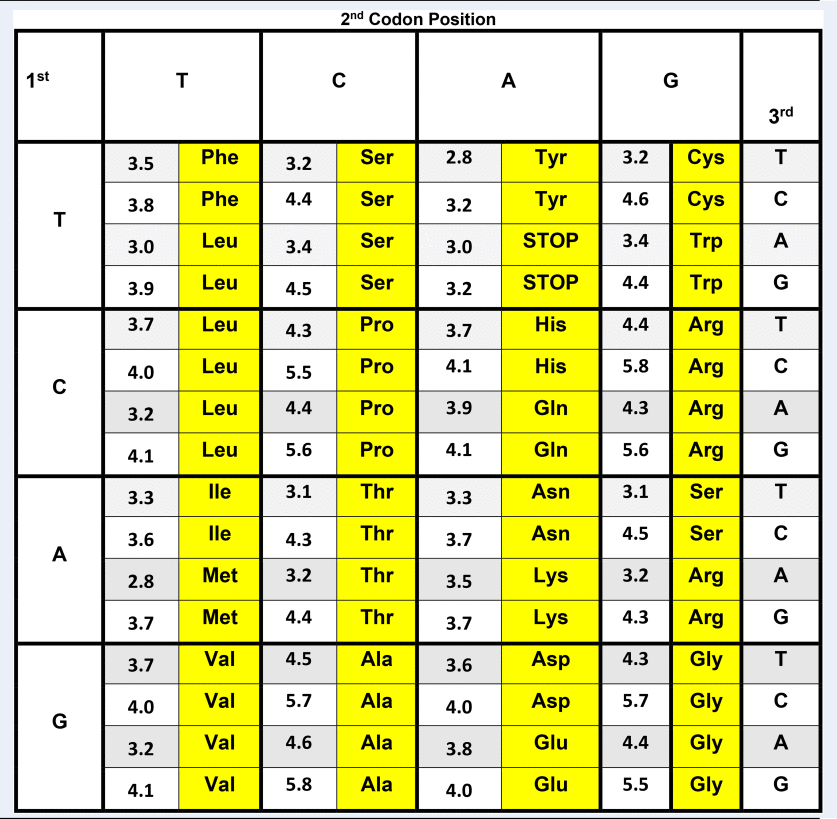 Table 1. Genetic code matrix annotated with trimeric duplex stabilities formed between codons and their corresponding, antiparallel complementary codons Genetic code matrix for human mitochondrial DNA annotated with stabilizing free energy values for the trimeric duplexes formed by antiparallel, complementary codons. The free energy values were calculated based on the calorimetrically determined nearest-neighbor dataset reported by Breslauer et al. (1986). To reduce the impact of end effects, the normalized, weighted average of the duplex free energies were calculated, with each terminus hypothetically ‘sealed’ by all possible base pairs/stacks. The net impact of this end effect correction is to dampen the variability between codons, and to reduce the dominance of the central base in the codon trimer caused by it being the only base with two neighbors. Significantly, however, aside from modest compression of the stability range, the rank order of the differential codon duplex stabilities listed, as well as the general correlations noted here remain unaltered, even when no end effect ‘correction’ is applied. Compilations of the codon free energy data employing any of the other commonly used nearest-neighbor databases results in some numerical and rank order differences, reflective of subtle differences in the numerical values assigned to nearest-neighbors in these different databases (e.g. Delcourt and Blake, 1991; Doktycz et al., 1992; SantaLucia et al., 1996; SantaLucia, 1998). That said, the nearest-neighbor data of Ritort and Bustamante, derived from force stretching experiments, exhibit the most concurrence with the relative trends reported here; specifically in terms of the free energy rank ordering of the codons, as well as the codon usage patterns (Huguet et al., 2010). Given Ritort's subsequent assessment of the differential impact of magnesium ion on the nearest-neighbor data, future studies should also consider measurements in a variety of counterion/cation environments (Huguet et al., 2017). The genetic code matrix shown in Table 1 corresponds to the human mitochondria code (Anderson et al., 1981; Breitenberger and RajBhandary, 1985). Note that the smallest interaction free energy (least stable), ΔG, of 2.8 kcal per mole triplet is between the alternating sequences ATA and TAT, and the highest (most stable) ΔG value is 5.8 kcal per mole triplet for the duplex formed by the alternating sequences GCG and CGC. |
|
|
|
Post by Admin on Feb 15, 2021 22:38:15 GMT
Energy dispersion of the codon/complementary codon, trimeric duplexes The 64 possible DNA triplet codons collectively constitute the minimal ‘words’ of the genetic code. When evaluated as bound to their fully complementary, antiparallel codons, they form 32 trimeric duplexes, each with a calculated stability. As summarized in Scheme 1 heat map, this process reveals a broad stability dispersion of the trimeric duplexes formed by the codon/complementary antiparallel codon. Such significant energy dispersion makes these differential energy profiles information-rich. 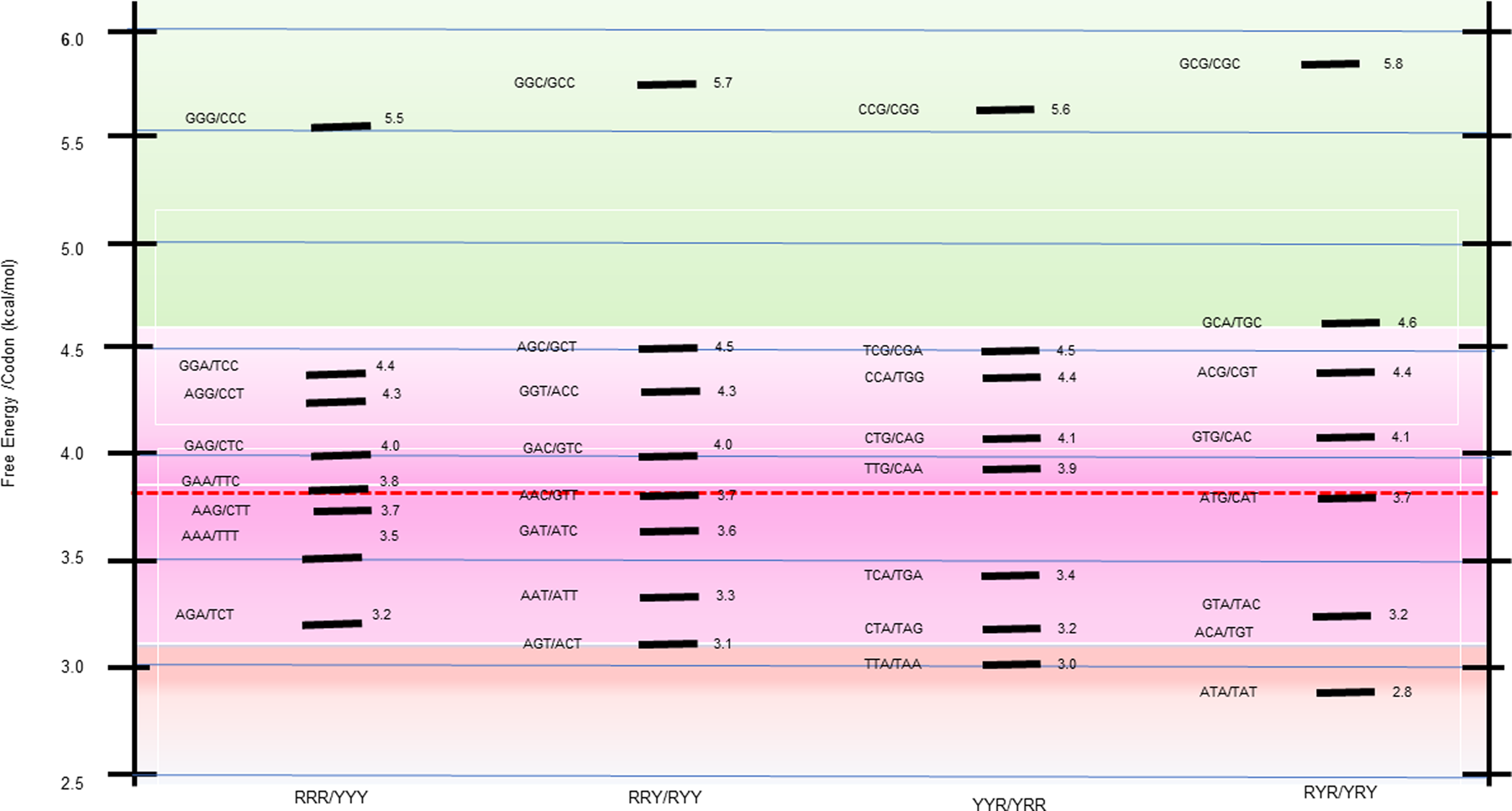 Scheme 1. Free energy distribution spectrum for the 32 trimeric duplexes formed by all 64 complementary codons. The stability distribution is color coded as a ‘heat map’, with the GC-rich most stable family (highest free energy of trimeric duplex formation) highlighted toward the top of the scheme in light green; the next most stable family is highlighted in light purple; and the less stable duplexes relative to the mean are highlighted in light red. The energy spectrum is formatted within four columns that reflect the purine (R)/pyrimidine (Y) sequence patterns designated at the bottom of the scheme. Correlations between codon usage frequencies and the stabilities of the duplexes formed by each codon and its antiparallel complementary codon Comparisons between the codon/complementary codon free energy data and the whole genome codon usage frequencies reported by Futcher and coworkers (Gardin et al., 2014) for yeast Saccharomyces cerevisiae reveal an intriguing coupling of properties. By combining the differential stabilities and Futcher's datasets, one observes a near linear correlation between the frequency with which a given codon is used for a particular amino acid and the corresponding codon/complementary codon free energy. To be specific, save for isoleucine, of the 17 out of 20 amino acids for which Futcher reports sufficient data density, we observe that codons with lower free energies (less stable) are used more frequently than codons for the same amino acid with higher free energies (more stable). This coupling of a fundamental physio-chemical property with the outcome of a complex biological process is illustrated for several amino acids in the plots shown in Fig. 1. Such empirical correlations reinforce reductionists' efforts to rationalize complex biology in terms of fundamental chemical principles. 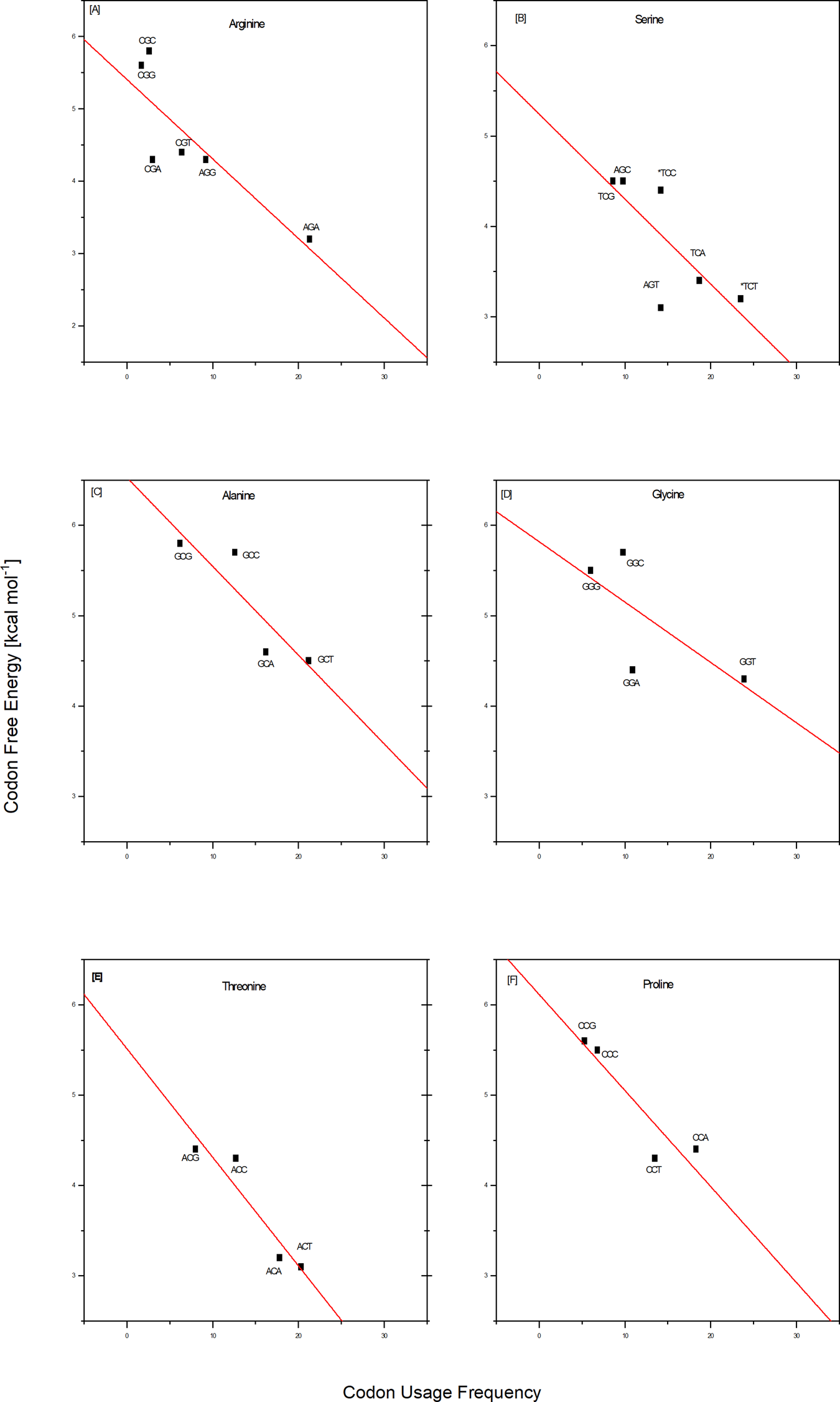 Fig. 1. Empirical correlations between whole genome codon usage frequencies in S. cerevisiae taken from the work of Futcher and coworkers (Gardin et al., 2014) and the corresponding codon/complementary codon free energies of this study. Each red line represents a best fit to the equation for a straight line of these two independently derived data sets. The result shown here are for two of the three amino acids encoded by six codons, and for four of the five amino acids encoded by four codons. This selection corresponds to that subset of the amino acids judged most ancient, based on a meta-analysis reported by Trifonov (2000, 2004). With the exception of isoleucine, and the insufficient data density for methionine and tryptophan, all of the amino acids encoded by only two codons also show a preference for higher codon usage frequency that correlates with lower codon free energy. For a thermodynamic argument, one strictly should use a log scale plot for the usage frequency. However, over the small data range assessed here, we have confirmed that one cannot distinguish between linear and log linear, with the log plot simply compressing the data. Based on this coupling, one might speculate, that the degeneracy associated with the use of multiple codons of differential stabilities to code for the same amino acid reflects a form of thermodynamic selection; one in which codon energetics is more determinatory of usage frequency than a codon's chemical syntax alone. It will be instructive to probe the extent to which this empirical correlation between codon stabilities and codon usage frequencies is universal across all organisms and genes, as well as to define the biological implications. For now, this correlation provides an example of insights that can be gained by parsing the iconic genetic code in terms of energy differentials. Conversely, one also might posit that the usages of codons reflect biologically relevant features of those DNA sequences containing a statistical overabundance of energetically favorable or unfavorable codons. The altered energy profiles of such DNA sequences relative to a statistically expected distribution of codons/energies may reflect the existence of biological constraints that do not apply to an average sequence. In other words, codon usage that deviates from the average expected distribution (either positive or negative) may reflect altered biological constraints. |
|
|
|
Post by Admin on Feb 16, 2021 1:09:50 GMT
‘Evolving’ all codons from each other through sequential series of transitions and transversions To illustrate the cycles associated with interconverting codons for the entire genetic code, the 64 codons presented in Table 1 and Scheme 1 can be arranged into a total of eight ‘octets’. Each octet is composed of codons located at one of the eight apices of a cube. Each cube corresponds to one of the purine (R)/pyrimidine (Y) sequence patterns designated in Scheme 1. The resultant eight octet cubes are then positioned at the corners of a master scaffolding cube to create a ‘hypercube’ as shown in Fig. 2. This hypercube illustrates the full cascade of all of codon interconversions via sequential site changes over all codon sequence space. 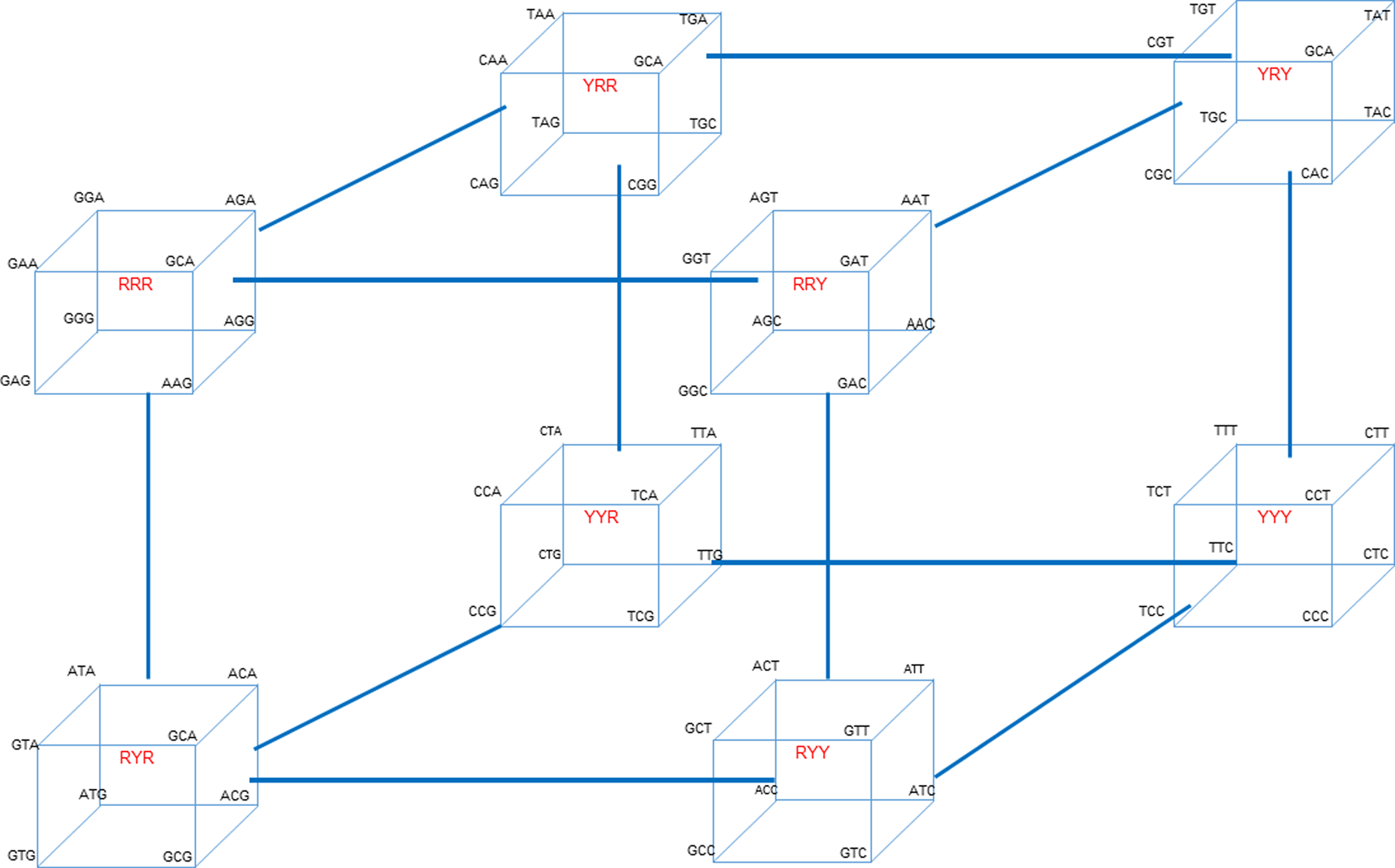 Fig. 2. The hypercube of all eight cube octet sequence classes (shown in red) located at each apex of the hypercube, illustrating the interconnectedness of the cycles associated with the full cascade of codon interconversions via sequential site changes. Transition mutations occur within a cube, while transversion mutations link one cube to another. Note that the eight octet cubes shown in the hypercube of Fig. 2 are inter-cube related by codon transversion mutations, whereas codons within a given octet cube are intra-cube related by transition mutations. These stepwise interconversions create interlocking cycles that allow one to traverse/”evolve” the entire genetic code. This stepwise generation/”evolution” of all 64 codons via sequential transition and transversion mutations, starting from any codon, may reflect a differential, energy-modulated evolution of the genetic code. As such, it might correspond to a biophysical basis for what Eigen referred to as ‘molecular Darwinism’. At the heart of molecular Darwinism is the generation of genetic variation. Genetic variants can result from DNA sequence alterations at local levels; from rearrangement of DNA segments intragenomically; and by gene transfer of foreign DNA. The hypercube illustrates how local sequence changes can interconvert all 64 codon variants via sequential cascades of transition and transversion steps that Table 1 shows exhibit differential free energies, thereby creating codon variants; a characteristic at the heart of molecular Darwinism. |
|





