|
|
Post by Admin on Feb 8, 2022 20:20:21 GMT
Uniparental genetics and kinship analyses
Both Bk-II and Bk-III show phylogeographic signals for their maternal and paternal lineages. Accordingly,
Bk-II is mostly defined by mtDNA connections to the region of present-day Poland and its surroundings,
whereas Bk-III has more diverse maternal composition, see Supplementary Information section 2.1. Male
lineages in both Bk-II and Bk-III are mostly defined by Y chromosome haplogroup I2a-L1229 (Table 1), for
which network analysis (Supplementary Information section 2.2) narrowed down regional affinities to the
North European plain and shows continuity between these two horizons. Uniparental diversity makeup points
to a patriarchal social structure similar to previously reported Bronze Age findings9,11,15. Results are highly
similar to previous observations on Encrusted Pottery culture at the Jagodnjak site (Croatia)10. The kinship
network of Bk-II follows the distribution of grave groups (Fig. 1) which were likely established along family
relationships and chronology. Individuals buried in the Bk-III mass grave only show a few blood relations, like
a half-brother and father-son and a dizygotic twin, the latter is the most archaic detection to date to our
knowledge. However, Bk-III as an extended family group can not be excluded. For further details, see
Supplementary Information section 2 and Supplementary Tables S1-S3.
Table 1 Summary of the investigated samples. MtDNA and ChrY denote mitochondrial haplogroup and Y
chromosome haplogroup. In column “Kinship” 1st and 2nd mean the degree relations. For the feature, grave
IDs and details on newly reported radiocarbon dates see the Supplementary Table S1.
Genetic disorders and pigmentation
Investigating genetic disorders in archaic datasets is potentially valuable for history of health and medicine,
and also highlights the overall genetic health of past populations. Inherited genetic disorders, if accompanied
with severe phenotypic anomalies, could also explain unusual burial practises, as it was described in cases
of dwarfism16. For detailed results of this topic, see Supplementary Information section 3.
Aneuploidies
The abnormal number of chromosomes result in a few well known diseases which we tested thoroughly. We
found one individual, S10, the only child burial in Bk-II, with XYY gonosomal genotype, described as Jacob’s
syndrome. Although this syndrome remains in most cases silent as it is relatively frequent (~0.1%) in today’s
populations, it occasionally comes with a wide scale of symptoms17, which may be linked to its separate
burial, but due to poor bone preservation for S10 no further assessments could be made.
Mitochondrial DNA diseases
We examined the clinical significances of the polymorphisms that can be found in the mtDNA by using the
mitopathotool software on the AmtDB database18, and found that individual S1 (40+ years old) from Bk-II had
one of the defining mutations (T14484C) of Leber’s hereditary optic neuropathy (LHON) causing complete
vision loss in ~10% of females and in ~50% of males between 20-40 years of age, rarely accompanying with
other neuropathies19.
Pigmentation
Pigmentation patterns highly differ between horizons, as Bk-I mostly possess variants for light pigmentation,
blue eyes and blonde hair, while Bk-II is more similar to populations of Neolithic Europe (Fig. 2), although
some variants for lighter pigmentation exist within this group too. Members of Bk-III on the other hand show a
wide range from dark to light tones and even the presence of variants for red hair (Supplementary Table S5,
Supplementary Information section 3.2.1).
 |
|
|
|
Post by Admin on Feb 8, 2022 21:47:33 GMT
Nuclear variants with clinical significance
We also examined the nuclear genomes to find regions with clinical significance. Since a complete panel for
determining disease susceptibility only exists in commercial DNA kits, and detailed description for the 1240k
SNP set is not available, we created our own SNP calling panel (included in PAPline) focusing on various
conditions including amyotrophic lateral sclerosis, Alzheimer disease, autism, Crohn’s disease, diabetes,
lactose intolerance, tumor markers, mental disorders, Parkinson disease, schizophrenia and ulcerative
colitis. For this study we used 3,874 clinically significant SNPs, which were marked as “pathogenic” or “likely
pathogenic” in the ClinVar database20, by ignoring deletion, duplication and copy number variants, as well as
SNPs with questionable (“reported”, “conflicting reports”, etc.) contribution to diseases. From this set we
found ~2,200 SNPs which covered at least one individual, out of these 27 positions showed clinically
relevant substitutions. However, test runs on database data resulted in numerous positive hits for pathogenic
variants most likely related to DNA damage, which highlights the unreliability of low coverage SNP data for
variant identification (Supplementary Information section 3.2.2). To overcome this issue, we considered
positions with more than 0.99 genotype likelihood (GL) values calculated using ANGSD v0.93121
(Supplementary Table S6) or when skeletal features supported results. A variant for Lig4 syndrome
(rs10489442122, GL=0.999) in individual S15 was detected, and some of his skeletal features (e.g. congenital
hip dysplasia) show possible onset of symptoms (Supplementary Information section 3.2.2.1). In another
case the physical manifestation of hereditary spastic paraplegia is likely for S11 and S6, father and son but
the genotype likelihood is lower (0.67; see Supplementary Information section 3.2.2.3). Interestingly, a tumor
marker on the BRCA2 gene (rs80358920, GL=0.999) in individual S9 is nowadays only prevalent in Asian
populations22. For further discussions, see Supplementary Information section 3.2.
Whole genome composition and genetic ancestry
Balatonkeresztúr site samples
According to the principal component analysis (PCA) based on 590k nuclear SNPs (Fig. 3 a.) Bk-I is clearly
separated from Bk-II and Bk-III, where Bk-II has a strong shift towards HG samples23 overlapping with only a
fraction of known archaic samples23 and Bk-III. Admixture and qpAdm analyses for assessing genetic
components (Supplementary Tables S9, S12-16; Supplementary Information sections 5.2 and 5.5.2)
revealed ~17% HG, ~40% European farmer, and ~43% steppe ancestry for Bk-I, similar to average Bronze
Age Europeans. Bk-I is most likely derived from a single source that is genetically related to a Poland
Southeast Bell Beaker culture (BBC) associated population (p = 0.784) in line with archaeological
observations24. Bk-II comprises a unique makeup of ~42% HG, ~41% European farmer, and ~17% steppe
ancestries. qpAdm analysis revealed most plausible sources of Neolithic Sweden Funnel Beaker culture
(~32±8%), Poland Southeast BBC (~41±6%) and an extra HG (~29±3%) ancestry of yet unknown origin.
Despite the lower supported models Bk-I can not be excluded as an ancestry component for Bk-II, while the
affinity of Sweden Funnel Beaker culture associated population likely reflects a more closely related group,
such as population related to the Poland Globular Amphora culture, see Supplementary Information section
5.5.2.2 and Supplementary Tables S12 and S15. Bk-III shows a shift in ancestry composition (~29% HG,
~46% European farmer, ~25% steppe). qpAdm analyses revealed that the main ancestry component for
Bk-III is Bk-II (~53±5%), while “dilution” of Bk-II to Bk-III is mostly driven by population events that are yet to
be uncovered.
Genetic outliers from previous studies
Many samples were defined as genetic outliers in their genetic context by previous studies from Bronze Age
Europe. We selected such outlier individuals with high HG ancestry components to assess whether they are
related to Bk-II. Selection was based on previous observations and also by using Dixon’s Q-test25 at 90%
confidence interval on HG component’s upper deviation using results of the Admixture analysis. First, we ran
f4-statistics in the form of f4(W=test outlier, X=corresponding population, Y=Bk-II, Z=Yoruba)26. This test
resulted in positive values for some outliers (W) meaning that these are genetically closer to Bk-II (Y) than its
presumed population (X). However, Z-scores are low in many cases, and false positives may appear solely
by high HG component, not by true relationship (see Supplementary Table S10 and Supplementary
Information section 5.3). To check true relationship between Bk-II and groups/samples with high HG
ancestry, we performed an outgroup f3-statistics in the form of f3(X=Bk-II, Y=test HG-s, Z=Yoruba) for all
relevant archaic populations and outliers23 (Supplementary Table S11) that resulted a table of allele
frequency based distances between test pairs Xs and Ys. Euclidean clustering based on the results of
f3-statistics revealed that a number of samples and even three populations from the Baltic (Fig. 3 b,
Supplementary Information section 5.4) from AADR23 form a cluster with Bk-II and Bk-III, suggesting actual
genetic connection via a common HG ancestry source.
|
|
|
|
Post by Admin on Feb 9, 2022 2:52:10 GMT

Origin of high HG ancestry component in BK-II and the Kisapostag population
f4-statistics in the form of f4(W=Sweden HG, X=Serbia IronGates HG, Y=Bk-II, Z=Yoruba) revealed that
while Pitted Ware culture associated individuals have similarly high HG levels to Bk-II from Scandinavian
HG-s, in Bk-II this type of HG component is significantly negative meaning no or minimal shared ancestry
(Supplementary Information section 5.3). Results of outgroup in f3-statistics in the form of f3(X=Bk-II,
Y=HG-s, Z=Yoruba) showed that HGs fit the best to Bk-II are geographically widely distributed
(Supplementary Table S11), thus we conclude that the true source for this component is yet to be described.
We can take into consideration the outgroup f3-statistics results, chronology, timing of HG admixture
according to Freilich 202110 between ~3400-2400 BCE, qpAdm results and the geographical distribution of
groups and outliers of similar HG makeup. These suggest a dated migration pattern for this undescribed
population with dominant HG genetic ancestry from what is present-day Bulgaria to the Baltic through the
Eastern borders of the Carpathians (Fig. 3 b). Ancestors of Bk-II likely branched off from this migration route
and started to move towards West, by at least around ~2500 BCE, subsequently intermixing with various
groups. Interestingly, the phylogeography of mitochondrial haplogroup U4b1b1 (individual S15) perfectly fits
this scenario. Notably, strontium (87Sr/86Sr ratio) isotope results from molars (Fig. 4, Supplementary
Information section 1.9) indicate that every individual of this study was raised and lived at least close
proximity to Balatonkeresztúr site in their childhood and early adolescence, suggesting that Bk-II group does
not represent the first generation of the Kisapostag culture associated population in Transdanubia (Western
Hungary). Correspondingly, the high number of supported models in qpAdm results could be the result of
subsequent local admixtures.
|
|
|
|
Post by Admin on Feb 10, 2022 19:55:25 GMT
A Late Copper Age outlier individual from Balatonlelle site
We included a Late Copper Age individual from Balatonlelle site to this study for its high HG genomic
ancestry component. Mitogenome of BAD002 (K1a4a1) shows affinity to Iberian BBC associated individuals
(Supplementary Information Fig. S.2.1.1), and his Y chromosomal haplogroup belongs to I-M170. On
genomic PCA this sample groups with Iberian and French Neolithic individuals, due to higher HG component
(~34%) than known Neolithic and Copper Age populations in the Carpathian Basin have, and due to the lack
of steppe related ancestry. qpAdm estimates pointed to a source of Neolithic communities from present-day
(Northwestern) France (87±8%)27 with a minor extra HG component (13±8%) with p=0.948 (Supplementary
Information section 5.5, Supplementary Tables S12 and S13). Pigmentation pattern of BAD002 shows
resemblance to average Neolithic Europeans. The foreign cultural traits of the boy’s jewellery is in line with
his outlier genetic composition in the study region28. Notably, further tests (outgroup f3-statistics, qpAdm)
excluded contribution nor any connection of BAD002 to Bk-II (Supplementary Tables S11, S12 and S13,
Supplementary Information section 5). Therefore we conclude that this individual testifies large-scale
migration in the Copper Age, providing research questions for future studies.
Discussion
The Carpathian Basin was inhabited by the Baden cultures’ population at the end of the Copper Age, and
their genetic composition was represented by an early farmer and a slightly increased HG genetic
component, compared to the previous Neolithic populations of the region6. Here we demonstrated that in the early phase of this culture, a group of Western European origin appeared in Transdanubia, diversifying what we knew about the region’s Late Copper Age substrate up to now.
The Carpathian Basin experienced the influx of steppe-related genetic ancestry at the dawn of the Bronze
Age5,8, and this transformation was already detectable at Balatonkeresztúr-Réti-dűlő site as well, where we
could examine multiple populations. The earliest Bronze Age horizon Bk-I (representative of the
Somogyvár-Vinkovci culture) shows similarities to Poland Southeast BBC associated population with high
steppe ancestry that was replaced by the Kisapostag culture associated Bk-II around the 23-22th century
BCE, while at least some sort of genetic ancestry of Bk-I in this population can not be excluded. According to
our results, the Bk-II population had outstandingly high HG genetic ancestry levels, compared to other
Bronze Age groups of the region, which can be traced back to today’s Ukraine, Belarus, Moldavia or
Romania, pointing to a long standing previously unsampled population with dominant HG ancestry.
Calculated admixture dates10 suggest the presence of a genetically pure or at least highly HG specific
population in Eastern Europe as late as the end of the Copper Age. Part of this group subsequently admixed
with populations of mainly steppe (likely Poland Southeast BBC) and early farmer (most likely a Globular
Amphora culture related) ancestry during their westward migration on a Northern route, leaving genetic
traces in Corded Ware culture, BBC, and other Bronze Age populations. The paternal lineage of BK-II was
likely linked to the farmer component, as I2a-M223 (upstream to I2a-L1229) was a frequent paternal lineage
among Globular Amphora culture and related populations. Looking for the possible source areas of the
Kisapostag culture, a number of archaeological theories need to be considered. The pottery decoration
technique originated either from Corded Ware in the Middle Dnieper region (Ukraine), epi-Corded Ware
groups (northern Carpathians), e.g. Chłopice-Veselé or Nitra groups (Slovakia), the latter two is also
supported by inhumation practises31–36. However, connections with the Litzenkeramik or
Guntramsdorf-Drassburg group (eastern Austria, Slovenia, western Croatia) were also raised37,38. Pottery
forms were connected to local development of communities with eastern (Makó–Kosihy–Čaka) or southern
(Somogyvár–Vinkovci) origins, too39. BBC influence was also mentioned based on connections of pottery
and craniometry data (so called Glockenbecher or brachycran skull type40–42). The results of this study fit best
with the Middle Dnieper area origin of BK-II, especially when we consider individual I4110 from Dereivka I
(Ukraine Eneolithic) as one of the earliest representatives of their genomic makeup.
Strontium isotope (87Sr/86Sr ratio) data, representing through nutrition the bioavailable Sr in the area where
people lived in a certain age interval, shows local values for both sexes in both Bk-II and Bk-III. These results
push back the timing of their arrival a few generations, meaning that local or southern impact of cultural traits
and maybe even genetic admixtures likely occurred during this short period as well, which also could explain
the culture’s archaeological heterogeneity.
The population of Bk-III was the direct descendant of Bk-II, forming not just cultural (Encrusted pottery) but
also genetic continuity for at least ~500 years, even if the radiocarbon sequences allow a few decades of
hiatus at the studied site. Continuous female-biased admixture with various groups occurred during this
period according to our and previous genetic10 and archaeological31,43 evidence, diluting the BK-II genetic
ancestry.
In both periods, the homogeneity of paternal lineages suggest a similar social organisation described in9,10 of
a patrilocal residence system. However, strontium isotope data shows local values for both sexes, which
along with similar genomic makeup of females and males suggest exogamy most probably between villages
of the same population. Two pairs of half-sibling graves in the two periods may indicate polygamy, although
remarriage for high female mortality is more plausible. Notably, almost none of the uniparental markers are
identical even at the haplogroup level with individuals from the Croatian Encrusted Pottery culture Jagodnjak
site, despite high similarities in cultural traits, social structure and genomic composition of the communities.
This points to clan-like or patriarchal superfamily structure of Kisapostag and Encrusted Pottery groups. The
relatively limited presence of female and children burials in both Bk-II and Bk-III periods may suggest
distinctive treatment or another (here undiscovered) burial group for women and children at the same site.
Although, in other Bronze Age cemeteries, e.g. Ordacsehi and Bonyhád in Hungary, males, females and
children were buried close to each other, suggesting high variance of burial practises34,35,44.
While low genomic coverage did not allow fine SNP recovery, we did find evidence for malignant variants
within all of our tested groups, and undoubtedly showed the presence of LHON and Jacob’s syndrome within
Bk-II. Additionally, the disease panel we created can be extended and used in future studies, providing
insight into past population health qualities.
Considering the unstructured age and kinship distribution in the mass grave Bk-III compared to Bk-II, the
coetaneous death of eight people at least, the absence of traumatic or ritual events on bones, and
non-cremated nature of the burial all signals a sudden tragic event in the Encrusted pottery period, most
likely an epidemic, as first suggested based on the anthropological analyses45. Interestingly, comparative
strontium isotope analyses on the first and third molar of the individuals in the BK-III mass grave indicate that
subadult males - including a severely disabled individual (S15) with hip dysplasia - left their community for a
while and then returned to their birthplace prior to their death, raising further questions for future studies on
prehistoric lifeways and social organisations.
|
|
|
|
Post by Admin on May 1, 2022 18:04:07 GMT
Genetic analysis of male Hungarian Conquerors: European and Asian paternal lineages of the conquering Hungarian tribes Erzsébet Fóthi, Angéla Gonzalez, Tibor Fehér, Ariana Gugora, Ábel Fóthi, Orsolya Biró & Christine Keyser Archaeological and Anthropological Sciences volume 12, Article number: 31 (2020) Abstract According to historical sources, ancient Hungarians were made up of seven allied tribes and the fragmented tribes that split off from the Khazars, and they arrived from the Eastern European steppes to conquer the Carpathian Basin at the end of the ninth century AD. Differentiating between the tribes is not possible based on archaeology or history, because the Hungarian Conqueror artifacts show uniformity in attire, weaponry, and warcraft. We used Y-STR and SNP analyses on male Hungarian Conqueror remains to determine the genetic source, composition of tribes, and kin of ancient Hungarians. The 19 male individuals paternally belong to 16 independent haplotypes and 7 haplogroups (C2, G2a, I2, J1, N3a, R1a, and R1b). The presence of the N3a haplogroup is interesting because it rarely appears among modern Hungarians (unlike in other Finno-Ugric-speaking peoples) but was found in 37.5% of the Hungarian Conquerors. This suggests that a part of the ancient Hungarians was of Ugric descent and that a significant portion spoke Hungarian. We compared our results with public databases and discovered that the Hungarian Conquerors originated from three distant territories of the Eurasian steppes, where different ethnicities joined them: Lake Baikal-Altai Mountains (Huns/Turkic peoples), Western Siberia-Southern Urals (Finno-Ugric peoples), and the Black Sea-Northern Caucasus (Caucasian and Eastern European peoples). As such, the ancient Hungarians conquered their homeland as an alliance of tribes, and they were the genetic relatives of Asiatic Huns, Finno-Ugric peoples, Caucasian peoples, and Slavs from the Eastern European steppes. Introduction According to the general belief, the Finno-Ugric-speaking Hungarians left Magna Hungaria, their original homeland in the Western Ural Mountains (i.e., present-day Bashkortostan), for Levedia, on the edge of the Khazar Khaganate, and then moved to Etelköz, an area between the Don and Danube Rivers. They conquered their current homeland in the Carpathian Basin in 895 CE and defeated the Moravian Kingdom led by Svatopluk in 902 CE (see Fig. 1). Fig. 1 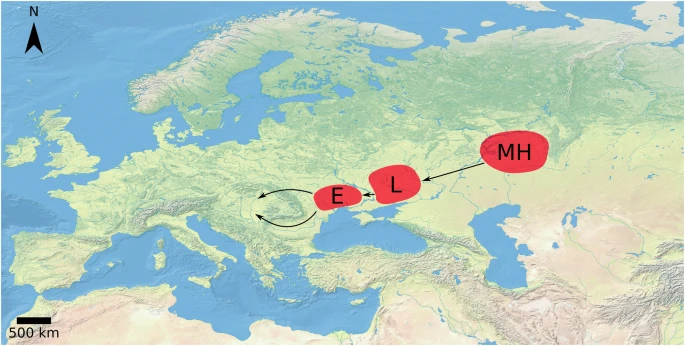 The nomadic ancient Hungarians from the Eastern European steppes consisted of an alliance of seven tribes, which was joined by the Kabars that had splintered off from the Khazars. At the end of the ninth century CE, following confrontations with the Turkic Pechenegs, Árpád led the tribes out of Etelköz to the Hungarians’ current home in the Carpathian Basin. Linguists agree that the Hungarian language is essentially Finno-Ugric, but it was also influenced by Iranian, Turkic, and Slavic languages. As a result, the currently accepted view among researchers is that the Hungarian Conquerors spoke a Finno-Ugric language but looked and lived like Turkic peoples (Róna-Tas 1995; Bálint 2006). The term “Hungarian Conquerors” is well-known and is traditionally applied in Hungarian scientific life from history to archaeology and anthropology, and is widely used in the field of archaeogenetics (Tömöry et al. 2007; Csányi et al. 2008; Csősz et al. 2016; Neparáczki et al. 2017, 2018). See Online Resource 1 (ESM_1) for more information on the origin stories, history, linguistics, and archaeology of the ancient Hungarians. Although several thousand Hungarian Conqueror graves have been discovered, containing rich skeletal and artifact assemblages, the origin of the ancient Hungarians has remained unclarified. Early anthropological research placed the ancient Hungarian homeland in the Volga-Kama region (Lipták 1983). Tóth (1981) and Éry (1983) theorized that the classic Hungarian Conquerors came from farther east and they placed the origin of most Asian elements of the Hungarian tribes in the Khazak steppes. Based on Fóthi’s (2014) interpretation, the Hungarian Conquerors had ties to the Northern Pontus and Volga-Kama regions during the migration period; however, the closest anthropological parallels were found among the Central Asian and Southern Siberian nomadic Turkic tribes from the 5th–7th centuries CE. The results of the first genetic tests increased the number of answerable questions. Based on analyses using classical markers, the modern Hungarian population is very similar to other European populations, especially Slavic- and Germanic-speaking ones, while the frequency of genetic variants typical around the Ural Mountains is only 13% (Guglielmino et al. 2000). Mitochondrial DNA population tests yielded similar results; the majority of the maternal genetic lineages were identical to those of the surrounding European peoples (Lahermo et al. 2004; Semino et al. 2000). The first archaic mtDNA analyses also supported this finding (Tömöry et al. 2007). Tambets et al. (2018) studied the autosomal genetic variation of Uralic-speaking populations and found that the recent gene pool of Hungarians did not form genetic clusters with other Uralic speakers but instead grouped with a wide range of geographically adjacent samples. Recent studies have dedicated greater attention to the so-called classic Hungarian Conqueror samples contrary to previous, random testing, which mostly included remains from the tenth century CE commoner cemeteries. A new archaic mtDNA analysis (Csősz et al. 2016) suggests that both Finno-Ugric roots and Central Asian Turkic influxes were possible genetic influences on the conquerors’ mixed genetic composition. Neparáczki et al. (2018) suggested that at least one-third of the Hungarian Conquerors’ maternal lineage is descended from Central-Inner Asia, most likely from the Asiatic Huns. Genetic testing of the modern Hungarian populations’ paternal lineage showed either no or very low frequency of the Y-chromosomal N-Tat haplogroup (Zerjal et al. 1997; Semino et al. 2000; Csányi et al. 2008; Fehér et al. 2015; Pamjav et al. 2017), which is dominant in all the other Finno-Ugric populations (Zerjal et al. 1997). On the other hand, it is noteworthy that in one study, the N-Tat (M46) Y-chromosomal marker was found in 50% of the investigated tenth century CE classic Hungarian Conquerors (Csányi et al. 2008). However, a recent study by Post et al. (2019) confirmed that a small number of N3a4-Z1936 lineages are shared not only by Hungarians and Mansi but also by the other Ob-Ugric group, Khanty. The exclusivity of this Ugric connection is that it is missing from other European populations and its non-Ugric appearance is restricted to the Volga-Ural region, notably Bashkirs and Tatars, corresponding to the territory of “Magna Hungaria” where proto-Magyars lived before the conquest of the Hungarian Basin and where Friar Julian found a surviving Hungarian-speaking group in the thirteenth century before the Mongol raids (Vásáry 2015). It is known that many nomadic tribes practiced (and still practice) patrilocal exogamy, in which the male members choose wives from other tribes. In that context, to study the genetic makeup of nomadic tribes, the analysis of maternal lineages is less meaningful than the paternal ones. For this reason, we conducted Y-chromosomal DNA analysis to determine the genetic origins of the Hungarian Conquerors. The purposes of the present study were (i) to determine the genetic makeup of the Hungarian Conquerors, (ii) to find out if there were any ancestral differences among the Hungarian Conqueror components, and (iii) to investigate whether the single Uralic homeland hypothesis stands, or other geographical regions might have played a role as a genetic source for some Hungarian tribes. |
|



