|
|
Post by Admin on Sept 26, 2021 0:28:07 GMT
RESULTS We extracted DNA from the petrous portion of the temporal bone and tooth samples of 86 individuals, created double-stranded DNA libraries, and used shotgun or an in-solution capture approach to retrieve the entire mitochondrial genome and up to ~1.24 million single-nucleotide polymorphisms (SNPs) across the human genome from each individual (table S1A and Materials and Methods). Following criteria for authentication, we restricted all downstream analyses to specimens with low nuclear and mtDNA contamination, typical aDNA damage patterns, and SNP counts above 31,000 (table S1A). These quality controls resulted in a final sample set of 82 individuals that were grouped on the basis of their radiocarbon dates and genetic affinities into three time intervals: 48 individuals from 800 to 1 BCE (Iron Age and Roman Republic), 6 individuals from 1 to 500 CE (Imperial period), and 28 individuals from 500 to 1000 CE (12 from central Italy and 16 from southern Italy). While we would have preferred to mirror conventional historical chronologies (Etruscan to ~300 BCE and Republican to 27 BCE), neither the large radiocarbon dating intervals nor the genetic results warrant a division of the dataset in this way (Fig. 2B and table S2F). 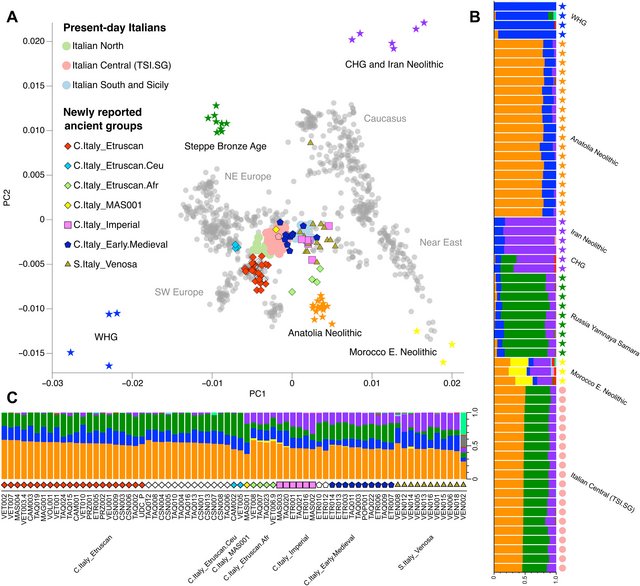 Fig. 2. Genetic map and clustering of ancient and present-day populations. (A) PCA inferred from genomic variation of 60 West Eurasian populations genotyped on the Human Origins array (gray dots with geographic regions labeled in gray) including present-day Italians (circles without outline) onto which we projected the newly reported ancient individuals (symbols with outline, not filled for nonradiocarbon-dated individuals) and comparative ancient populations (stars without outline). (B) Unsupervised admixture (K = 11) of comparative ancient individuals and 20 present-day central Italians (TSI.SG). (C) Unsupervised admixture (K = 11) of newly reported ancient individuals excluding genetically related individuals. WHG and CHG refer to Western and Caucasus hunter-gatherers, while SW and NE Europe refer to southwestern and northeastern Europe, respectively. Of individuals associated with the first time interval, the vast majority (40 of 48) form a genetic cluster here named “C.Italy_Etruscan” that overlaps with present-day Spanish individuals in a principal components analysis (PCA) built with West Eurasian populations from the Human Origins dataset (Fig. 2A) (21). Across this temporal interval (800 to 1 BCE), three groups of PCA outliers are identified, i.e., four individuals shifted toward northern African populations (C.Italy_Etruscan.Afr), three individuals shifted toward central European populations (C.Italy_Etruscan.Ceu), and one individual shifted toward Near Eastern populations (C.Italy_Etruscan_MAS001) (table S1A). To further inspect the genetic clustering of the central and southern Italian populations studied, we performed unsupervised ADMIXTURE on 71 individuals (Fig. 2, B and C) after the exclusion of genetically related individuals (table S1B and fig. S2). C.Italy_Etruscan individuals harbor the three genetic ancestries associated with Anatolian Neolithic farmers, European hunter-gatherers, and Bronze Age pastoralists from the Pontic-Caspian Steppe. C.Italy_Etruscan.Ceu carries a higher proportion of “steppe-related ancestry,” while C.Italy_MAS001 shows a genetic component maximized in Iranian Neolithic farmers. The latter is also present in C.Italy_Etruscan.Afr individuals alongside an ancestry component identified in an Early Neolithic Moroccan group. Since the spread of the steppe-related ancestry into Europe during the Late Neolithic and Early Bronze Age has been linked to the diffusion of Indo-European languages (15, 19) and is consistent with linguistic evidence for a “Steppe” homeland (22, 23), we attempted to formally estimate the proportion of steppe-related ancestry in Etruscan individuals who are putatively associated with the non–Indo-European Etruscan language. We first grouped 21 dated and genetically unrelated individuals from the C.Italy_Etruscan cluster (Fig. 3A) and modeled them with qpAdm (P > 0.05) as a mixture of steppe-related ancestry, represented by Bronze Age pastoralists from Samara in western Russia (Yamnaya), and Neolithic or Copper Age populations from Italy (table S4B). This analysis demonstrated around 25% ancestry from such a distal steppe-related source, which reached around 50% when comparative populations were reduced to those more proximate in time and space than the Yamnaya, e.g., central European Bell Beakers (Fig. 3B). Moreover, C.Italy_Etruscan can be modeled successfully as having derived its entire ancestry from other European populations such as the earlier Bell Beaker group from northern Italy and Iron Age populations from southern Europe (Iberia, Croatia, and Greece) (table S4A). PCA reveals a complete overlap between Iron Age and Roman Republic individuals from Tuscany and Lazio, including the ancient city of Rome (17), indicating that substantial levels of steppe-related ancestry were widespread and homogenized in the multilingual context known to include both Indo-European (i.e., Italic and Celtic) and non–Indo-European (i.e., Etruscan) speakers across central Italy by the Iron Age. |
|
|
|
Post by Admin on Sept 26, 2021 20:10:21 GMT
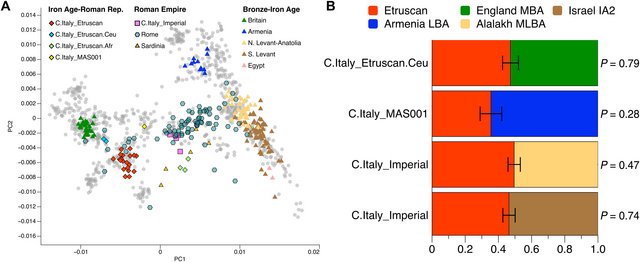 Fig. 3. Genetic placement and modeling of Iron Age and Roman Republic individuals. (A) PCA of Italian individuals from the Iron Age and Roman Republic (symbols with outline) plus Copper and Bronze Age individuals from Italy and other source populations (symbols without outline) used in qpAdm. Gray dots represent present-day West Eurasian individuals (see Fig. 2A). (B) Proportion of steppe-related ancestry in C.Italy_Etruscan and C.Italy_Etruscan.Ceu as estimated with qpAdm (table S4B). MN and EBA refer to Middle Neolithic and Early Bronze Age, respectively. The other contemporaneous ancestry groups identified in this study, albeit represented by small numbers of individuals, add detail and complexity to this picture. Two C.Italy_Etruscan.Ceu unrelated individuals (VET005 and CAM002) are characterized by a higher proportion of Yamnaya-related ancestry (40%) and are consistent with deriving from those Chalcolithic or Bronze Age European populations from both central and southern Europe that carried a high fraction of steppe-related ancestry (Fig. 3, A and B, and table S4, A and B). This signal is confirmed by the f4-statistic f4(Onge, Test; C.Italy_Etruscan.Ceu, C.Italy_Etruscan) that is significantly negative (z score > |3|) when the Test population comprises eastern hunter-gatherer (EHG) individuals representing about half of the Yamnaya-related ancestry (table S2C) (19). This indicates higher affinity of EHG to C.Italy_Etruscan.Ceu than to the main C.Italy_Etruscan cluster, while the opposite is observed when the Test population is restricted to southern European Neolithic groups without steppe-related ancestry. We then tested whether C.Italy_Etruscan.Ceu carries a signature of admixture with local ancestry. Statistics of the form f3(C.Italy_Etruscan, Test; C.Italy_Etruscan.Ceu) do not reveal evidence of C.Italy_Etruscan.Ceu resulting from admixture between C.Italy_Etruscan and any of the 255 ancient Test populations in our dataset (table S3A). Moreover, while it is possible to model C.Italy_Etruscan.Ceu in qpAdm as a mixture between the Etruscan-related individuals and central European Bell Beakers (Fig. 3B), the model still holds when C.Italy_Etruscan is moved to the reference set and is replaced by Neolithic populations, consistent with an incoming northern ancestry that did not admix locally (table S4C) (24). Last, we estimated the admixture dates for two C.Italy_Etruscan.Ceu individuals using the software DATES (25). Individual VET005 resulted in an unfeasible date (negative value), whereas CAM002 provided an admixture date of 19.7 ± 8.6 generations, which corresponds to 572 ± 249 years before the individual’s date (seventh century BCE). These results suggest that the two tested C.Italy_Etruscan.Ceu individuals represent an incoming ancestry that is not the result of recent local admixture. Those individuals originate from two different archeological sites and are radiocarbon dated to the seventh and third century BCE, respectively. Therefore, rather than persisting across several centuries, this distinct ancestry profile might have arrived independently from northern regions into central Italy multiple times. The three individuals grouped in C.Italy_Etruscan.Afr—after exclusion of a genetic outlier (Materials and Methods, Fig. 4A, and table S2A)—cannot be modeled as a mixture between Neolithic-related and Bronze Age–related European ancestries (table S4B). These individuals are dated to around 300 BCE and were excavated at two archeological sites separated by a distance greater than 100 km (Tarquinia and Vetulonia). Statistics in the form f3(C.Italy_Etruscan, X; C.Italy_Etruscan.Afr) and f4(Onge, X, C.Italy_Etruscan, C.Italy_Etruscan.Afr) show evidence of C.Italy_Etruscan.Afr being a mixture of Etruscans and ancient or modern-day individuals (as a proxy) carrying high proportions of north African or sub-Saharan ancestries (tables S2, B and C, and S3, A and B). Using qpAdm, all admixture models including C.Italy_Etruscan as one of the sources are rejected (table S4D). However, given the limited availability of genomes from northern Africa with comparable ages (26), we caution that ancestry proportions might be more correctly estimated as additional data from this region become available. |
|
|
|
Post by Admin on Sept 27, 2021 4:19:37 GMT
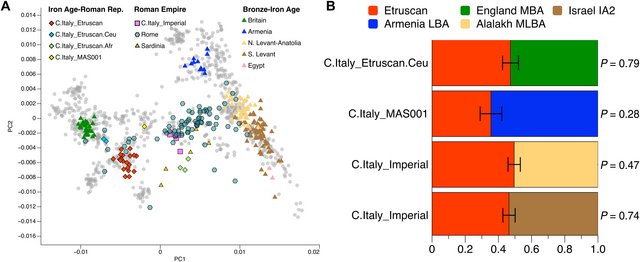 Fig. 4. Genetic placement and modeling of Iron Age and Imperial period individuals. (A) PCA of Italian individuals from the Iron Age to the Imperial period (symbols with outline) plus Bronze and Iron Age individuals (symbols without outline) used as source populations in qpAdm. Gray dots represent present-day west Eurasian individuals (see Fig. 2A). (B) Proportion of C.Italy_Etruscan ancestry in Iron Age outlier groups and in C.Italy_Imperial as estimated with qpAdm. For each test, the reported non-C.Italy_Etruscan population is the one providing a proportion that is closest to the median among all fitting models (table S4D). MBA, MLBA and LBA refer to Middle, Middle-Late and Late Bronze Age, respectively, and IA refers to Iron Age. Contrary to previously reported findings from Bronze Age Sicily and Iron Age Sardinia (27, 28), we do not find evidence for Iranian-related ancestry in individuals from central Italy older than 2000 years (fig. S3). We were able to model C.Italy_Etruscan and C.Italy_Etruscan.Ceu as a mixture between three distal sources [Anatolia_Neolithic, Western hunter-gatherers (WHG), and Yamnaya_Samara] even when Neolithic Iranian individuals were placed in the reference set of qpAdm (table S4H). This suggests that the genetic history of Sicilians and Sardinians during the Bronze and Iron Ages was substantially different from that of populations on the Italian mainland, as confirmed by the distinctive spheres of interaction observed in the archeological record (29). The C.Italy_Etruscan_MAS001 individual represents a single exception in our dataset showing a shift in PCA space toward Near Eastern populations ~200 BCE (Fig. 4A). While f-statistics do not significantly reject ancestry continuity with the C.Italy_Etruscan cluster (table S2C), an admixture model between Neolithic- and steppe-related ancestries does not fit the genetic profile of this individual (table S4B). Instead, C.Italy_Etruscan_MAS001 can be modeled as a mixture between the C.Italy_Etruscan cluster and populations from the Caucasus, such as Bronze Age Armenians (Fig. 4B), indicating the sporadic presence of Iranian-related ancestry in Etruria at least by the second century BCE. During the first half of the first millennium CE, we observe a marked shift in PCA space of all studied individuals toward the Near Eastern cline (Fig. 4A), distributed across the genetic space occupied by present-day southeastern European populations. We grouped nonoutlier individuals dating between 1 and 500 CE into the “C.Italy_Imperial” cluster (table S2A). Formal f4-tests reveal its higher affinity than C.Italy_Etruscan to ancient groups from Iran, Africa, and the Near East (table S2C). We then used qpAdm to quantify this group’s ancestry components, where C.Italy_Imperial was modeled as a mixture of the sources C.Italy_Etruscan and 158 published European and Near Eastern genomes from the Bronze and Iron Ages. As a result, the models that were found to fit the data best are those with a 38 to 59% contribution from Levantine or Anatolian populations into the local/preexisting C.Italy_Etruscan gene pool (Fig. 4B and table S4D). Substantial gene flow from the eastern Mediterranean was also reported in ancient individuals from Rome dated to the Imperial period (17). Despite our limited number of data points from the first five centuries CE, the new results suggest that the contribution of nonlocal ancestry in Rome was larger than in Etruria (Fig. 4A). However, this large-scale genetic impact of incoming groups during the Imperial period was not only limited to the metropolitan area around Rome but also extended into the neighboring and more distant regions considered here. Regarding the last temporal interval of our ancient genomic transect (500 to 1000 CE), we observe that individuals grouped in the “C.Italy_Early.Medieval” cluster are generally shifted toward central European groups compared to C.Italy_Imperial and largely overlap with present-day populations from central Italy (TSI.SG) (Fig. 5A) (30). Using f4-tests, we can show that this transition is confirmed by a reduced affinity of C.Italy_Early.Medieval toward eastern Mediterranean populations compared to C.Italy_Imperial (table S2D). Moreover, the C.Italy_Early.Medieval cluster can be modeled successfully in qpAdm as a mixture between the preceding C.Italy_Imperial group and Late Antique or Medieval groups from northern and eastern Europe (among the 59 populations tested) in estimated proportions of 60 to 90% and 10 to 40%, respectively (table S4E). Notably, among the best supported models are those that feature individuals associated with Longobard cemeteries from Hungary and northern Italy (31). If we specifically restrict the analyses to those Longobard-related individuals carrying unadmixed northern European genetic ancestry (Piedmont_N.Longobard), then we obtain a ~20% contribution to the C.Italy_Early.Medieval cluster (Fig. 5B). This finding is consistent with a genetic input of northern European ancestry in central Italy during the Longobard period. However, the influence of other Germanic tribes in Italy like the Ostrogoths could also have enhanced the observed genomic shift. |
|
|
|
Post by Admin on Sept 27, 2021 22:12:43 GMT
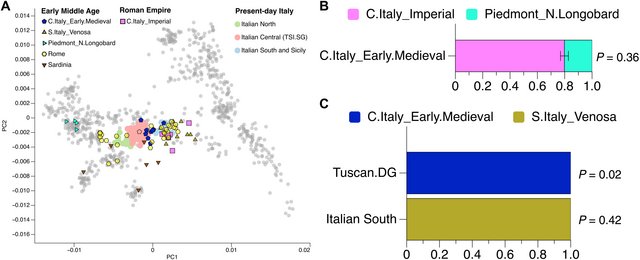 Fig. 5. Genetic placement and modeling of Early Middle Age and present-day individuals. (A) PCA of Italian individuals from the Imperial to the Early Medieval periods (symbols with outline) and present-day Italians (symbols without outline). Gray dots represent present-day west Eurasian individuals (see Fig. 2A). (B) Proportion of northern ancestry from Longobard-related individuals in C.Italy_Early.Medieval as estimated with qpAdm (table S4E). (C) Proportion of C.Italy_Early.Medieval and S.Italy_Venosa ancestry in modern-day central Italians (Tuscan.DG) and southern Italians, respectively, as estimated with qpAdm (table S4G). Since modern-day central Italians largely overlap in PCA space with C.Italy_Early.Medieval individuals (Fig. 5A), we tested the consistency of the former group deriving from the latter. To enhance resolution, qpAdm was implemented with present-day worldwide populations in the reference set. No present-day Italian populations are consistent with deriving from the C.Italy_Early.Medieval cluster (P values below 0.05), although high-coverage genomes from Tuscany (Tuscan.DG) (32) yielded no grounds for strong rejection of genetic continuity (P = 0.02) (Fig. 5C and tables S2E and S4G). This suggests that the genetic makeup of present-day central Italian populations was largely formed at least by 1000 CE. To investigate whether an analogous picture is observed in contemporaneous individuals from southern Italy, data from the Early Medieval archeological site of Venosa in Basilicata were similarly analyzed. With the exception of VEN002, all Venosa individuals (S.Italy_Venosa) broadly overlap modern-day southern Italian populations in PC space and can be jointly modeled in qpAdm as deriving from the same stream of ancestry (P = 0.42) (Fig. 5, A and C). In PCA space, most Medieval and Early Modern individuals from Rome fall in an intermediate position between Early Medieval groups from Tuscany and Basilicata (Fig. 5A). This distribution is thus consistent with the current north-south genetic cline that mirrors geography (33, 34) (fig. S4), with Italy bridging the genetic gap between Europe and the eastern Mediterranean. To investigate the potential influence of sex biases in these genomic transformations, we computed the frequency of uniparental markers through time. The mtDNA diversity does not seem to change substantially before and after year 1 CE (fig. S5A). By contrast, the newly reported central Italian individuals from 800 to 1 BCE show ~75% frequency of the Y-chromosome haplogroup R1b, mostly represented by the R1b-P312 polymorphism and its derived R1b-L2, that diffused across Europe alongside steppe-related ancestry in association with the Bell Beaker complex (16). This suggests that this R1b Y-chromosome lineage spread into the Italian peninsula with steppe-related movements during the Bronze Age. In the first millennium CE, its frequency is reduced to ~40% with higher occurrence of Near Eastern–associated Y-chromosome lineages such as J (fig. S5B). While we cannot rule out substantial female mobility, the marked shift in Y-chromosome haplogroup frequency indicates that male mobility played an important role in the observed genetic turnovers from the Imperial period onward. |
|
|
|
Post by Admin on Sept 28, 2021 0:12:06 GMT
DISCUSSION
Genomic analyses of 82 ancient individuals spanning ~2000 years of Italian history from Tuscany, Lazio, and Basilicata have revealed major episodes of genetic transformation. Across the first interval of our central Italian temporal transect (800 to 1 BCE), most individuals form a homogenous genetic cluster (C.Italy_Etruscan), indicating that the sporadic presence of individuals with ancestry tracing back to other regions did not leave a substantial local genetic legacy. In particular, and contrary to previous suggestions [see, e.g., (9, 10)], the Etruscan-related gene pool does not seem to have originated from recent population movements from the Near East. Etruscans carry a local genetic profile shared with other neighboring populations such as the Latins (table S2F) from Rome and its environs despite the cultural and linguistic differences between the two neighboring groups (17). A large proportion of the C.Italy_Etruscan genetic profile can be attributed to steppe-related ancestry, confirming a trend observed in most other European regions, that this genetic component also reached central Italy during the Bronze Age (18) but earlier than it reached Sardinia (27, 35). Since it is estimated that the split between Proto-Italic and descendant languages (e.g., Latin, Oscan, and Umbrian) took place during the second millennium BCE (36), their presence in Italy cannot directly be correlated with the at least one millennium–older movement of Yamnaya-related groups out of the Steppe. Instead, with the assumption that incoming steppe-related groups were responsible for the initial arrival of Indo-European languages in Europe, more evolved forms of Indo-European, including Italic, may have spread across Italy at a later stage. The only three Beaker complex–related individuals sequenced so far from northern Italy dated to around 2000 BCE reveal a nonubiquitous presence of steppe-related ancestry, suggesting that the admixture process was ongoing (16). The historically documented persistence of the non–Indo-European Etruscan language in Etruria indicates that this speech community was maintained despite a large-scale admixture, a situation similar to the Basque region in Iberia where a non–Indo-European language endures today (24). This linguistic persistence, combined with a genetic turnover, challenges simplistic assumptions that genes equal languages and suggests a more complex scenario that may have involved the assimilation of (early) Italic speakers by the Etruscan speech community, possibly during a prolonged period of admixture over the second millennium BCE. This scenario is made plausible by the recent finding of steppe-related ancestry in central Italy as early as 1650 BCE followed by an increase of this component through time (18). Although Etruscan is considered a relict language, which survived in central Italy until the Imperial Period, it was not isolated. Instead, Etruscan seems to be linked to both Rhaetic, a language documented in the eastern Alps in a population that ancient historians claim to have migrated from the Po valley, and to Lemnian, a language putatively spoken on ancient Lemnos in the Aegean Sea. This leaves the question open as to whether these enigmatic “Tyrsenian” languages may somehow relate to sea-borne expansions from the eastern Mediterranean (37). However, the lack of Iranian-related ancestry in C.Italy_Etruscan might also suggest that the close linguistic affinity across the Mediterranean Sea could represent population movements departing from the Italian peninsula.
The earliest individual in our dataset with a nonlocal genetic signature is radiocarbon dated to the seventh century BCE (CAM002) and exhibits a central European genetic profile. During the Early Iron Age, the Hallstatt culture associated with Celtic-related groups occupied the region north of the Alps. Although there is archeological evidence for exchange of goods and techniques between the Etruscan civilization and northern cultural groups from the eighth century BCE, extensive direct contacts are reported only later, during the La Tène cultural phase, when Celtic-related groups spread into northern Italy bordering Etruscan territories (38–40). In the presented dataset, we find another individual (VET005) radiocarbon dated to the third century BCE with a genetic profile that overlaps with individual CAM002, despite their ~400-year discrepancy in radiocarbon age. This suggests continuity in the source of the central European genetic ancestry sporadically found in Etruria from the Hallstatt to the La Tène cultural periods.
In the last four centuries BCE, we identify a higher proportion than in the previous four centuries of individuals carrying nonlocal genetic ancestries, which show the greatest affinities to the Near East and northern Africa. This could be explained by increased interconnections between Etruria and other regions, not only for those societies associated with harbors but also for those in the hinterland. An emblematic case of such cross-continental connectivity is observed at the archeological site of San Germano in Vetulonia (VET), where even within the same tomb, there is a clear genetic transition from a local genetic profile in the eighth to sixth century BCE to central European– and north African–related ancestries in the fourth to third century BCE. During the latter period, a similar northern African genetic signal is observed in two other individuals from a distantly located site [Tarquinia (TAQ)]. While more data from this temporal horizon are needed to determine whether these findings represent a general phenomenon, it is possible that the arrival of this ancestry was influenced by the expansion of the Carthaginian Empire across the Mediterranean (41, 42).
The vast majority of individuals from the first millennium BCE, however, show large levels of genetic continuity for more than 800 years from the post-Villanovan period to the end of the Roman Republic. While in Etruria we do not detect increased affinity toward central European ancestry, we are unable to exclude the possibility that admixture took place across neighboring regions between populations with similar genetic profiles, such as with Latin-associated groups (table S2F). However, the marked genetic stability in Etruria over almost a millennium accords with the historical record, which describes its assimilation into the Roman Republic as a political rather than a demographic process, further evidenced by the maintenance of Etruscan culture and its language in the region for centuries (3).
By sharp contrast, all analyzed individuals from the Roman Imperial and Late Antique periods (1 to 500 CE) show a marked shift in ancestry toward populations of the eastern Mediterranean. While the strength of this shift might be influenced by the changing frequency of different burial practices—such as cremation and inhumation—among groups through time (43, 44), it clearly depicts the role of the Roman Empire in the large-scale displacement of people in a time of enhanced upward or downward socioeconomic and geographic mobility. In central Italy, including around Rome itself (17), the incoming ancestry detected so far mainly originated from the Near East rather than other areas of the Empire. The genetic replacement of ~50% of the preceding Etruscan-related gene pool was likely influenced by the movement of slaves and possibly soldiers, along with a larger pattern of human mobility from the eastern Mediterranean toward Italy (45–49). In the Roman Empire, citizenship was progressively extended to more classes of free people until the Edict of Caracalla in 212 made it universal among them, and expanding citizenship likely facilitated intermixture between local and other populations. Our new data from Etruria show that the influx of Near Eastern ancestry spread far beyond the greater capital region itself and suggest that this broader pattern of population movement may have affected larger portions of the Italian peninsula.
Continuing our genomic transect into the Early Middle Ages (500 to 1000 CE), we observe an additional genetic transition in some of the former Etruria territories through the spread of northern European–related ancestries. Admixture models are consistent with this genetic component being introduced from previously published individuals associated with the Longobard culture, although other cultural groups may have contributed as well. Thus, settlers expanding across large parts of the Italian peninsula after the collapse of the Western Roman Empire and the establishment of the Longobard Kingdom might have left a traceable impact on the genetic landscape of central Italy. Last, our analyses identify broad population continuity between the Early Medieval times and today in the regions of Tuscany, Lazio, and Basilicata, suggesting that the main gene pool of present-day people from central and southern Italy was largely formed at least 1000 years ago.
In conclusion, our study sheds light on five main aspects of Italian population history. First, individuals associated with the Etruscan culture carried a high proportion of steppe-related ancestry, despite speaking a non–Indo-European language. If the Etruscan language was indeed a relict language that predated Bronze Age expansions, then it would represent one of the rare examples of language continuity despite extensive genetic discontinuity (24, 50). The steppe-related ancestry in Etruscans may have been mediated by Bronze Age Italic speakers, possibly through a prolonged admixture process resulting in a partial language shift. Second, after the Bronze Age admixture, the Etruscan-related gene pool remained generally homogeneous for almost 800 years, notwithstanding the sporadic presence of individuals of likely Near Eastern, northern African, and central European origins. Third, eastern Mediterranean ancestries replaced a large portion of the Etruscan-related genetic profile during the Roman Imperial period. Fourth, a substantial genetic input from northern European ancestries was introduced during the Early Middle Ages, possibly through the spread of Germanic tribes into the Italian peninsula. Fifth, the genetic makeup of present-day populations from central and southern Italy was mostly in place by the end of the first millennium CE.
Although a broader geographic analysis of aDNA across Italy would be needed to substantiate the above conclusions, the observation of extremely similar ancestry shifts in Tuscany and northern Lazio to those reported for the city of Rome and its surroundings implies that historical events during the first millennium CE determined large-scale genetic transformations over an extended portion of the Italian peninsula. An analogous turnover is observed in Iberia between Iron Age and modern-day populations (24). This suggests that the Roman Empire might have left a long-lasting demographic contribution to the genetic profile of southern Europeans, bridging the gap between European and Near Eastern populations on the genetic map of western Eurasia. Additional archeogenetic datasets from other regions of the Empire will be pivotal to better pinpoint the genetic origins of incoming groups and to discern regionally specific patterns of admixture.
|
|




