|
|
Post by Admin on Mar 11, 2022 20:20:09 GMT
Fig. 3: Admixture graph modeling for aMMD groups using qpGraph. 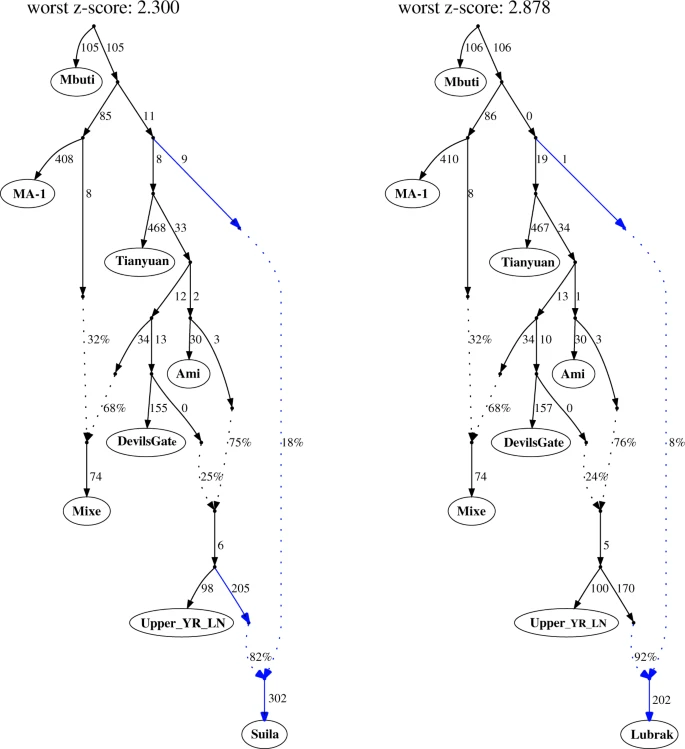 Although deriving 80–92% of their ancestry from a lineage related to Upper_YR_LN (Supplementary Table 4), the aMMD and present-day Sherpa/Tibetans are not adequately modeled as a sister clade to Upper_YR_LN, as expected given the unique genetic components of Tibetans not shared with lowlanders, including the EPAS1 allele from a Denisovan-related admixture. Rather, the remaining 8–20% of their ancestry derives from a deep part of the population graph near the split between Western and Eastern Eurasian branches (Fig. 3; Supplementary Fig. 9). This source, however, does not derive from archaic hominins (Neanderthals or Denisovans, who contribute <0.5% genome-wide ancestry), and our results reject previously suggested sources of gene flow into the Tibetan lineage13,35,36, including deeply branching Eastern Eurasian lineages, such as the 45,000-year-old Ust’-Ishim individual from southern Siberia, the 40,000-year-old Tianyuan individual from northern China, and Hoabinhian/Onge-related lineages in southeast Asia (Supplementary Fig. 10), suggesting instead that it represents yet another unsampled lineage within early Eurasian genetic diversity. This deep Eurasian lineage is likely to represent the Paleolithic genetic substratum of the Plateau populations. Two-routes of dispersal of the Tibeto-Burman speakers to the Himalayas The south-facing slopes of the Himalayas harbor many ethnolinguistic groups that show a striking pattern of stratification across altitudes: Indo-Iranian speaking South Asian populations occupy the lowlands, Sherpa/Tibetans occupy the highlands, and various non-Tibetan Tibeto-Burman speaking groups, such as the Tamang and Gurung, occupy the middle altitude range37,38. While Sherpa/Tibetans in Nepal likely arrived in the Himalayas from the Plateau (i.e., the northern route)39, a previous genetic study suggested a separate southern route of migration for the middle altitude Tibeto-Burman groups37. However, how non-Tibetan Tibeto-Burman speaking groups are related to each other and to the Tibetan lineage has remained unclear. Here we utilize Lubrak, the most representative ancient group within the Tibetan lineage, to investigate the genetic history of Tibeto-Burman speaking populations. Specifically, we model Sherpa/Tibetans and other Tibeto-Burman groups using Nepalese Tibetans from Tsum as one source and Upper_YR_LN/YR_MN as the other, while using Lubrak as a key outgroup to distinguish the Tibetan lineage from lowlander ancestries with high resolution. Consistent with a previous report18, we observe that the Tibetan groups from the Plateau and the Himalayas form a genetic cline. First, Nepalese Tibetans from the Mustang and Gorkha districts (Upper Mustang, Nubri, Tsum), which are cladal to each other, and Sherpa from the Solukhumbu district derive 87–92% of their ancestry from the Tibetan lineage, which is represented by Tsum (Fig. 4; Supplementary Table 5). Second, Tibetans relatively close to the Himalayas (e.g., Lhasa, Shigatse, Shannan) derive a major proportion of their ancestry from the Tibetan lineage (76–86%). Last, Tibetan groups further to the east or northeast have much higher contributions from the lowlander lineage (21–58%). While we know from radiocarbon dating that the two poles of this cline, represented by aMMD and Upper_YR_LN, were already present by ca. 1420 BCE, the admixture process between the two poles that formed the present-day cline may have occurred later. Additional archaeogenetic studies in the Plateau are needed to understand when the cline began to form and how it developed over time across the Plateau. |
|
|
|
Post by Admin on Mar 11, 2022 21:30:31 GMT
Fig. 4: A genetic cline of Tibeto-Burman groups. 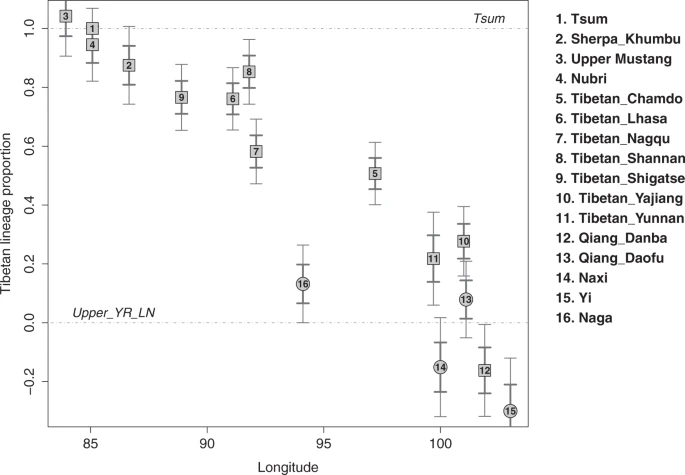 We model Tibeto-Burman groups using Nepalese Tibetan from the Tsum region (“Tsum”) and Upper_YR_LN as the two sources using qpAdm. Tibetans from the plateau and Tibetans close to the Himalayas derived the majority of their ancestry from the Tibetan lineage, while Tibeto-Burman groups further to the east derived a much higher proportion of their ancestry from the lowlander lineage. The numbered circles/rectangles represent point estimates from qpAdm, and the thick and thin vertical segments represent ±1 and ±2 standard error measures (SEM) estimated by 5 cM block jackknifing, respectively. With respect to the non-Tibetan Tibeto-Burman populations, we infer genetic links among them along the circum-Plateau route (Fig. 5). We describe the results beginning with the southeast edge of the Plateau with the Naxi and Yi and proceed clockwise to the southwest (Fig. 5). First, Naxi and Yi from southwestern China have a genetic profile that closely resembles that of YR_MN but distinct from that of Upper_YR_LN (Supplementary Fig. 11). Using qpAdm, we model Naxi/Yi as a sister clade of YR_MN with no contribution from the Tibetan lineage required (Supplementary Table 5). Models using Upper_YR_LN as a proxy fail by returning ancestry coefficients larger than 1 from Upper_YR_MN. Naga from northeastern India are modeled as a mixture of 68–78% YR_MN/Naxi/Yi and 22–32% Tibetan lineage (Fig. 5; Supplementary Tables 5–6). Finally, Tamang and Gurung from the mid-altitude region of the southern Himalayas have even higher levels of their ancestry from the Tibetan lineage (60–63%), as well as a South Asian influx (9–19%) in addition to the YR_MN-like ancestry (Supplementary Table 7). Models using Naxi/Yi/Naga as a source instead of YR_MN also fit (Supplementary Table 7). This same three-way admixture model, Tibetan lineage + YR_MN/Naga + South Asian, also adequately fits the 16 Bhutanese Himalayan groups previously published24, with heterogeneous levels of contribution from the YR_MN/Naga (21–47% YR_MN or 32–75% Naga) and Tibetan lineages (20–82%; Supplementary Fig. 12; Supplementary Data 9). Overall, the South Asian contribution is small but non-negligible for many Bhutanese groups, ranging from 0 to 7%. Interestingly, for the populations in Nepal with substantial South Asian ancestry (e.g., Baram, Chantyal, Chepang, Gurung), south Indian tribal groups (e.g., Pulliyar) better represent their South Asian ancestry than northern Indian groups (Supplementary Data 9). These results highlight the complexity and multi-layered admixture history of Tibeto-Burman populations in the Himalayas. |
|
|
|
Post by Admin on Mar 12, 2022 19:08:16 GMT
Fig. 5: Genetic links between Tibeto-Burman speakers. 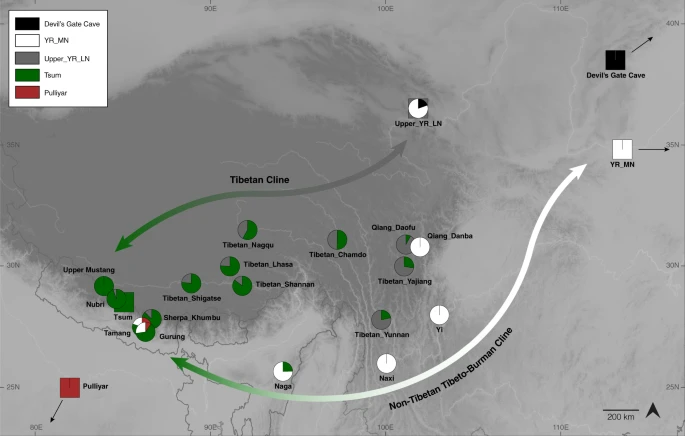 Tibetan groups from the Plateau and the Himalayas form a genetic cline, with the two poles represented by present-day Nepalese Tibetans (as well as aMMD) and Upper_YR_LN (“the Tibetan cline”). The non-Tibetan Tibeto-Burman cline reflects admixture along the circum-Plateau route and includes mid-altitude populations such as Naxi, Yi, Naga, Tamang and Gurung. Naxi and Yi cannot be modeled as a part of the Tibetan cline, i.e., Tsum+Upper_YR_LN; instead, YR_MN alone adequately models them. Non-Tibetan Tibeto-Burman speakers have higher contribution from the Tibetan lineage (represented by Nepalese Tibetan Tsum), and far-western mid-altitude populations Tamang and Gurung further have South Asian influx. Squares indicate the source populations used in ancestry models (circles). Prolonged positive selection on the EPAS1 and EGLN1 regions in Tibetans Our previous study reported that derived alleles for positively selected SNPs in the EPAS1 gene were observed only in the later Samdzong individuals but not the older Chokhopani and Mebrak individuals21. Including our new aMMD genomes, we still do not detect derived alleles in the EPAS1 haplotype block in the Chokhopani and Suila individuals, but we observe them at intermediate frequency in the other five sites (25–58%). Interestingly, the derived allele frequency in the ancient samples overall is lower than in present-day Tibetans (75%), indicating that selection still acted upon these alleles in the recent past (Supplementary Fig. 13; Supplementary Tables 8, 9). We also attempted to investigate the frequency changes at two adaptive nonsynonymous alleles in the EGLN1 gene: rs12097901, which is common among East Asians, and rs186996510, which is virtually unique to Tibetans16,40. Unfortunately, unfavorable capture conditions limit coverage for these two SNPs. Nevertheless, reads from shotgun sequencing suggest that the frequency of derived alleles in the genomic window spanning the EGLN1 gene in the aMMD samples is similar to that of present-day Tibetan populations (Supplementary Table 8); whether this finding indicates that selection on the EGLN1 alleles did not extend over the time period covered by the aMMD samples or is simply due to the sparsity of the sequence data is unclear. We next took advantage of 18 shotgun sequenced individuals in this study and our previous study18 to perform a genome-wide selection scan with window-based f3-statistics41 (Fig. 6; Supplementary Table 10). The method quantifies allele frequency differences between ancient and present-day Tibetans, using Han Chinese as an outgroup, and therefore aims to detect positive selection in present-day Tibetans since the time of the aMMD specimens. Combining 17 ancient individuals (excluding one individual due to relatedness), the genomic windows overlapping the EPAS1 gene show the strongest signals, supporting the continued positive selection at this locus. The genomic windows overlapping the EGLN1 gene show the second strongest signals. Interestingly, the elevated f3-statistics in these windows are not driven by the nonsynonymous SNPs rs12097901 and rs186996510 that had already reached high frequency in aMMD, but instead by SNPs that are common in both aMMD and Han but are rare in present-day Tibetans (Supplementary Data 10). Next, we looked at the overlap between the signals found in this selection scan (using z-score threshold of 4) and a previous set of signals (top 0.1% PBS values genome-wide) identified by using only contemporary population data23. All but three of the overlapping signals appear to be contributed to by the strong signature at the EPAS1 and EGLN1 loci (Supplementary Data 11). Of the three remaining regions, two span the PET112 and MCL1 genes, which are not well-established candidates for the response to hypoxia, and one contains the AKT3 gene, which is involved in angiogenesis and is implicated in the control of red blood cell traits in a candidate gene study42. |
|
|
|
Post by Admin on Mar 12, 2022 20:11:29 GMT
Fig. 6: Genome-wide selection scan using outgroup-f3 statistics with sliding windows. 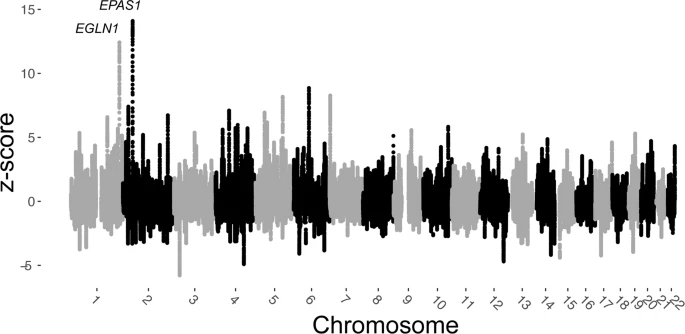 We computed f3 (Tibetans; aMMD, Han) using a sliding window approach with a window size 500 kb and a step size 10 kb. Z-scores for each window were calculated with a resampling approach (see Methods). Windows spanning the EPAS1 and EGLN1 genes harbor the two top signals. Discussion In this study, we analyze the genetic profile of 38 ancient Himalayan individuals and show that the ancestry found today among high altitude East Asians (i.e., Tibetans and Sherpa) was already distinctly diverged from lowlanders by 1494–1317 BCE. This pushes back the earliest evidence for the Tibetan gene pool at least by 500 years from our previous reports on Chokhopani21. Leveraging these early genomes, we illuminate key features of the genetic history of Tibetans and their relatives in the Tibetan Plateau and its periphery. We find that the Tibetan lineage is well-modeled as a mixture of two genetic ancestry sources: one is an ancient and previously uncharacterized Paleolithic substratum which accounts for up to 20% of contemporary Tibetan ancestry, and the other is related to lowlanders living at the northeastern fringe of the Plateau during the Late Neolithic. The Paleolithic substratum appears to have contributed exclusively to the Tibetan gene pool among the present-day populations studied to date. Our extensive modeling of present-day Tibetan and non-Tibetan Tibeto-Burman speakers identifies two genetic clines to explain their genetic history. These clines presumably reflect two distinct routes of population dispersal that are reflected in the distribution of diverse Tibeto-Burman languages in the Himalayas: one traversing the Plateau from its northeastern fringe to the Himalayas (the “northern” route), and the other along the periphery of the Plateau and the southern fringe of the Himalayas (the “southern” route) (Fig. 5). We provide a formal admixture modeling of the Tibetan populations along the northern cline and corroborate our previous report of this cline18,43. The genetic, cultural and linguistic diversity of present-day Tibeto-Burman speakers along the southern slope of the Himalayas reflects the confluence of ancient populations arriving via these two routes following their separation since the Late Neolithic. The unique features of the Tibetan genetic profile have long puzzled researchers, leading to wildly different and often incompatible population history models, ranging from Tibetans representing a sister clade that split from Han Chinese less than 3000 years ago14, to Tibetans branching off from a Han Chinese-related lineage more than 9000 years ago with gene flow from Paleolithic Siberians (Ust’-Ishim) or even from an unknown archaic hominin13,35. Moreover, these previous models, contradicting to each other, were developed on the basis only of present-day Tibetans and Han Chinese data and accepted an overly simplistic assumption that both populations are representative of the ancient groups ancestral to the two major branches of the Sino-Tibetan language family, i.e., Tibeto-Burman and Sinitic, respectively. Here we utilized ancient genomes from key time periods and geographic locations, which are better representatives of the lineages being modeled than present-day populations, to perform a direct test of the proposed demographic models. In our study, we show that ancestors of present-day Tibetans have been present in the Himalayas since at least ca. 1420 BCE, when the earliest direct evidence for sustained human presence appears at aMMD sites such as Suila and Lubrak. Moreover, we confirm the close relationship between early Himalayan populations and Late Neolithic groups living along the northeastern fringe of the Plateau around 2300–1800 BCE (Upper_YR_LN). Neolithic groups in the Gansu-Qinghai region likely include the ancestral population of those who later expanded onto the Plateau; however, the precise timing of the expansion is not clear. Barley cultivation, which is more suitable to the cooler and drier climate of the Plateau than millet, has long been argued to have allowed the Neolithic expansion onto the Plateau. While our results may ostensibly fit the long-held hypothesis of barley-driven expansion onto the Plateau ca. 1650 BCE, such a massive demic diffusion from Qinghai to the Himalayas in only about 200 years is unlikely to be a sole explanation for the ancient genetic link between the Plateau and the Gansu-Qinghai region. We propose an alternative scenario in which the genetic link between Plateau and lowland populations may have formed much earlier and therefore may not have been related to the introduction of barley or other domesticated plants or animals of Western Eurasian origin. The Karou site in eastern Tibet (ca. 5000–3000 BP) and the Qugong site near Lhasa (ca. 3800–3000 BP) show an indigenous archaeological tradition, and have assemblage composition and ceramic motifs distinct from those at Qijia2. In addition, the evidence from the Zongri site (ca. 2600–2000 BCE) suggests that Plateau hunter-gatherers traded for millets with lowlanders much earlier than the presumed introduction of barley44. The absence of an EGLN1 selection signature in the Upper_YR_LN combined with an estimated EGLN1 selective sweep dated to around 8000 BP40,45 suggests that the two populations may have already split long before the arrival of barley in Gansu-Qinghai. Barley was cultivated as a minor crop in Gansu-Qinghai as early as ca. 2000 BCE, leaving open the possibility for an earlier barley-driven expansion prior 1650 BCE, but archaeological evidence to support such a scenario is lacking. We acknowledge that our present data cannot completely reject the barley hypothesis; therefore, we call for a search of ancient genomes from the Plateau older than 1650 BCE to directly test it. Finally, our study shows the prolonged effects of natural selection in shaping the gene pool of high altitude East Asians. Of note, the increase in the EPAS1 allele frequency over the time period spanning the aMMD samples and present-day Tibetans highlights the slow but steady action of positive selection on this Denisovan-derived genetic variant. Future studies on additional ancient genomes across the Tibetan Plateau will be able to lead us toward a comprehensive understanding of the evolutionary history of the two Tibetan signature genes, EGLN1 and EPAS1, as well as to investigate further the polygenic signatures of adaptation suggested by the study of present-day genomes23. Supplementary Information static-content.springer.com/esm/art%3A10.1038%2Fs41467-022-28827-2/MediaObjects/41467_2022_28827_MOESM1_ESM.pdfPeer Review File static-content.springer.com/esm/art%3A10.1038%2Fs41467-022-28827-2/MediaObjects/41467_2022_28827_MOESM2_ESM.pdf |
|
|
|
Post by Admin on Mar 15, 2022 21:02:05 GMT
A dual origin of Tibetans: evidence from mitochondrial genomes Yu-Chun Li, Jiao-Yang Tian & Qing-Peng Kong Journal of Human Genetics volume 60, 403–404 (2015) Substantial progress on how modern humans settled and adapted to the Tibetan Plateau has been achieved in the past few years. In particular, genetic evidence suggests that peopling the Plateau was attributed mainly to the neolithic immigration initiated from northern China ~7 kilo years ago (kya).1, 2 Furthermore, a very small proportion of genetic components showed a restricted distribution in the Tibetans and were estimated to be extraordinarily ancient, thus plausibly reflecting an early dispersal event onto the Plateau during the Late Pleistocene.1 This observation is in good agreement with the previous archeological records.3 Recently, a more extensive study, by analyzing archeological materials collected from 53 Neolithic and Bronze sites in northeast Tibetan Plateau, provides solid evidence in support of the Neolithic immigration into Tibet and, furthermore, suggests a permanent settlement on the high areas of the Plateau likely occurred till 3.6 kya.4 Although some inconsistency on dating the entrance exists between archeological and genetic studies, likely introduced by the methodology in time estimation, evidence from both disciplines supports the notion that the Tibetans can trace their origin to the Neolithic dispersal from northern China.1, 2, 3, 4 Since evidence from both mitochondrial DNA (mtDNA)1, 2 and Y chromosome2 also suggests the existence of genetic relic of the Late Pleistocene settlers in the Tibetans, a new question then arises: how did modern humans move onto the Plateau or, alternatively, how were these ancient genetic components introduced into the Tibetans? To provide more insights into this issue, we collected and analyzed mtDNA data of a total of 53 665 Asian individuals from 814 populations (Figure 1a and Supplementary Table S1). Then, we examined the distribution of haplogroup M62b, a representative of the suggested genetic relic of the Late Pleistocene immigrants, via a motif search strategy.5 Our result reveals that this haplogroup shows a restricted distribution in the Tibetans and has age ~24 kya (Figures 1b and c), in concordance with the previous studies.1, 2 In contrast with the Neolithic-originated components that can find their closer sister lineages in the current northern Chinese populations,1, 2 only few M62b mtDNAs are present sporadically in some surrounding populations, which all locate merely at the terminal branch (Figure 1c) and thus are most likely introduced from the Tibetan populations via recent gene flow. Taking into account the observation of M62a (~15–24 kya; Figures 1b and c), the sister clade of M62b, in India,6 it is then plausible that M62b might have been introduced into the Tibetans via a different way from the northeast Tibetan Plateau entrance. Our extensively searching result confirms this and further reveals the presence of M62a in Bangladesh and Tibet (Figure 1c). A phylogenetic tree of haplogroup M62 is reconstructed on the basis of whole mitochondrial genomes. Evidently, the Indian M62a mtDNAs belong to the same clade, M62a1 (defined by variants 11914, 12793 and a reverse variant at 4561), whereas another clade in M62a, M62a2 (defined tentatively by variants 16491, 16487, 16452 and 16189), contains only a single Tibetan individual. This result, together with the specific distribution pattern of M62b, suggests haplogroup M62 most plausibly to be originated or differentiated in the Tibetans at ~30–45 kya (Figures 1b and c). Intriguingly, haplogroup M68 (with an age of ~60–80 kya7), the sister clade of M62, was observed in mainland Southeast Asia,7 strongly arguing for a south origin of haplogroup M62. Figure 1 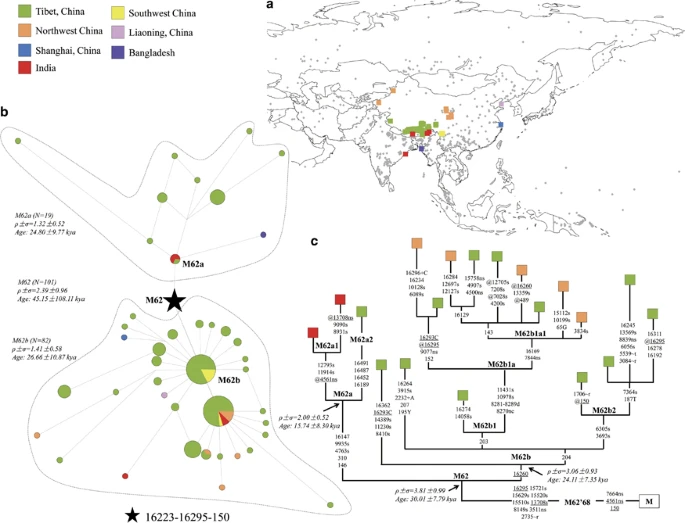 Phylogeographic structure of mtDNA haplogroup M62. (a) Sampling locations. Diamonds in gray indicate the 814 Asian populations (comprising of 53 665 individuals; see Supplementary Table S1), with the one bearing haplogroup M62 being highlighted by square. (b) Median joining network of haplogroup M62. Coalescent ages are calibrated based on mutation rate of control region transitions on segment 16090–1636510 by rho statistic method. (c) Phylogenetic tree of haplogroup M62. The tree is reconstructed by using 19 reported whole mtDNA genomes (see Supplementary Table S2). Nucleotide position numbers are consistent with the revised Cambridge reference sequence (rCRS11). Suffixes C, G and T refer to transversions, ‘d’ means a deletion, ‘s’ means synounymous mutation, ‘ns’ means nonsynounymous mutation and ‘+’ indicates an insertion; recurrent mutations are underlined; ‘@’ means a reverse mutation; and ‘h’ means heterogeneity. Coalescent ages are calibrated based on mutation rate of coding region synonymous mutations10 by rho statistic method. A full color version of this figure is available at the Journal of Human Genetics journal online. |
|






