Post by Admin on Mar 30, 2021 22:17:03 GMT
S5. Y chromosome analysis
A list of informative SNP sites was obtained from AMY-tree v2.1 [62], using a snapshot of
PhylotreeY [63]. We excluded all non-SNP sites, transition sites (to avoid deamination
damage), and A/T and G/C SNPs (to avoid strand misidentification). This left us with 550
sites after filtering. All these sites were checked in ATP12-1420 and ATP2 for bases with a
minimum quality of 30 and a minimum mapping quality of 30. We also excluded sites
showing two different alleles since those are likely sequencing errors or contamination, this
excluded 1 SNP for ATP12-1420 and 14 SNPs for ATP2.
S5.1. ATP2
ATP2 displayed the derived allele for nine Y chromosome markers (Table S5); with all of the
markers providing phylogenetic support for ATP2 belonging to haplogroup H2. These
markers included: L985, L1013 and L1053 (A1); M235, P159 and P187 (F); L279, L281 and
P96 (H2). Previously labeled haplogroup F3, H2 was recently redefined on the basis of an
overlap between the datasets of [64] and [65] [63]. It was found that the two haplogroups, HM69 and F3-M282, shared a root defined by the marker M3035. While only a few H2 individuals have ever been found, the haplogroup appears to have a west Eurasian distribution; with a low level Middle Eastern presence in modern-day Iran, Turkey, Bahrain, Kuwait and Qatar (Family Tree DNA, [66]), as well as minor occurrences in modern-day England, France, Sardinia, Sweden and the Netherlands (Family Tree DNA, [64,65,67]). H2 also seems to occur at low frequencies in Neolithic samples [68].
S5.2. ATP12-1420
ATP12-1420 displayed the derived allele for five Y chromosome markers (Table S6); with all
of the markers providing phylogenetic support for ATP12-1420 belonging to haplogroup
I2a2a. These markers included: L1053 (A1); P124 (IJ); L460 (I2a); L34 and P221 (I2a2a).
While the almost European-specific haplogroup I arose approximately 20000 to 25000 years
ago [67,69], haplogroup I2a2a may have diverged as a subclade, around 15000 years ago
[69], possibly during the recolonization of Europe following the Last Glacial Maximum (LGM).
Unlike the more common subclades of I1 and I2a1, haplogroup I2a2a appears at relatively
low frequencies across much of Europe. Its highest levels (10-12%) are found in modern-day
Germany and the Netherlands, with frequencies of around 5%, notably occurring in parts of
modern-day France as well as Mordvin in the Volga region of central Eastern Europe [69].
Other members of haplogroup I have been discovered previously in ancient individuals; e.g.
I* in Mesolithic Scandinavians [19], I1 in Hungary [70], I2a in Neolithic individuals from
Hungary [71] and France [46], I2a1 in Neolithic Croatia [70] and a late hunter-gatherer from
Sweden [20], and I2a1b in Mesolithic individuals from Luxembourg and Sweden [19].
Other haplogroups found among ancient specimens include C* in Upper Paleolithic Russia
[72] and Mesolithic Spain [73], C6 in Neolithic Hungary [71], E1b1b1 in Neolithic Spain [45],
F* in Neolithic Germany [44] and Neolithic Hungary [70], G2 in Neolithic Hungary [70], G2a
in Neolithic France [46], Neolithic Germany [44], Neolithic Hungary [70], Chalcolithic Italy
[74], and Neolithic Spain [45], J2a1 in Bronze Age Hungary [71], K (xLT) in Upper Paleolithic
western Siberia [75], N in Iron Age Hungary [71], R* in south central Upper Paleolithic
Siberia [76], R1a in Neolithic Germany [77], and Neolithic R1b in Germany [78].
S6. Phenotypic/selection analysis
In order to get an idea about the appearance of our individuals and their allelic status at sites
identified to evolve under selection in modern-day Europeans, we checked reads covering
five specific SNPs and compare them to a Mesolithic Iberian [73]. We only included reads
with a mapping score of at least 30 and sites with a minimum base quality of 30 into the
analysis. A more comprehensive study of selection targets and phenotypes would require a
larger sample size and higher coverage per individual especially since most of these
phenotypes are highly polygenic. Furthermore, we note that the potential post-mortem
damage at transition sites might affect the conclusion of such surveys. Allelic states are
shown in Table S7.
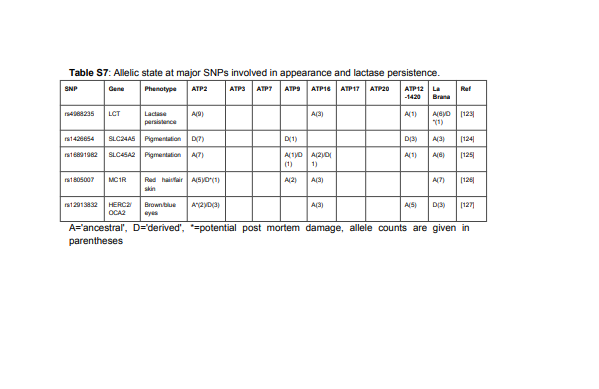
As shown before [99], the inhabitants of the El Portalon cave were probably all lactose
intolerant in adulthood. This suggests a much later spread of this variant that has been the
target of adaptation to a milk-rich diet in modern-day northwestern Europeans which also
occurs at reasonably high frequencies in Northern Spain and Basques [100]. All sequences
for the SLC24A5 (rs1426654) variant showed the derived state in the El Portalon individuals,
which together with two derived variants at SLC45A2 (rs16891982), suggest that the
pigmentation of the Chalcolithic Iberians was lighter than the Mesolithic LaBrana1 individual
who carried the ancestral states at these major pigmentation loci [73].
rs1805007 in MC1R which is associated with red hair and light skin is ancestral in all but one
sequence (out of eleven in all El Portalon individuals) but that single derived base call might
also be due to post-mortem damage. rs12913832, a SNP that explains more than 56% of
the variation between blue and brown eyes [101], has been shown to be derived in the
Mesolithic LaBrana1 [73]. Two of the El Portalon individuals show only ancestral alleles at
this site whereas one individual shows both variants suggesting the individual is heterozygot
at the site. These observations suggest some eye color variation but also a tendency
towards brown eyes in the Chalcolithic Iberians.
To summarize, Chalcolithic Iberian farmers seem to be lactose intolerant as the Mesolithic
inhabitants of the Peninsula. However, their pigmentation was fairer and their eyes were
darker than in the hunter-gatherer LaBrana1.
A list of informative SNP sites was obtained from AMY-tree v2.1 [62], using a snapshot of
PhylotreeY [63]. We excluded all non-SNP sites, transition sites (to avoid deamination
damage), and A/T and G/C SNPs (to avoid strand misidentification). This left us with 550
sites after filtering. All these sites were checked in ATP12-1420 and ATP2 for bases with a
minimum quality of 30 and a minimum mapping quality of 30. We also excluded sites
showing two different alleles since those are likely sequencing errors or contamination, this
excluded 1 SNP for ATP12-1420 and 14 SNPs for ATP2.

S5.1. ATP2
ATP2 displayed the derived allele for nine Y chromosome markers (Table S5); with all of the
markers providing phylogenetic support for ATP2 belonging to haplogroup H2. These
markers included: L985, L1013 and L1053 (A1); M235, P159 and P187 (F); L279, L281 and
P96 (H2). Previously labeled haplogroup F3, H2 was recently redefined on the basis of an
overlap between the datasets of [64] and [65] [63]. It was found that the two haplogroups, HM69 and F3-M282, shared a root defined by the marker M3035. While only a few H2 individuals have ever been found, the haplogroup appears to have a west Eurasian distribution; with a low level Middle Eastern presence in modern-day Iran, Turkey, Bahrain, Kuwait and Qatar (Family Tree DNA, [66]), as well as minor occurrences in modern-day England, France, Sardinia, Sweden and the Netherlands (Family Tree DNA, [64,65,67]). H2 also seems to occur at low frequencies in Neolithic samples [68].
S5.2. ATP12-1420
ATP12-1420 displayed the derived allele for five Y chromosome markers (Table S6); with all
of the markers providing phylogenetic support for ATP12-1420 belonging to haplogroup
I2a2a. These markers included: L1053 (A1); P124 (IJ); L460 (I2a); L34 and P221 (I2a2a).
While the almost European-specific haplogroup I arose approximately 20000 to 25000 years
ago [67,69], haplogroup I2a2a may have diverged as a subclade, around 15000 years ago
[69], possibly during the recolonization of Europe following the Last Glacial Maximum (LGM).
Unlike the more common subclades of I1 and I2a1, haplogroup I2a2a appears at relatively
low frequencies across much of Europe. Its highest levels (10-12%) are found in modern-day
Germany and the Netherlands, with frequencies of around 5%, notably occurring in parts of
modern-day France as well as Mordvin in the Volga region of central Eastern Europe [69].
Other members of haplogroup I have been discovered previously in ancient individuals; e.g.
I* in Mesolithic Scandinavians [19], I1 in Hungary [70], I2a in Neolithic individuals from
Hungary [71] and France [46], I2a1 in Neolithic Croatia [70] and a late hunter-gatherer from
Sweden [20], and I2a1b in Mesolithic individuals from Luxembourg and Sweden [19].
Other haplogroups found among ancient specimens include C* in Upper Paleolithic Russia
[72] and Mesolithic Spain [73], C6 in Neolithic Hungary [71], E1b1b1 in Neolithic Spain [45],
F* in Neolithic Germany [44] and Neolithic Hungary [70], G2 in Neolithic Hungary [70], G2a
in Neolithic France [46], Neolithic Germany [44], Neolithic Hungary [70], Chalcolithic Italy
[74], and Neolithic Spain [45], J2a1 in Bronze Age Hungary [71], K (xLT) in Upper Paleolithic
western Siberia [75], N in Iron Age Hungary [71], R* in south central Upper Paleolithic
Siberia [76], R1a in Neolithic Germany [77], and Neolithic R1b in Germany [78].
S6. Phenotypic/selection analysis
In order to get an idea about the appearance of our individuals and their allelic status at sites
identified to evolve under selection in modern-day Europeans, we checked reads covering
five specific SNPs and compare them to a Mesolithic Iberian [73]. We only included reads
with a mapping score of at least 30 and sites with a minimum base quality of 30 into the
analysis. A more comprehensive study of selection targets and phenotypes would require a
larger sample size and higher coverage per individual especially since most of these
phenotypes are highly polygenic. Furthermore, we note that the potential post-mortem
damage at transition sites might affect the conclusion of such surveys. Allelic states are
shown in Table S7.

As shown before [99], the inhabitants of the El Portalon cave were probably all lactose
intolerant in adulthood. This suggests a much later spread of this variant that has been the
target of adaptation to a milk-rich diet in modern-day northwestern Europeans which also
occurs at reasonably high frequencies in Northern Spain and Basques [100]. All sequences
for the SLC24A5 (rs1426654) variant showed the derived state in the El Portalon individuals,
which together with two derived variants at SLC45A2 (rs16891982), suggest that the
pigmentation of the Chalcolithic Iberians was lighter than the Mesolithic LaBrana1 individual
who carried the ancestral states at these major pigmentation loci [73].
rs1805007 in MC1R which is associated with red hair and light skin is ancestral in all but one
sequence (out of eleven in all El Portalon individuals) but that single derived base call might
also be due to post-mortem damage. rs12913832, a SNP that explains more than 56% of
the variation between blue and brown eyes [101], has been shown to be derived in the
Mesolithic LaBrana1 [73]. Two of the El Portalon individuals show only ancestral alleles at
this site whereas one individual shows both variants suggesting the individual is heterozygot
at the site. These observations suggest some eye color variation but also a tendency
towards brown eyes in the Chalcolithic Iberians.
To summarize, Chalcolithic Iberian farmers seem to be lactose intolerant as the Mesolithic
inhabitants of the Peninsula. However, their pigmentation was fairer and their eyes were
darker than in the hunter-gatherer LaBrana1.
