Post by Admin on Dec 21, 2021 20:06:24 GMT
Results and discussion
We assembled available sequences of C, D and FT lineages from worldwide surveys ensuring that common lineages were represented (Bergstrom et al. 2020; Karmin et al. 2015; Mallick et al. 2016; Meyer et al. 2012; Poznik et al. 2016), and supplemented them with additional sequences from known rare lineages potentially relevant to early divergences, specifically, Australian C (Bergstrom et al. 2016; Mallick et al. 2016), West African D0 (Haber et al. 2019), Andamanese D (Mondal et al. 2017), and F chromosomes from China (Mallick et al. 2016), Vietnam (Poznik et al. 2016) and Singapore (Wong et al. 2013): 1204 sequences in all. We then focussed on the phylogenetic structure of the early divergences within these three lineages, and their geographical distributions revealed by ancient DNA and present-day analyses.
The resulting Y-chromosomal tree (Fig. 1) depicts 50 lineages, with the African lineages (gold) represented only by the four major African haplogroups without including their subsequent branches, but with the non-African lineages represented more fully to include all those originating before 45,000 years ago and found in the sample of present-day Y chromosomes examined, together with some of the more abundant recent lineages. As expected from previous analyses, this phylogeny shows that the three initial lineages C, D and FT each underwent initial rapid expansions soon after 54,000 (95% highest posterior density [HPD], 44,400–64,100) years, so that by 50,000 (95% HPD, 43,700–64,100) years ago there were seven branches within C, 5 within D and 18 within FT (30 non-African lineages in all); by 45,000 (95% HPD, 40,200–64,100) years ago the number of branches within FT had increased to 24 (36 in all). The branching patterns, together with the present-day locations of the lineages derived from an analysis of 2319 sequences, provide insights into possible locations of the early expansions. Lineage C split into two, C1 and C2; C1 lineages are found today only in East, Southeast and South Asia plus Oceania, while C2 lineages are more widespread and are now found in East and South Asia and also North and Central/West Asia (Fig. 2, Supplementary Fig. 1). D lineages are entirely confined to East and Southeast Asia. FT lineages now have a worldwide distribution, but the earliest split was into F and GHIJK; F is known only from East and Southeast Asia (Fig. 1, Supplementary Fig. 1), while GHIJK and its descendants are found worldwide. These descendant lineages themselves often have more continent-specific distributions, but 14/15 GHIJK lineages originating before 50,000 (95% HPD, 43,700–63,300) years ago have distributions that include East, Southeast or South Asia, apart from a few that are specific to Oceania (Fig. 1). Only one (H2, represented by a single sample) is specific to Europe, and none to the region adjoining the likely exit routes from Africa, in the terminology used Central/West Asia, where less than half are now present in the samples examined.
Fig. 2
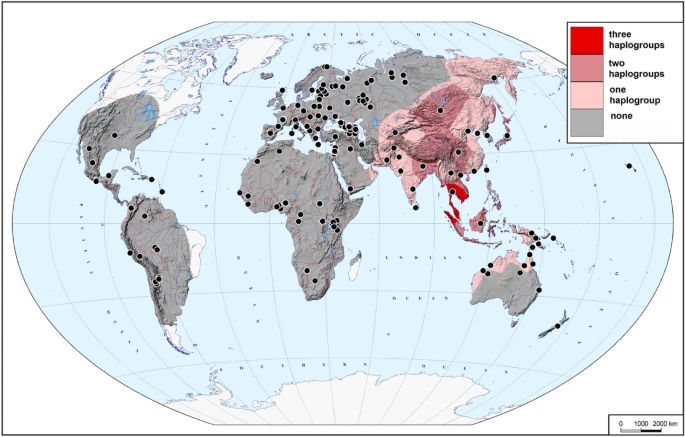
Presence of haplogroups C, D and F in 2302 present-day samples. The map demonstrates how many of the three haplogroups of interest (none, one, two, or all three) were found in different areas of the Old World and Near Oceania. Black dots indicate the locations of the studied populations
No ancient Y-chromosomal data earlier than 45,000 years ago have been reported, but 21 Asian or European males living 30,000–45,000 years ago are documented, and for 18 of them assignments to C, D or FT have been reported (Fig. 1, Supplementary Fig. 1, 2, Supplementary Table 1) (Fu et al. 2014, 2015, 2016; Seguin-Orlando et al. 2014; Sikora et al. 2017, 2019; Yang et al. 2017). Ten belong to the C lineage, six from North Asia and four from Europe. The remaining eight belong to FT, three from North Asia, one from East Asia and four from Europe. Although the data are limited, two conclusions can be drawn. First, none of the ancient samples carry Y lineages outside the 30 represented in Fig. 1 at 50,000 years ago. Second, C lineages (both C1a and C1b), now confined to East, Southeast and South Asia plus Oceania, were more widespread 30,000–40,000 years ago, including in Europe where they persisted until after 8000 years ago (Mathieson et al. 2018), although they have now been replaced in Europe by other lineages.
In a simple model of gradual human expansion from Africa to Asia and Oceania without subsequent continental-scale reshaping, we would expect the initial divergences in the Y-chromosomal phylogeny to have occurred in geographical locations close to Africa, and the present-day Y-chromosomal phylogeography to reflect this history by showing the presence of the early-diverging lineages within C, D and FT now being located geographically in Central/West Asia (Fig. 3a), with lower lineage diversity further east. In stark contrast, the observed distributions of these lineages all lie further to the east, suggesting that a simple model of this kind cannot explain the observed present-day data (Fig. 3b, Supplementary Fig. 3), a discrepancy we discuss further below.
Fig. 3
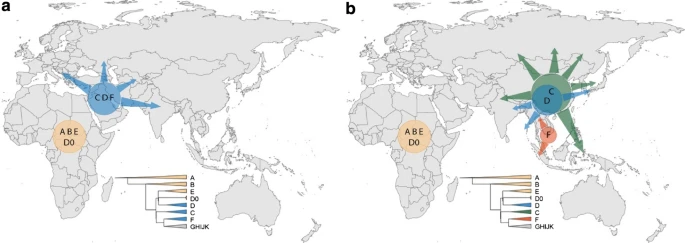
According to serial founder model, the earliest-branching non-African lineages are expected to expand and be present closer to Africa (a), but instead have expanded in East or Southeast Asia (b). Simplified Y tree is shown as reference for colours
The phylogeny of maternally inherited mitochondrial DNA (mtDNA), like that of the Y chromosome, also retains information from 50,000 to 70,000 years ago, although female-specific and with less detail because of its shorter length. Nevertheless, it provides a useful comparison. Outside Africa, the initial split inferred from a combination of ancient and present-day sequences was between lineages M and pre-N, with divergence within M dated to 44,000–55,000 years ago and within N to 47,000–55,000 years ago (Posth et al. 2016). Present-day geographical distributions of mtDNAs are less specific than Y chromosomes, and both of these major lineages are widespread worldwide, although M is absent from present-day Europeans with the exception of recent migrations (Gonzalez et al. 2007). Nevertheless, M was present in early Europeans until at least 28,000 years ago; moreover, the first branch within the pre-N/N lineage is between the pre-N mtDNA carried by Oase1 from Romania dating to 40,000 years ago (who, incidentally, showed increased Neanderthal admixture) and the remaining worldwide N mtDNAs. mtDNA thus shares with the Y chromosome a history of continental-scale change (loss of M from Europe), in the case of mtDNA dated to after 28,000 years ago. In addition, mtDNA N demonstrates the phylogeographic pattern expected from a simple expansion model, with its earliest divergence in the west.
How then can the present-day Y-chromosomal phylogeography be reconciled with an out-of-Africa expansion? It is well established that all known present-day Y-chromosomal lineages trace back to Africa at some point in human history (Jobling and Tyler-Smith 2017), but the current work demonstrates that the deepest rooting C, D and FT lineages now seen outside Africa are found in East/Southeast Asia. Without support from additional ancient DNA samples, it is difficult to make claims about the geographic origins of these deep-rooting lineages; however, this difficulty does not change the observation about their current location. The default explanation for the observed patterns is perhaps that the initial divergences within the Y-chromosomal phylogeny did indeed occur in the west, but that the deepest rooting lineages have now been lost from this part of the world, consistent with the lack of genetic continuity in West Eurasia seen in autosomal aDNA and the presence of Y haplogroup C lineages in West Eurasia until ~ 8000 years ago (Mathieson et al. 2018). In principle, this could be because C, D and F lineages all migrated east, together with some GHIJK lineages, leaving only GHIJK lineages in the west; or more plausibly that C, D and F were lost by genetic drift in the west, but not in the east. The first scenario would imply unprecedented levels of male-structured migration, and would be difficult to reconcile with subsequent divergences within GHIJK during the next few thousand years, whereby some of the descendent lineages such as G1, H1 and H3 would also need to have migrated east in a male-structured way. The second scenario is not easy to reconcile in a simple way with the inference that genetic effective population sizes have been lower in East Asia than in Europe (Gutenkunst et al. 2009; Kelleher et al. 2019), so less genetic drift is expected in the west. Further explanations should, therefore, also be considered; one such is that initial western Y chromosomes have been entirely replaced by lineages from further east (Fig. 3), perhaps on more than one occasion. This is supported by the observed patterns of early-diverging lineages of C, D and FT now being located in East and Southeast Asia, and, according to our present-day dataset of surviving lineages, the more likely origin of GHIJK in the east (Fig. 1). Formally, another explanation could be that selection has acted, for example, to favour the FT lineage to different extents in different regions, but positive natural selection has not been documented on the human Y chromosome (Jobling and Tyler-Smith 2017) and there are no candidate coding variants reported among annotated protein-coding genes (Poznik et al. 2016), so this seems unlikely. Nevertheless, the possible explanations for observed patterns cannot be reliably differentiated at present. Until aDNA data earlier than 45,000 years ago are available, future studies using spatial simulations with models that are able to adequately capture the complexity of the human past may help to explain the observed patterns in the present-day human Y-chromosomal data.
Ancient DNA studies are beginning to show some of the true complexity of human genetic history, including providing evidence for large-scale intercontinental movements in the last 30,000 years or so (Fu et al. 2014, 2015, 2016; Seguin-Orlando et al. 2014; Sikora et al. 2017, 2019; Yang et al. 2017). The out-of-Africa model requires major intercontinental movements 40,000–60,000 years ago, as well as later expansion into the Americas. From these perspectives, it is perhaps more likely that large-scale movements have continued throughout human prehistory than not, and replacement from the east is thus an explanation to consider. Ultimately, the prehistory of this period must encompass fossil, archaeological and multiple forms of genetic data, and reconcile them into a coherent overall understanding. The unique genetic properties of the Y chromosome may offer insights into movement during an early period that is currently difficult to investigate in other ways and provide a glimpse of this prehistory.
We assembled available sequences of C, D and FT lineages from worldwide surveys ensuring that common lineages were represented (Bergstrom et al. 2020; Karmin et al. 2015; Mallick et al. 2016; Meyer et al. 2012; Poznik et al. 2016), and supplemented them with additional sequences from known rare lineages potentially relevant to early divergences, specifically, Australian C (Bergstrom et al. 2016; Mallick et al. 2016), West African D0 (Haber et al. 2019), Andamanese D (Mondal et al. 2017), and F chromosomes from China (Mallick et al. 2016), Vietnam (Poznik et al. 2016) and Singapore (Wong et al. 2013): 1204 sequences in all. We then focussed on the phylogenetic structure of the early divergences within these three lineages, and their geographical distributions revealed by ancient DNA and present-day analyses.
The resulting Y-chromosomal tree (Fig. 1) depicts 50 lineages, with the African lineages (gold) represented only by the four major African haplogroups without including their subsequent branches, but with the non-African lineages represented more fully to include all those originating before 45,000 years ago and found in the sample of present-day Y chromosomes examined, together with some of the more abundant recent lineages. As expected from previous analyses, this phylogeny shows that the three initial lineages C, D and FT each underwent initial rapid expansions soon after 54,000 (95% highest posterior density [HPD], 44,400–64,100) years, so that by 50,000 (95% HPD, 43,700–64,100) years ago there were seven branches within C, 5 within D and 18 within FT (30 non-African lineages in all); by 45,000 (95% HPD, 40,200–64,100) years ago the number of branches within FT had increased to 24 (36 in all). The branching patterns, together with the present-day locations of the lineages derived from an analysis of 2319 sequences, provide insights into possible locations of the early expansions. Lineage C split into two, C1 and C2; C1 lineages are found today only in East, Southeast and South Asia plus Oceania, while C2 lineages are more widespread and are now found in East and South Asia and also North and Central/West Asia (Fig. 2, Supplementary Fig. 1). D lineages are entirely confined to East and Southeast Asia. FT lineages now have a worldwide distribution, but the earliest split was into F and GHIJK; F is known only from East and Southeast Asia (Fig. 1, Supplementary Fig. 1), while GHIJK and its descendants are found worldwide. These descendant lineages themselves often have more continent-specific distributions, but 14/15 GHIJK lineages originating before 50,000 (95% HPD, 43,700–63,300) years ago have distributions that include East, Southeast or South Asia, apart from a few that are specific to Oceania (Fig. 1). Only one (H2, represented by a single sample) is specific to Europe, and none to the region adjoining the likely exit routes from Africa, in the terminology used Central/West Asia, where less than half are now present in the samples examined.
Fig. 2

Presence of haplogroups C, D and F in 2302 present-day samples. The map demonstrates how many of the three haplogroups of interest (none, one, two, or all three) were found in different areas of the Old World and Near Oceania. Black dots indicate the locations of the studied populations
No ancient Y-chromosomal data earlier than 45,000 years ago have been reported, but 21 Asian or European males living 30,000–45,000 years ago are documented, and for 18 of them assignments to C, D or FT have been reported (Fig. 1, Supplementary Fig. 1, 2, Supplementary Table 1) (Fu et al. 2014, 2015, 2016; Seguin-Orlando et al. 2014; Sikora et al. 2017, 2019; Yang et al. 2017). Ten belong to the C lineage, six from North Asia and four from Europe. The remaining eight belong to FT, three from North Asia, one from East Asia and four from Europe. Although the data are limited, two conclusions can be drawn. First, none of the ancient samples carry Y lineages outside the 30 represented in Fig. 1 at 50,000 years ago. Second, C lineages (both C1a and C1b), now confined to East, Southeast and South Asia plus Oceania, were more widespread 30,000–40,000 years ago, including in Europe where they persisted until after 8000 years ago (Mathieson et al. 2018), although they have now been replaced in Europe by other lineages.
In a simple model of gradual human expansion from Africa to Asia and Oceania without subsequent continental-scale reshaping, we would expect the initial divergences in the Y-chromosomal phylogeny to have occurred in geographical locations close to Africa, and the present-day Y-chromosomal phylogeography to reflect this history by showing the presence of the early-diverging lineages within C, D and FT now being located geographically in Central/West Asia (Fig. 3a), with lower lineage diversity further east. In stark contrast, the observed distributions of these lineages all lie further to the east, suggesting that a simple model of this kind cannot explain the observed present-day data (Fig. 3b, Supplementary Fig. 3), a discrepancy we discuss further below.
Fig. 3

According to serial founder model, the earliest-branching non-African lineages are expected to expand and be present closer to Africa (a), but instead have expanded in East or Southeast Asia (b). Simplified Y tree is shown as reference for colours
The phylogeny of maternally inherited mitochondrial DNA (mtDNA), like that of the Y chromosome, also retains information from 50,000 to 70,000 years ago, although female-specific and with less detail because of its shorter length. Nevertheless, it provides a useful comparison. Outside Africa, the initial split inferred from a combination of ancient and present-day sequences was between lineages M and pre-N, with divergence within M dated to 44,000–55,000 years ago and within N to 47,000–55,000 years ago (Posth et al. 2016). Present-day geographical distributions of mtDNAs are less specific than Y chromosomes, and both of these major lineages are widespread worldwide, although M is absent from present-day Europeans with the exception of recent migrations (Gonzalez et al. 2007). Nevertheless, M was present in early Europeans until at least 28,000 years ago; moreover, the first branch within the pre-N/N lineage is between the pre-N mtDNA carried by Oase1 from Romania dating to 40,000 years ago (who, incidentally, showed increased Neanderthal admixture) and the remaining worldwide N mtDNAs. mtDNA thus shares with the Y chromosome a history of continental-scale change (loss of M from Europe), in the case of mtDNA dated to after 28,000 years ago. In addition, mtDNA N demonstrates the phylogeographic pattern expected from a simple expansion model, with its earliest divergence in the west.
How then can the present-day Y-chromosomal phylogeography be reconciled with an out-of-Africa expansion? It is well established that all known present-day Y-chromosomal lineages trace back to Africa at some point in human history (Jobling and Tyler-Smith 2017), but the current work demonstrates that the deepest rooting C, D and FT lineages now seen outside Africa are found in East/Southeast Asia. Without support from additional ancient DNA samples, it is difficult to make claims about the geographic origins of these deep-rooting lineages; however, this difficulty does not change the observation about their current location. The default explanation for the observed patterns is perhaps that the initial divergences within the Y-chromosomal phylogeny did indeed occur in the west, but that the deepest rooting lineages have now been lost from this part of the world, consistent with the lack of genetic continuity in West Eurasia seen in autosomal aDNA and the presence of Y haplogroup C lineages in West Eurasia until ~ 8000 years ago (Mathieson et al. 2018). In principle, this could be because C, D and F lineages all migrated east, together with some GHIJK lineages, leaving only GHIJK lineages in the west; or more plausibly that C, D and F were lost by genetic drift in the west, but not in the east. The first scenario would imply unprecedented levels of male-structured migration, and would be difficult to reconcile with subsequent divergences within GHIJK during the next few thousand years, whereby some of the descendent lineages such as G1, H1 and H3 would also need to have migrated east in a male-structured way. The second scenario is not easy to reconcile in a simple way with the inference that genetic effective population sizes have been lower in East Asia than in Europe (Gutenkunst et al. 2009; Kelleher et al. 2019), so less genetic drift is expected in the west. Further explanations should, therefore, also be considered; one such is that initial western Y chromosomes have been entirely replaced by lineages from further east (Fig. 3), perhaps on more than one occasion. This is supported by the observed patterns of early-diverging lineages of C, D and FT now being located in East and Southeast Asia, and, according to our present-day dataset of surviving lineages, the more likely origin of GHIJK in the east (Fig. 1). Formally, another explanation could be that selection has acted, for example, to favour the FT lineage to different extents in different regions, but positive natural selection has not been documented on the human Y chromosome (Jobling and Tyler-Smith 2017) and there are no candidate coding variants reported among annotated protein-coding genes (Poznik et al. 2016), so this seems unlikely. Nevertheless, the possible explanations for observed patterns cannot be reliably differentiated at present. Until aDNA data earlier than 45,000 years ago are available, future studies using spatial simulations with models that are able to adequately capture the complexity of the human past may help to explain the observed patterns in the present-day human Y-chromosomal data.
Ancient DNA studies are beginning to show some of the true complexity of human genetic history, including providing evidence for large-scale intercontinental movements in the last 30,000 years or so (Fu et al. 2014, 2015, 2016; Seguin-Orlando et al. 2014; Sikora et al. 2017, 2019; Yang et al. 2017). The out-of-Africa model requires major intercontinental movements 40,000–60,000 years ago, as well as later expansion into the Americas. From these perspectives, it is perhaps more likely that large-scale movements have continued throughout human prehistory than not, and replacement from the east is thus an explanation to consider. Ultimately, the prehistory of this period must encompass fossil, archaeological and multiple forms of genetic data, and reconcile them into a coherent overall understanding. The unique genetic properties of the Y chromosome may offer insights into movement during an early period that is currently difficult to investigate in other ways and provide a glimpse of this prehistory.
