Post by Admin on Aug 22, 2021 0:33:21 GMT
Genetic transformation following the Bronze Age
Neither the Jagodnjak nor the Dalmatian Bronze Age groups approximate present-day populations from the region in PCA space, indicating that further significant population changes have since occurred. Our single individual from Popova zemlja, Croatia_Pop_RomanP, provides rare genomic data for Croatia after the Bronze Age33. (Table 1, Fig. 4a, Supplementary Table S1). We find this individual clustering with present-day populations of Croatia, Bulgaria and Romania in PCA and UMAP space (Figs. 2, 3c).We investigated this clustering with f4-statistics and could confirm cladality of this individual with present-day Croatians compared to other ancient and present-day populations in Europe (Supplementary Table S7). We then tested population continuity with qpWave, and found Croatia_RomanP was consistent with forming a genetic clade with present-day Croatians, as well as Bulgarians or Hungarians (p = 0.78 respectively) (Supplementary Table S4). Although based on a single individual who may or may not be representative for the wider population in that time period, this data point indicates that a broadly present-day genetic signature had already formed by Roman times, and any further population turnovers were not as significant as previous ones.
Figure 4
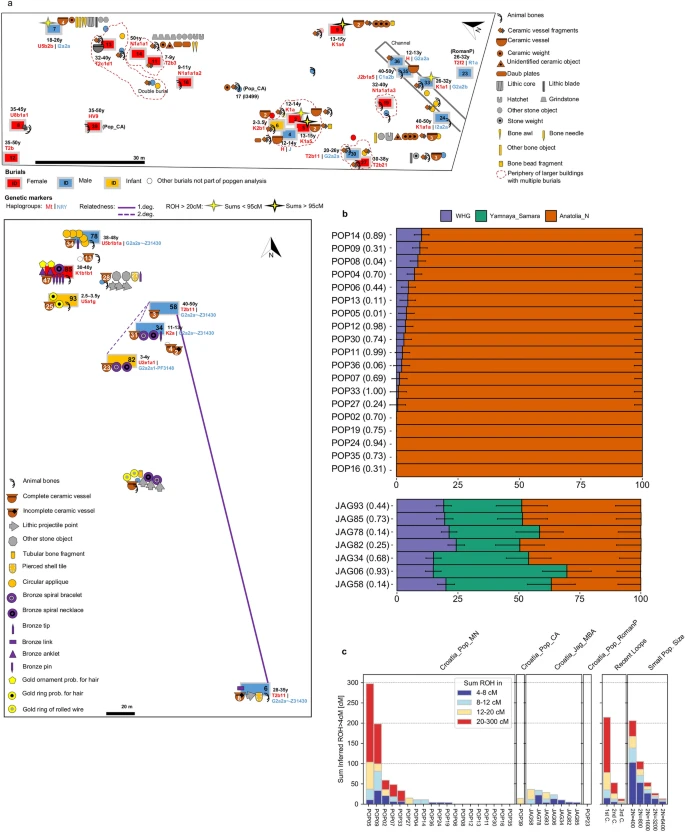
Burials and individual measures of genetic variation. (a) Plan of Popova zemlja (top) and Jagodnjak necropolis (bottom) showing inhumations, grave goods and demographic information. Burials with increased translucency indicate these were not sampled due to poor preservation or uncertain context. Burials without a fill colour indicate preservation was too poor for sex estimation. Number of grave good objects exceeding one are indicated. Estimated age at death is indicated as a range in years (y) for individuals included in population genetic analysis. Individuals with an estimated age at death of under 8 years are indicated as infants. (b) Stacked bar plots showing distal admixture models calculated with qpAdm for individuals from Croatia_Pop_MN (top) and Croatia_Jag_MBA (bottom), with p-values stated in brackets. Nested two-way models between WHG and Anatolia_N are shown for Croatia_Pop_MN. Three-way admixture models between WHG, Anatolia_N and Yamnaya_Samara are shown for Jagodnjak. Standard error bars are shown as one standard error in each direction (Source data in Supplementary Table S4). Plot produced using R 3.5.2.102 (c) Sums of inferred runs of homozygosity (ROH) greater than 4 cM calculated for each individual with hapROH (Source data in Supplementary Table S10). “Recent loops” and “Small Pop. Size”, generated by hapROH, show the expected distribution of ROH from simulated data for close parental relationships (C = cousin) and small effective population sizes respectively. Plot produced using R 3.5.2102.
Intra-population genetic diversity, kinship and demography
We next investigated intra-population genetic heterogeneity and patterns of demography by analyzing individual ancestry, haplotypic diversity, consanguinity and runs of homozygosity (ROH) (Fig. 4a–c).
Individual ancestry modelling with qpAdm confirms high genetic homogeneity among all Middle Neolithic individuals, with a large majority exhibiting either no or low Western hunter-gatherer ancestry introgression at p > 0.01 (given multiple testing) and no significant differences between burial rites (Fig. 4a-b, Supplementary Table S4). Jagodnjak MBA individuals also display unstructured ancestry but a greater degree of heterogeneity particularly in the proportion of steppe ancestry (26–55 ± 10%).
Mitochondrial haplotypes assigned with Haplogrep (Methods) identified high haplotypic diversity in the Neolithic (Table 1, Fig. 4a, Supplementary Table S1). A single individual carries an mtDNA haplogroup associated with European hunter-gatherer populations (U5)3, while almost 60% of haplotypes belong to branches K and T2. These, together with clades N1a and J account for most of the variation reported in early Neolithic Starčevo and Linearbandkeramik farming communities of northern Croatia and the adjacent Carpathian Basin3, pointing to genetic continuity. Y chromosomal haplogroups assigned with Yleaf (Methods), similarly show a high degree of diversity, with seven males represented by four different haplogroups (Table 1, Fig. 4a, Supplementary Table S1). Two of these, C and I, are found in Mesolithic populations, while G2a is commonly associated with the Neolithic expansion3. We also detected high mtDNA haplotypic diversity in Jagodnjak, with two males carrying the same defining mutation for T2b11, while three U subclades and two K subclades, present in Mesolithic populations, are also represented. Y chromosomal haplogroups are restricted to the G2a clade however, four of which belong to the same haplotype G2a2a-Z31430. The fifth individual lacks a read covering the determining mutation and can be assigned the next upstream haplotype. These shared haplotypes indicate relatedness between individuals, which was further explored with genome-wide kinship analysis.
Pairwise genome-wide mismatch rates (Fig. 4a, Supplementary Fig. S8, Supplementary Table S9, Methods) identified JAG58 as a first degree relative of JAG06, and their shared mtDNA and Y chromosomal haplotypes (Table 1, Fig. 4a, Supplementary Table S1) are consistent with the interpretation that these men were full siblings rather than father and son. Additionally, JAG58 is a second degree relative of JAG34 and JAG82. JAG06 and JAG34 also have lowered pairwise mismatch rates indicative of a third-degree or more distant consanguineous relationship, as do JAG78 and JAG93. An adult woman, JAG85, is the only individual not part of any pedigree.
We did not identify first or second degree genetic kin among the Middle Neolithic individuals, however POP05 has a lowered pairwise mismatch rate indicative of a more distant consanguineous relationship with another two subadults (POP02 and POP04), all buried in the same pit house. POP05 also has a lowered pairwise mismatch rate with POP24, an older adult male deposited in the channel. Three of these individuals also share the same maternal K1a haplogroup (Table 1, Fig. 4a, Supplementary Fig. S8, Supplementary Tables S1 and S9). Although high inbreeding coefficients can inflate kinship coefficients, (see below), taken together, this suggests they are part of the same matrilineal pedigree. POP24, deposited in the channel, also appears distantly related to POP02, POP04 and POP07, with whom he shares the same Y chromosomal haplotype I2a2a-M223.
We estimated runs of homozygosity greater than 4 cM (centiMorgans) with hapROH (Methods) to assess levels of inbreeding and infer past mating practices (Fig. 4a and c, Supplementary Fig. S9, Supplementary Table S10). Long ROH > 20 cM indicate recent consanguineous mating, whilst many short runs suggest more distant limitations on effective population size. Eight Middle Neolithic individuals have no ROH, pointing to a lack of inbreeding and thus a large mating pool. Among the remainder however, two individuals, POP05 and POP09, harbour many long ROH > 20 cM that strikingly sum to over 95 cM. This is consistent with them being the offspring of first cousins or equivalent, a rarity in the ancient DNA record34. A further three individuals have fewer long ROH > 20 cM indicative of their parents being second cousins or equivalent, while the remainder display ROH pointing to relatedness within the last ten generations. Contrastingly, individuals from Jagodnjak exhibit much lower sums of ROH, with no long runs > 20 cM, and some short runs among all individuals suggestive of more distant relatedness. Infant JAG93 and JAG58 have ROH > 12 cM suggestive of parental relatedness up to five generations ago, although shorter ROH is found in JAG58’s first degree relative JAG06, demonstrating some heterogeneity that is mirrored in their admixture profiles. Copper Age and Roman Period individuals show few to no ROH greater than 4 cM.
Finally we analysed a panel of functional SNPs associated with phenotypic traits under recent selection35,36,37,38,39,40 (Supplementary Table S11, Methods) and found that derived alleles for lighter skin pigmentation (SLC45A2 and SLC24A5) and lighter eye colour (HERC2)35,36,37 are present in individuals from all time periods, consistent with previous findings that point to the derived allele for SLC24A5 rapidly increasing in frequency during the Neolithic as a result of migration38,39 . In addition, all individuals carry ancestral alleles for the SNP responsible for lactase persistence into adulthood among Europeans (LCT rs4988235) 39,40, suggesting they may not have been able to digest lactose as adults, and is concordant with previous findings that lactose tolerance in Europe remained at low frequency into the Bronze Age38,39.
Neither the Jagodnjak nor the Dalmatian Bronze Age groups approximate present-day populations from the region in PCA space, indicating that further significant population changes have since occurred. Our single individual from Popova zemlja, Croatia_Pop_RomanP, provides rare genomic data for Croatia after the Bronze Age33. (Table 1, Fig. 4a, Supplementary Table S1). We find this individual clustering with present-day populations of Croatia, Bulgaria and Romania in PCA and UMAP space (Figs. 2, 3c).We investigated this clustering with f4-statistics and could confirm cladality of this individual with present-day Croatians compared to other ancient and present-day populations in Europe (Supplementary Table S7). We then tested population continuity with qpWave, and found Croatia_RomanP was consistent with forming a genetic clade with present-day Croatians, as well as Bulgarians or Hungarians (p = 0.78 respectively) (Supplementary Table S4). Although based on a single individual who may or may not be representative for the wider population in that time period, this data point indicates that a broadly present-day genetic signature had already formed by Roman times, and any further population turnovers were not as significant as previous ones.
Figure 4

Burials and individual measures of genetic variation. (a) Plan of Popova zemlja (top) and Jagodnjak necropolis (bottom) showing inhumations, grave goods and demographic information. Burials with increased translucency indicate these were not sampled due to poor preservation or uncertain context. Burials without a fill colour indicate preservation was too poor for sex estimation. Number of grave good objects exceeding one are indicated. Estimated age at death is indicated as a range in years (y) for individuals included in population genetic analysis. Individuals with an estimated age at death of under 8 years are indicated as infants. (b) Stacked bar plots showing distal admixture models calculated with qpAdm for individuals from Croatia_Pop_MN (top) and Croatia_Jag_MBA (bottom), with p-values stated in brackets. Nested two-way models between WHG and Anatolia_N are shown for Croatia_Pop_MN. Three-way admixture models between WHG, Anatolia_N and Yamnaya_Samara are shown for Jagodnjak. Standard error bars are shown as one standard error in each direction (Source data in Supplementary Table S4). Plot produced using R 3.5.2.102 (c) Sums of inferred runs of homozygosity (ROH) greater than 4 cM calculated for each individual with hapROH (Source data in Supplementary Table S10). “Recent loops” and “Small Pop. Size”, generated by hapROH, show the expected distribution of ROH from simulated data for close parental relationships (C = cousin) and small effective population sizes respectively. Plot produced using R 3.5.2102.
Intra-population genetic diversity, kinship and demography
We next investigated intra-population genetic heterogeneity and patterns of demography by analyzing individual ancestry, haplotypic diversity, consanguinity and runs of homozygosity (ROH) (Fig. 4a–c).
Individual ancestry modelling with qpAdm confirms high genetic homogeneity among all Middle Neolithic individuals, with a large majority exhibiting either no or low Western hunter-gatherer ancestry introgression at p > 0.01 (given multiple testing) and no significant differences between burial rites (Fig. 4a-b, Supplementary Table S4). Jagodnjak MBA individuals also display unstructured ancestry but a greater degree of heterogeneity particularly in the proportion of steppe ancestry (26–55 ± 10%).
Mitochondrial haplotypes assigned with Haplogrep (Methods) identified high haplotypic diversity in the Neolithic (Table 1, Fig. 4a, Supplementary Table S1). A single individual carries an mtDNA haplogroup associated with European hunter-gatherer populations (U5)3, while almost 60% of haplotypes belong to branches K and T2. These, together with clades N1a and J account for most of the variation reported in early Neolithic Starčevo and Linearbandkeramik farming communities of northern Croatia and the adjacent Carpathian Basin3, pointing to genetic continuity. Y chromosomal haplogroups assigned with Yleaf (Methods), similarly show a high degree of diversity, with seven males represented by four different haplogroups (Table 1, Fig. 4a, Supplementary Table S1). Two of these, C and I, are found in Mesolithic populations, while G2a is commonly associated with the Neolithic expansion3. We also detected high mtDNA haplotypic diversity in Jagodnjak, with two males carrying the same defining mutation for T2b11, while three U subclades and two K subclades, present in Mesolithic populations, are also represented. Y chromosomal haplogroups are restricted to the G2a clade however, four of which belong to the same haplotype G2a2a-Z31430. The fifth individual lacks a read covering the determining mutation and can be assigned the next upstream haplotype. These shared haplotypes indicate relatedness between individuals, which was further explored with genome-wide kinship analysis.
Pairwise genome-wide mismatch rates (Fig. 4a, Supplementary Fig. S8, Supplementary Table S9, Methods) identified JAG58 as a first degree relative of JAG06, and their shared mtDNA and Y chromosomal haplotypes (Table 1, Fig. 4a, Supplementary Table S1) are consistent with the interpretation that these men were full siblings rather than father and son. Additionally, JAG58 is a second degree relative of JAG34 and JAG82. JAG06 and JAG34 also have lowered pairwise mismatch rates indicative of a third-degree or more distant consanguineous relationship, as do JAG78 and JAG93. An adult woman, JAG85, is the only individual not part of any pedigree.
We did not identify first or second degree genetic kin among the Middle Neolithic individuals, however POP05 has a lowered pairwise mismatch rate indicative of a more distant consanguineous relationship with another two subadults (POP02 and POP04), all buried in the same pit house. POP05 also has a lowered pairwise mismatch rate with POP24, an older adult male deposited in the channel. Three of these individuals also share the same maternal K1a haplogroup (Table 1, Fig. 4a, Supplementary Fig. S8, Supplementary Tables S1 and S9). Although high inbreeding coefficients can inflate kinship coefficients, (see below), taken together, this suggests they are part of the same matrilineal pedigree. POP24, deposited in the channel, also appears distantly related to POP02, POP04 and POP07, with whom he shares the same Y chromosomal haplotype I2a2a-M223.
We estimated runs of homozygosity greater than 4 cM (centiMorgans) with hapROH (Methods) to assess levels of inbreeding and infer past mating practices (Fig. 4a and c, Supplementary Fig. S9, Supplementary Table S10). Long ROH > 20 cM indicate recent consanguineous mating, whilst many short runs suggest more distant limitations on effective population size. Eight Middle Neolithic individuals have no ROH, pointing to a lack of inbreeding and thus a large mating pool. Among the remainder however, two individuals, POP05 and POP09, harbour many long ROH > 20 cM that strikingly sum to over 95 cM. This is consistent with them being the offspring of first cousins or equivalent, a rarity in the ancient DNA record34. A further three individuals have fewer long ROH > 20 cM indicative of their parents being second cousins or equivalent, while the remainder display ROH pointing to relatedness within the last ten generations. Contrastingly, individuals from Jagodnjak exhibit much lower sums of ROH, with no long runs > 20 cM, and some short runs among all individuals suggestive of more distant relatedness. Infant JAG93 and JAG58 have ROH > 12 cM suggestive of parental relatedness up to five generations ago, although shorter ROH is found in JAG58’s first degree relative JAG06, demonstrating some heterogeneity that is mirrored in their admixture profiles. Copper Age and Roman Period individuals show few to no ROH greater than 4 cM.
Finally we analysed a panel of functional SNPs associated with phenotypic traits under recent selection35,36,37,38,39,40 (Supplementary Table S11, Methods) and found that derived alleles for lighter skin pigmentation (SLC45A2 and SLC24A5) and lighter eye colour (HERC2)35,36,37 are present in individuals from all time periods, consistent with previous findings that point to the derived allele for SLC24A5 rapidly increasing in frequency during the Neolithic as a result of migration38,39 . In addition, all individuals carry ancestral alleles for the SNP responsible for lactase persistence into adulthood among Europeans (LCT rs4988235) 39,40, suggesting they may not have been able to digest lactose as adults, and is concordant with previous findings that lactose tolerance in Europe remained at low frequency into the Bronze Age38,39.
