|
|
Post by Admin on Feb 7, 2023 19:46:13 GMT
Early lineages of Asia
Though present-day Asians and Australasians are more closely related to each other than to present-day Europeans, genetic comparisons highlight deep separations between mainland East and Southeast Asians, island Southeast Asians, and Australasians. Philippine Negritos, Papuans, and aboriginal Australians share a close genetic relationship to each other relative to mainland East and Southeast Asians [11,15,59,75] and are collectively referred to here as belonging to an Australasian (AA) lineage (Box 2). In contrast, mainland East and Southeast Asians and other Pacific islanders (e.g., Austronesian speakers) are closely related to each other [9,15,16] and here denoted as belonging to an East and Southeast Asian (ESEA) lineage (Box 2). In South Asia, present-day populations are highly admixed, but recent sequencing of ancient DNA indicated the presence of a deeply diverged Asian ancestry only distantly related to populations associated with the AA or ESEA lineages [59]. This distinct South Asian ancestry, denoted as the Ancient Ancestral South Indian (AASI) lineage (Box 2), was only found in a small percentage of ancient and present-day South Asians. Present-day Onge from the Andamanese Islands are the best reference population to date, but Narasimhan et al. used qpGraph to show that the divergence between the AASI lineage and the ancestry found in present-day Onge was very deep [59]. Ancestry associated with the AASI lineage was found at low levels in almost all present-day Indian populations, particularly southern Indians [59,60], emphasizing the high impact of the AASI lineage in South Asia.
The AA, ESEA, and AASI lineages showed a closer genetic relationship to each other than lineages observed in present-day Europeans [59] and together represent the main branches of Asian-related ancestry sampled to date (Figure 1B). The relationship of the AA, ESEA, and AASI lineages to each other is not well resolved, in part due to high levels of admixture in present-day Asian and Australasian populations, either with archaic humans [26,27,32,37], populations carrying other non-African ancestries [44,51], or with each other [9]. Recently, Hajdinjak et al. used ancient DNA capture targeting a 3.8M SNP panel (Box 1) to sequence DNA for three 45,000-year-old individuals from Bacho Kiro Cave in Bulgaria [76]. They found that the Bacho Kiro individuals were genetically more similar to present-day Asians than to present-day Europeans in an outgroup f3-analysis (Box 1), extending the range in which ancestry associated with present-day Asian populations has been found as far west as Bulgaria.
|
|
|
|
Post by Admin on Feb 8, 2023 19:53:00 GMT
Differentiation and admixture in Asia 45,000–10,000 years ago
To date, no Upper Paleolithic individuals sampled have been associated with the AASI and AA lineages, though ADMIXTURE analyses (Box 1) on present-day populations associated with these lineages in South Asia and Australasia indicate the presence of complex structure and admixture [11]. Genetic data from ancient individuals have been informative about the ESEA lineage. All humans sampled to date from the Upper Paleolithic (45,000–10,000 years ago) in Asia were excavated in China, Mongolia, and Russia, in part because preservation of genetic material in human specimens is more likely in colder and drier regions [77]. In the next sections, I review how ancient DNA has helped to clarify genetic structure and admixture in populations associated with the ESEA lineage in the Upper Paleolithic.
Early Upper Paleolithic: 45,000 to 20,000 years ago
Three individuals from northern China [61,66,78] and northeastern Mongolia [65] dating to 40,000 to 33,000 years ago (Figure 1B) were sequenced using ancient DNA capture targeting a 1.2M (AR33K) or 2.2M (Tianyuan, Salkhit) SNP panel. Comparison with present-day populations using outgroup f3- and D-statistics showed that they were genetically most similar to present-day East Asians and Southeast Asians, thus showing that Tianyuan, Salkhit and AR33K are associated with the ESEA lineage, rather than the AA or AASI lineages [61,65,66,78]. In a test of genetic continuity, Tianyuan was shown to derive from a population distinct from the one that contributed to present-day East and Southeast Asians, indicating differentiation of the ESEA lineage in the Early Upper Paleolithic [66]. D-statistic analyses showed that AR33K and Salkhit were more closely related to Tianyuan than to present-day East and Southeast Asians [61,65]. Thus, by 40,000–33,000 years ago, one or more populations in northern China and Mongolia were genetically differentiated from the source population that contributed to present-day East and Southeast Asians. I denote the shared ancestry between Tianyuan, AR33K, and Salkhit individuals as Tianyuan ancestry (Box 2), which diverged prior to the shared common ancestor of present-day East and Southeast Asians.
No DNA has been retrieved from 40,000–20,000-year-old humans in southern regions of Asia, but sampling of 8,000–4,000-year-old hunter-gatherers [63] associated with the Hòabìnhian culture in Laos and Malaysia reveals another ancestry that must have existed in the Early Upper Paleolithic. Ancient DNA capture on the 1.2M SNP panel and assessment of relative genetic similarity using D-statistics showed that these Southeast Asian hunter-gatherers were most closely related to Tianyuan and present-day East and Southeast Asians [63]. However, D-statistic comparisons also showed that they were as genetically different from Tianyuan as they were from present-day East and Southeast Asians [63,68] and are denoted here as associated with Hòabìnhian ancestry (Box 2). Together, the genetic patterns described above show that the ESEA lineage differentiated into at least three distinct ancestries: Tianyuan ancestry which can be found 40,000-33,000 years ago in northern East Asia, ancestry found today across present-day populations of East Asia, Southeast Asia, and Siberia, but whose origins are unknown, and Hòabìnhian ancestry found 8,000-4,000 years ago in Southeast Asia, but whose origins in the Upper Paleolithic are unknown.
DNA sampling of ancient humans in western Siberia, the Near East, and Central Asia showed that populations existed in the past in the western regions of Asia that are not directly represented today. For instance, Fu et al. [70] retrieved and sequenced 1.86 gigabases of the autosomal genome from a femur found at Ust’-Ishim, a settlement in western Siberia (Figure 1B). Using D-statistics, they showed that the Ust’-Ishim individual was similarly related to Upper Paleolithic European hunter-gatherers and present-day East Asians [66,70,79]. After targeting 1.2M SNPs [42,43] in 12,000–7,000-year-old individuals from the Levant and Iran, Lazaridis et al. found that Ust’-Ishim and present-day East Asians shared more alleles with each other than with ancient individuals from the Levant and Iran [71], which indicates that they diverged basally from other non-Africans. In a principal component analysis (PCA, Box 1), ancient individuals from the Levant and Iran clustered at opposite ends of a Near Eastern cline, indicating high genetic differentiation between these two populations [59,71]. Ust’-Ishim and the 12,000–7,000-year-old individuals from the Levant and Iran represent deeply diverged non-African ancestries (denoted here as Basal ancestries, Box 2) that are partially or not represented in present-day populations.
In northeastern Siberia, Upper Paleolithic populations were not associated with the ESEA, AA, or AASI lineages. Two individuals (Yana 1 and 2) were sampled in northeastern Siberia at the Yana Rhinoceros Horn site off the Yana River dating to 31,000 years ago [69], and another three (Mal’ta 1 and Afontova Gora 2/3), dating to 24,000–17,000 years ago, were sampled in south central Siberia [51] (Figure 1B). Comparison to ancient and present-day populations in Europe and Asia using f4-statistics (non-normalized version of the D-statistic, Box 1) showed that these individuals were genetically more similar to European hunter-gatherers than to Tianyuan or present-day East and Southeast Asians [51,69], which suggests that Upper Paleolithic Siberians were split at an early point from a population that contributed to ancient European hunter-gatherers [79]. The ancestry represented by the Yana individuals is denoted as Ancient Northern Siberian (ANS) ancestry (Box 2, Figure 1B), based on Sikora et al. [69]. Mal’ta1 and Afontova Gora 2/3 [51], often referred to as representing Ancestral North Eurasian ancestry, is closely related to ANS ancestry found in Siberia 31,000 years ago and is suggested to have originated from ANS ancestry [69]. The persistence of ANS ancestry in Siberia from 31,000-17,000 years ago shows that ANS ancestry played an essential role in the early history of human migration across northern Asia. These patterns in Siberia and East and Southeast Asia show that 40,000-20,000 years ago in the Early Upper Paleolithic, eastern Eurasia was replete with populations that were genetically distinct from one another. Both ANS and ESEA ancestries were instrumental in shaping human genetic history.
Interaction between populations associated with Tianyuan ancestry and ANS ancestry played a key role in shaping the distribution of humans in East Asia and Siberia. For instance, a symmetry test using D-statistics showed that Salkhit shares more genetic similarity with the ANS-related Yana individuals than either Tianyuan or AR33K [61,65], which suggests that populations in Mongolia associated with Tianyuan ancestry likely interacted more with populations associated with ANS ancestry than those in the Amur River region and the North China Plain. D-statistic analyses additionally show that Salkhit and Tianyuan share more alleles with a 35,000-year-old European hunter-gatherer from Goyet Cave, Belgium (GoyetQ116-1) than with present-day East and Southeast Asians [61,65,66]. Yang et al. [66] could not fully explain the population dynamics leading to a genetic connection between western European and northern Asian individuals, but a similar affinity to GoyetQ116-1 recently observed in the 45,000-year-old Bacho Kiro individuals from Bulgaria using qpGraph showed that the extended connections were potentially facilitated by Asian-related populations that once lived in Europe [76]. The interconnections among Early Upper Paleolithic European and Asian populations show that these Early Upper Paleolithic hunter-gatherers were not totally isolated--admixture and migration played a fundamental role in their ancestral makeup.
|
|
|
|
Post by Admin on Feb 13, 2023 17:42:02 GMT
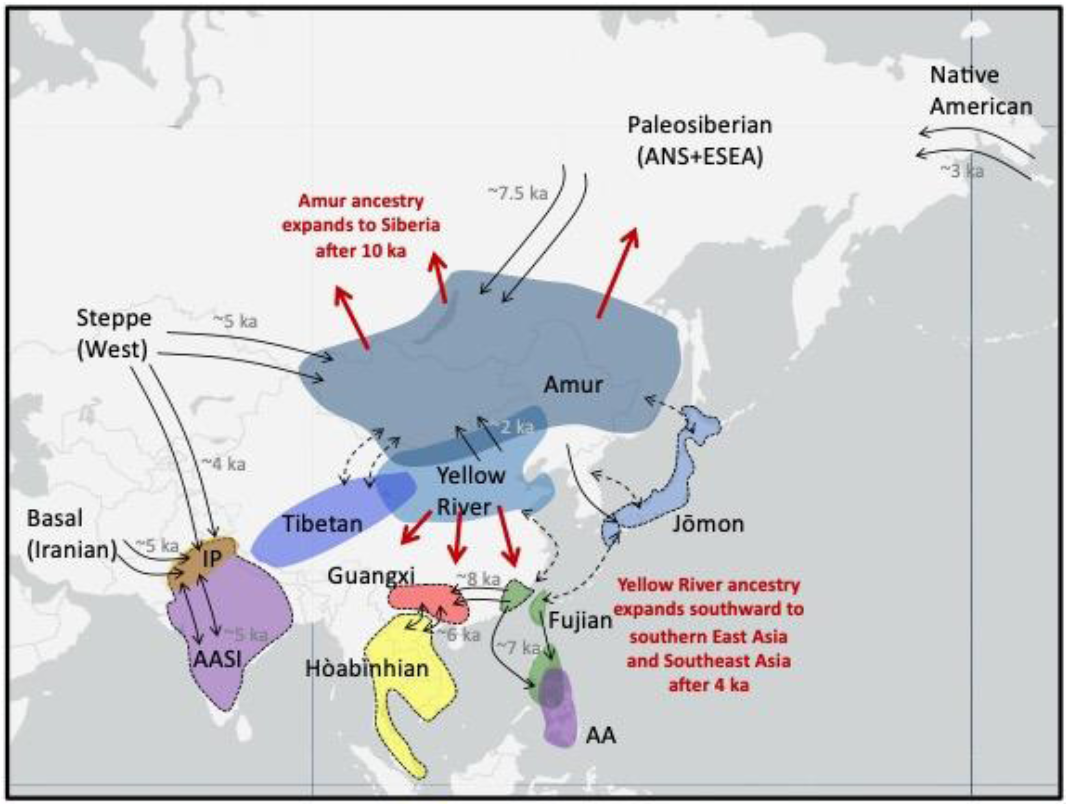 Figure 2 Schematic depicting major ancestries in Asia from the last 15,000 years. Black labels indicate ancestries and lineages described in the text and in Box 2. Highlighted regions show where ancient individuals associated with the labeled ancestry have been sampled. Excepting Indus Periphery (IP) ancestry, ancestries not associated with a highlighted region do not derive complete from the AA, AASI, or ESEA lineages. Purple regions indicate non-ESEA lineages (AASI and AA lineages). All other labeled ancestries are subsets of ESEA lineage, where Hòabìnhian ancestry (yellow region) is deeply diverged from all other labeled ancestries. A dashed border indicates ancestries that are only partially (AASI, Jōmon, Fujian in mainland Asia) or not (Guangxi, Hòabìnhian) represented in present-day populations. Black arrows with a date in gray (ka = thousand years ago) indicate documented gene flow related to those ancestries, while black arrows with a dashed line indicate that connections were observed but the underlying demographic history is not well-known. Red arrows and associated red text describe expansion of ancestry associated with northern East Asians associated with Amur or Yellow River ancestry. A question persists regarding which populations in East Asia mixed with populations associated with ANS ancestry, resulting in the signature of genetic admixture found in Paleosiberians and Native Americans. Using shotgun sequencing, Sikora et al. [69] generated whole genomes from six 8,000-year-old individuals from Devil’s Gate Cave in the Primorye region of Siberia (DevilsCave_N) who were previously partially sequenced [82]. They showed that these individuals grouped with present-day East Asians, and represented the best Asian source population for Kolyma1 [69]. In a later study, Mao et al. [61] targeted the 1.2M SNP panel for a 14,000-year-old individual from the Amur River region (AR14K) and found using a qpAdm mixture analysis (Box 1] and qpGraph modeling analysis that AR14K was a better source population for Asian ancestry in Kolyma1 than the DevilsCave_N individuals. Using f4-statistics, both DevilsCave_N and AR14K share a close genetic relationship to each other and group phylogenetically with other ancient northern East Asian individuals rather than ancient southern East Asian individuals [61,68]. Mao et al. argue that populations in the Amur River region likely played an important role in interactions with populations associated with ANS ancestry [61]. Denoted here as Amur ancestry (Box 2, Figure 2), the ancestry associated with AR14K and DevilsCave_N found in the Amur River and Primorye region greatly affected the human landscape in Siberia and the Americas. Sikora et al. also estimated split times among Siberian and East Asian populations using fastsimcoal2. With a mutation rate of 1.25 x 10-8 per generation per site and a generation time of 29 years, they estimated that the Asian source that contributed to Kolyma1 diverged around 24,000 years ago [69]. If that Asian source was Amur ancestry, then 24,000 years ago may also indicate the split time separating Amur ancestry from other Asian ancestries. Moreno-Mayar sequenced a high coverage genome of an 11,500-year-old individual (USR1) at Upward Sun River, Alaska [72] who shares a close relationship with present-day Native Americans. They also estimated split times by modeling the evolutionary relationship between USR1 and present-day East Asians, Siberians, and Native Americans with diCal2 and momi2 (Box 1). Using the same mutation rate and generation time as Sikora et al., they estimated that Native Americans and present-day Siberians separated 36,000–25,000 years ago, and that USR1 separated from other Native Americans around 20,000 years ago. They used these estimates to suggest that admixture related to ANS ancestry likely occurred between 25,000 and 20,000 years ago [72]. These results are consistent with qpGraph analyses including Paleosiberians, Native Americans, and East Asians [69,73], where the Asian source for Native Americans separated earlier than the Asian source for Paleosiberians. Further south, the 1.2M SNP panel was used to analyze ancient DNA from a 10,500-year-old individual (Longlin) from Longlin Cave in Guangxi, a region in southern China [62] between the Fujian region and Southeast Asia. Both phylogenetic analyses using Treemix and tests of relative genetic similarity using f4-statistics showed that Longlin was more closely related to 9,000–4,000-year-old East Asians from northern and southern coastal China than with Tianyuan or Hòabìnhians [62]. However, these same analyses showed that Longlin was an outgroup of the 9,000–4,000-year-old northern and southern East Asians. The ancestry associated with Longlin, denoted as Guangxi ancestry (Box 2, Figure 2), was not observed in either historical samples from Guangxi or present-day East and Southeast Asians [62], which suggests that Guangxi ancestry did not persist to the present-day. Inferences using ancient DNA from 8,000–3,000-year-old humans in Japan indicated that a population in the Japanese archipelago was also differentiating during the end of the Upper Paleolithic. These ancient individuals were associated with the Jōmon period of Japan, the first cultural period found in the archaeological record dating from 16,000 to 2,800 years ago. They were originally from the northern reaches of Hokkaido [83], the central and northern regions of Honshu [63,81,84,85], and the southern island of Kyushu [64]. These Jōmon individuals consistently cluster together in a PCA and show high genetic similarity to each other distinct from that found in other Asian populations; their associated ancestry is denoted here as Jōmon ancestry (Box 2, Figure 2). Like Longlin, they are more closely related to 9,000–4,000-year-old East Asians from coastal China than to Tianyuan or Hòabìnhians, but are an outgroup of these northern and southern East Asians. Some have argued for the presence of excess connections to Hòabìnhians by fitting the data to a graph that includes admixture with a Hòabìnhian-related population and finding different f4 patterns for Hòabìnhians compared to younger Southeast Asians in comparisons to a Jōmon individual [63]; however, alternative admixture graphs and f4-statistic comparisons do not show evidence for this connection [68,85,86]. A lack of Upper Paleolithic samples representing South Asia or Southeast Asia makes it difficult to assess the ancestral patterns within these regions. While sampling of ancient humans in the Upper Paleolithic is still sparse, the findings to date reveal a diverse set of ancestries across Asia with high genetic structure across East Asia and interactions between distinct populations in northern Asia. |
|
|
|
Post by Admin on Feb 15, 2023 18:48:37 GMT
Rapid Human Dispersals Within Asia in the Last 10,000 Years
Ancient DNA capture on the 1.2M SNP panel has been performed on multiple individuals across Asia dating to the last 10,000 years, providing insight on the population dynamics leading to the genetic composition observed today. In the following section, I review demographic patterns determined from analysis of humans dating to the last 10,000 years.
Population dynamics in northern East Asia, Siberia, and the Tibetan Plateau
A PCA including present-day populations of East Asian ancestry in northern East Asia, Siberia, and the Tibetan Plateau shows two major clines [61,62,67]. One cline includes present-day populations in the Amur River region (e.g., the Ulchi) and inland across the eastern steppe (e.g., the Xibo), whereas another cline represents populations on or near the Tibetan Plateau. Analysis of ancient individuals from these regions revealed a more complex history than observed with present-day East Asians.
In the Late Upper Paleolithic of the Amur River region and Siberia, ancient individuals sampled were associated with Amur ancestry (e.g., AR14K) or an admixed Paleosiberian ancestry (e.g., UKY and Kolyma1). In the Amur River region, 14,000–3,000-year-old individuals shared greater genetic similarity with each other, and in a PCA, they and the 8,000–7,000-year-old individuals at Devil’s Gate Cave and Boisman-2 cemetery from the coastal Primorye region adjacent to the Amur River region [69,81,82] clustered together and with present-day populations from the region, particularly the Tungusic and Mongolic speakers [61,69,80–82]. Thus, shared ancestry has persisted within this region from at least 14,000 years ago to the present-day.
Further west in central Siberia, DNA sampling of ancient humans from 8,500 years ago to historical times from the Baikal region, Mongolia, and Inner Mongolia has consistently revealed high genetic diversity and interaction in central Siberian populations. In mixture models using qpAdm [61,69,74,81,87], most 8,500–3,000-year-old populations from central Siberia show a mixture of Amur ancestry (e.g., DevilsCave_N) and ANS ancestry (e.g., from Mal’ta1 or Kolyma1). Several ancient individuals dating from 8,000-5,000 years ago in eastern Mongolia and Inner Mongolia [68,87] and Early Iron Age individuals dating to 3,000 years ago in Central Mongolia (i.e., Slab Grave) [87] show little to no evidence of admixture with populations carrying ANS ancestry, which suggests that Mongolia was an interaction zone for diverse populations. From 5,000 to 3,500 years ago, central and western steppe populations associated with the Yamnaya and Afanasievo cultures, followed by the Sintashta culture, migrated to Mongolia and contributed to Mongolian and southern central Siberian populations (Figure 2). They brought an admixed ancestry related to populations associated with ANS ancestry, basal ancestry from the Levant and Iran, and ancestry related to hunter-gatherers from the Caucasus region of eastern Europe [42,59,81,87], denoted here as Steppe ancestry (Box 2). Beginning around 2,000 years ago in Mongolia, however, historical individuals were more closely related to the present-day Han than individuals associated with Amur ancestry [87], indicating interactions with populations south of Mongolia (Figure 2).
Based on PCA, several 8,000–5,000-year-old individuals in the Baikal region, Mongolia, and Inner Mongolia shifted towards the Tibetan cline rather than following the northern Asian cline like most Amur and Primorye region samples [68,81]. In D-statistic and Treemix analyses, 3,000–1,000-year-old individuals from Nepal who shared a close genetic relationship with present-day Tibetans [67] tended to group more with ancient individuals from the Shandong region associated with Yellow River ancestry than with more southern Fujian individuals [68]. Present-day Tibetans show a similar genetic pattern [81]. Thus, the currently sampled Tibetan populations fall within the genetic diversity observed in northern East Asians, although sampling of ancient individuals is needed to determine if Tibetans possess deeply diverging ancestry local to the plateau, as has been suggested previously [88]. Thus far, sparse sampling on the Tibetan Plateau have made it difficult to clarify which demographic processes could explain the shift in the PCA towards the Tibetan cline in populations associated predominantly with Amur ancestry (Figure 2).
Present-day northern and northeastern Siberians (e.g., Even) share a close genetic relationship to northern East Asians, particularly those associated with Amur ancestry. With low levels of Paleosiberian ancestry in present-day Siberians as estimated using qpAdm mixture analysis, Sikora et al. concluded that population movement from northern East Asia into Siberia must have occurred [69] (Figure 2). On the Siberian coast by the Beringian Sea, 3,000–2,000-year-old individuals are associated with Native American and Paleosiberian ancestries, suggesting back migration of populations from the Americas that re-entered and mixed with the remaining populations associated with Paleosiberian ancestry still found in the far reaches of northeastern Siberia [69] (Figure 2).
Along the middle to lower reaches of the Yellow River, ancient DNA was retrieved from 9,500–2,000-year-old individuals [68,80,81]. D-statistic, outgroup f3-analysis, and phylogenetic analyses show that these ancient individuals shared a close genetic relationship to each other and ancient individuals from the Amur River region [68,80,81]. In a PCA, they fall at the base of the northern East Asian cline [61,68,80,81]. They are closely related to each other to the exclusion of Amur River populations, indicating a shared ancestry distinct from that found in the Amur River region, denoted here as Yellow River ancestry (Box 2, Figure 2). Ancient individuals living between the Amur and Yellow River regions showed varying affinities to populations from these two regions, indicating high levels of interaction between populations associated with these two ancestries [80]. In tests of genetic similarity, Yang et al. found that ancient individuals from the lower reaches of the Yellow River in the Shandong region, who were associated with Yellow River ancestry, shared more alleles with present-day East and Southeast Asians [68] than ancient individuals from further south. These results suggested that populations along the Yellow River may have played a major role in the formation of present-day populations of East and Southeast Asia.
|
|
|
|
Post by Admin on Feb 18, 2023 2:25:53 GMT
Ancestry and admixture in southern East Asia, Southeast Asia, and the Japanese archipelago
In Fujian, a second 9,000-year-old individual from the Qihe cave site and 8,000–7,000-year-old individuals from Liang Island [68] off the coast of Fujian shared a close genetic relationship with Qihe3, suggesting that they are all associated with the same ancestry [62], denoted here as Fujian ancestry (Box 2, Figure 2). These ancient individuals also share a close genetic relationship with individuals from other Fujian sites dating to 4,000 years ago, showing that Fujian ancestry persisted at high levels from 12,000-4,000 years. Yang et al. [68], used tests of genetic differentiation to show that ancient individuals associated with Yellow River and Fujian ancestries were more genetically differentiated from each other than from present-day northern and southern East Asians. This was largely due to present-day East and Southeast Asians sharing more alleles with individuals associated with Yellow River ancestry than individuals associated with Fujian ancestry. Estimates of mixture proportions using qpAdm showed that most present-day East and Southeast Asians are a mixture of Yellow River, Fujian, and Paleosiberian ancestries [68]. The ancestry of Southeast Asian farmers postdating 4,000 years [63,89], unlike those associated with the deeply diverged Hòabìnhian ancestry, is primarily related to present-day East Asians, indicating that migration southwards from East Asia from 4,000 years ago profoundly influenced the genetic makeup of Southeast Asians (Figure 2). Taken together, these patterns support a hypothesis that migration and admixture between ancient populations associated with Yellow River and Fujian ancestries greatly influenced the current human landscape across East and Southeast Asia.
In the islands of Taiwan and Southeast Asia, sampling of ancient and present-day individuals associated with an Austronesian expansion that eventually occupied far reaching islands of Polynesia are associated with Fujian ancestry. Among present-day populations, Austronesians share the most genetic similarity with 8,000–4,000-year-old individuals from Fujian, showing that unlike mainland Asia, Fujian ancestry persisted at high levels across island Austronesian populations [68]. Some studies have suggested that the Austronesian dispersal moved from Taiwan into the islands of Polynesia [90], but others claim more than one migratory route, including through mainland Southeast Asia [91,92]. Larena et al. [75] recently sampled present-day Cordillerans in the Philippines. Using qpAdm mixture analysis, they observed that the ancient individuals from Fujian and present-day Austronesian speakers in Taiwan showed admixture with individuals associated with Yellow River ancestry, a pattern not observed for the Cordillerans. They used this pattern to argue that Cordillerans must have derived from another migration, perhaps through mainland Southeast Asia [75]. In the islands of Southeast Asia and Oceania, ancient and present-day populations have shown varying levels of ancestry associated with the AA and CSEA lineages [75,93,94], revealing complex layers of migration and interaction between deeply divergent populations in the islands of Southeast Asia and Southwest Pacific (Figure 2).
In the Guangxi region, the Late Upper Paleolithic Longlin was found to be an outgroup to a clade containing ancient individuals associated with Yellow River and Fujian ancestries and associated with Guangxi ancestry. Guangxi ancestry is not associated with any present-day East and Southeast Asians, and 8,000–6,000-year-old individuals from Guangxi (Dushan, Baojianshan) hint at large genetic shifts as the explanation. Dushan and Baojianshan have a closer genetic relationship to individuals associated with Fujian ancestry (Qihe3) than individuals associated with Guangxi ancestry (Longlin), and estimates of mixture proportions using qpAdm showed that they were best described as a mixture of populations associated with Guangxi and Fujian ancestries [62] (Figure 2). In Treemix and qpAdm analyses, the 6,000-year-old Baojianshan additionally showed evidence of admixture with a Southeast Asian hunter-gatherer group, the first detection of Hòabìnhian ancestry in southern China [62] (Figure 2). Historical individuals in Guangxi, dating to 1,500 and 500 years ago, are genetically most similar to present-day southern East Asians, and thus show high levels of admixture from populations associated with Yellow River ancestry.
In the Japanese archipelago, 7,000–3,000-year-old individuals sampled to date associated with the Jōmon culture are all closely related to each other and are associated with Jōmon ancestry. Using Treemix and f4-ratio analyses (Box 1), Adachi et al. [64] found that present-day Japanese populations had 10% Jōmon ancestry. This finding broadly supports a core tenet of the dual structure hypothesis for the population history of Japan, wherein migrants from mainland Asia, likely through the Korean peninsula, moved to the Japanese archipelago starting 3,000 years ago, and mixed with indigenous Jōmon populations [86,95]. Adachi et al. also estimated that present-day Korean and Ulchi populations in northeast Asia show 5%–8% Jōmon ancestry [64]. Furthermore, in f4-statistics, Jōmon individuals show connections to present-day Austronesians and 8,000–7,000-year-old individuals from coastal southern East Asia and Siberia [85,86]. These ties to coastal and island populations suggest that the Jōmon may not have been completely isolated after their migration into the Japanese archipelago (Figure 2).
|
|

