Post by Admin on May 4, 2016 22:47:03 GMT
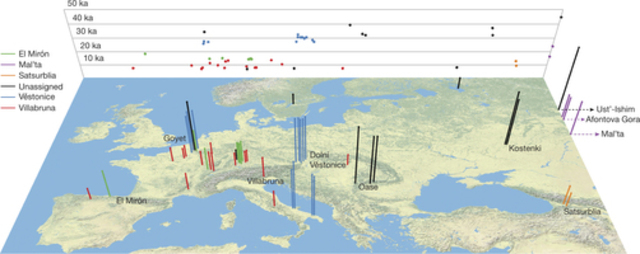
Figure 1: Location and age of the 51 ancient modern humans.
Modern humans arrived in Europe ~45,000 years ago, but little is known about their genetic composition before the start of farming ~8,500 years ago. Here we analyse genome-wide data from 51 Eurasians from ~45,000–7,000 years ago. Over this time, the proportion of Neanderthal DNA decreased from 3–6% to around 2%, consistent with natural selection against Neanderthal variants in modern humans. Whereas there is no evidence of the earliest modern humans in Europe contributing to the genetic composition of present-day Europeans, all individuals between ~37,000 and ~14,000 years ago descended from a single founder population which forms part of the ancestry of present-day Europeans. An ~35,000-year-old individual from northwest Europe represents an early branch of this founder population which was then displaced across a broad region, before reappearing in southwest Europe at the height of the last Ice Age ~19,000 years ago. During the major warming period after ~14,000 years ago, a genetic component related to present-day Near Easterners became widespread in Europe. These results document how population turnover and migration have been recurring themes of European prehistory.
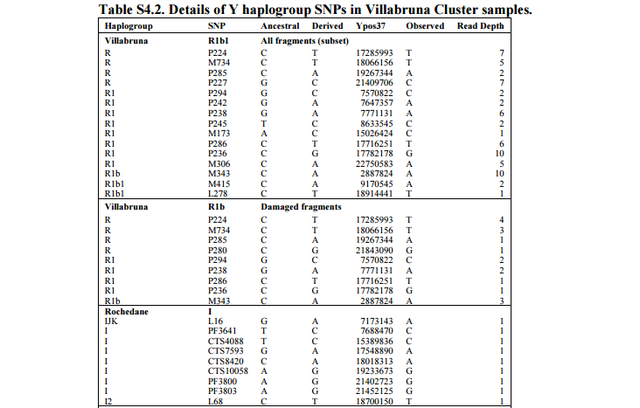
Previous studies assumed that Kostenki14 was the oldest European man with Basal Eurasian ancestry, who survived the Ice Age, but Fu et al. (2016) put it more accurately. Kostenki14 was likely to be a mixed-race individual between populations related to East Asians (hg C) and the ancestors of Europeans (hg U2). This new study also found that some of the Villabruna Cluster individuals such as Loschbour and LaBrana1 are closely associated with Han Chinese but not all Villabruna individuals have an affinity with East Asians. The ancestors of Loschbour and LaBrana1 in the Villabruna Cluster may have interbred with ancient populations related to East Asians as well, which was why excess allele sharing with East Asians was detected. Moreover, the ‘Villabruna Cluster’ is composed of 15 post-Last Glacial Maximum individuals from 14,000–7,000 years ago. Of these fifteen Villabruna samples, older twelve samples belonged Haplogroup R or R1 and haplogroups R1b and R1b1 only appear in more recent three samples, which are dated around 7,000 years ago (Table S4.2. Details of Y haplogroup SNPs in Villabruna Cluster samples).
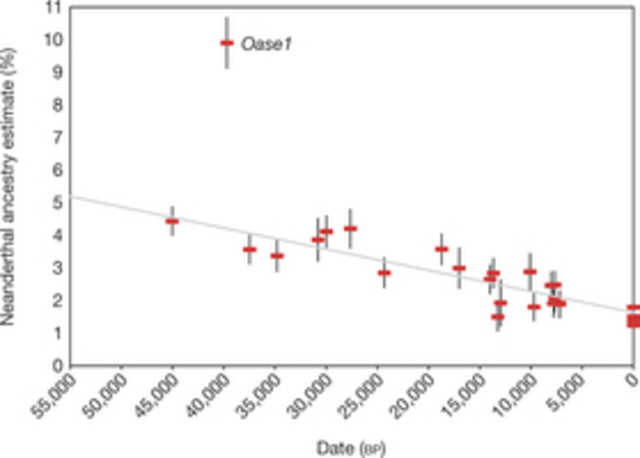
Figure 2: Decrease of Neanderthal ancestry over time.
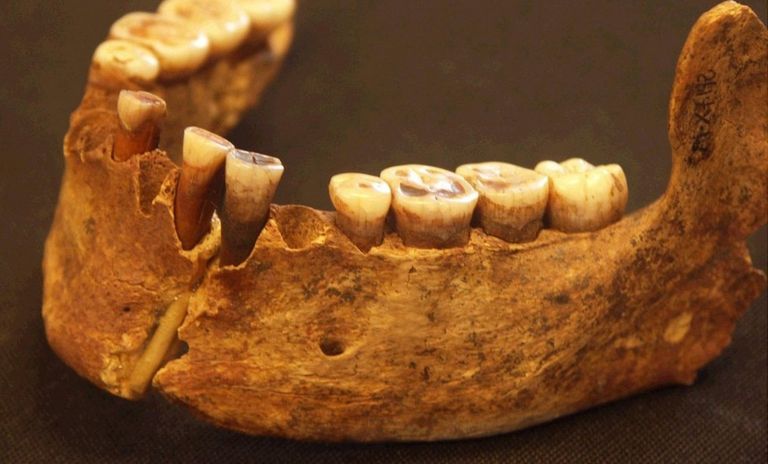
The burial of Riparo Villabruna was discovered in 1988 by A. Broglio in the small rockshelter named Riparo Villabruna A in the Veneto Dolomites. It contains a partial skeleton with lower limbs severed at the distal femoral shafts associated with burial goods of the Epigravettian culture50. The date quoted here comes from the skull51, whereas the genetic analysis is of a left femur. This individual bears the earlier known example of treatment of dental caries52. We were surprised to assign Villabruna to R1b1 (Table S4.2). When we restrict to damaged sequences, we still assign it to R1b.
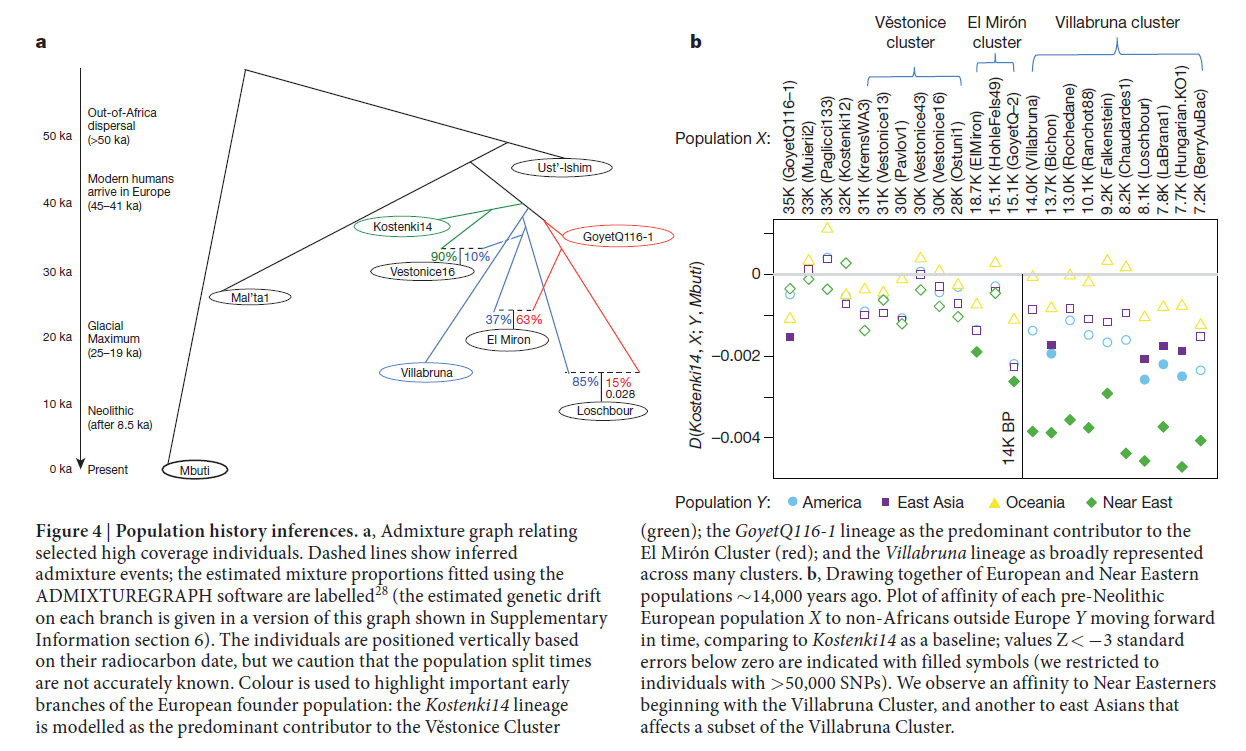
Gene flow linking the Villabruna Cluster and the Near East
We investigated the relationship between pre-Neolithic Europeans and present-day as well as ancient populations using statistics of the form: D(European1, European2; Test, Mbuti). Here, the Test populations are Native Americans, East Asians, Oceanians and Near Eastern populations from the Simons Genome Diversity Project (SGDP) panel.
Affinities of pre-Neolithic Europeans to the Near East
When neither of the two pre-Neolithic Europeans analysed in the statistic is in the Villabruna Cluster—that is, both are older than about 14,000 BP—they tend to be symmetrically related
to populations outside Europe including present-day and ancient Near Easterners. However, when one lived prior to the Villabruna Cluster (e.g. Vestonice16, ElMiron, Kostenki14,
KremsWA3, and GoyetQ116-1) and the other is in the Villabruna Cluster (e.g. BerryAuBac, Bichon, Falkenstein, Hungarian.KO1, LaBrana1, Loschbour, Ranchot88, Rochedane and Villabruna), there is a distinct attraction of the Villabruna Cluster samples to Near Eastern populations (Figure 4b; Extended Data Figure 3). Table S11.1 shows the statistics when the Near Eastern population is Iraqi_Jew.

There are several possible explanations for these findings. One is gene flow between relatives of Near Easterners and pre-Neolithic Europeans after ~14,000 years ago, beginning
with the Villabruna Cluster. A second is population substructure in Europe. In this scenario, after post-glacial re-peopling of Europe, the balance of ancestry could have shifted toward populations that were more closely related to Near Easterners. In either case, however, major population turnovers must have occurred.
The affinity of pre-Neolithic Europeans to Near Easterners beginning around 14,000 years ago is distinct from the affinity to East Asians in Mesolithic Europeans Seguin-Orlando et al.1 documented that statistics of the form D(Kostenki14; Mesolithic Europeans; East Asians, Outgroup) are significantly less than 0. In Supplementary Information section 8, we show that this is likely due to gene flow between the ancestors of East Asians and the ancestors of Mesolithic Europeans. These patterns are evident in Figure 4b and Extended Data Figure 3, which show that a subset of Villabruna Cluster samples including Mesolithic Europeans show a significant affinity to East Asians. This pattern does not go hand-in-hand with the affinity to the Near East that is present in all Villabruna Cluster
samples, and thus the two signals must therefore reflect at least two distinct historical events.
The Satsurblia and Villabruna Clusters are not particularly closely related
What is the nature of the West Eurasian genetic affinities in the Satsurblia Cluster samples? We observe significantly positive statistics of the form D(Villabruna Cluster, Early Upper
Palaeolithic Europeans; Satsurblia Cluster, Mbuti), showing that Satsurblia Cluster samples share more alleles with Villabruna Cluster samples—for example, Villabruna, Bichon, Loschbour, LaBrana1, and Hungarian.KO1—than with Early Upper Palaeolithic Europeans (Kostenki14, GoyetQ116-1, Vestonice16 and ElMiron) (Table S12.2). This suggests the possibility that a component of the non-Basal Eurasian ancestry in the Satsurblia Cluster may be related to the ancestry that appears in our European sample series with the Villabruna Cluster. In other words, migrations involving populations related to the Satsurblia Cluster could be responsible for the genetic link between the Near East and the Villabruna Cluster (Supplementary Information section 11).
To explore the relationship between the non-Basal Eurasian ancestry in the Satsurblia cluster and the Near Eastern related ancestry in the Villabruna Cluster in more detail, we fit Satsurblia into the Admixture Graph of Supplementary Information section 6 that includes Mbuti, UstIshim, Malta1, GoyetQ116-1, Kostenki14, Vestonice16, and ElMiron (Figure S6.3). In all fitting models, Satsurblia is inferred to harbor ~32% ancestry from a Basal Eurasian lineage that branched before UstIshim (Extended Data Figure 4). These results are consistent with the finding that Satsurblia Cluster samples have Basal Eurasian ancestry1, while also rejecting the previous model in which Satsurblia is of entirely Basal Eurasian ancestry1. Our fitted model specifies that the majority of ancestry in Satsurblia is from a West Eurasian, lineage providing an explanation both for why Satsurblia has Basal Eurasian ancestry, while at the same time sharing alleles with all Europeans beginning with Kostenki14.
All three of the best fitting models in Extended Data Figure 4 specify that the majority ancestry component in Satsurblia branched very deeply in the tree of West Eurasian populations, forming a clade with Malta1. Further evidence for a deep connection to Malta1 and populations with admixture of Malta1-related ancestry comes from the observation in Table S12.2 that D(Motala12/Malta1, Early Upper Palaeolithic Europeans; Satsurblia Cluster, Mbuti) is significantly positive. In a simple model of this type, a prediction is that statistics of the form D(Villabruna Cluster, Early Upper Palaeolithic Europeans; Malta Cluster, Mbuti), would be significantly positive, as Malta1 would share more alleles with Villabruna Cluster samples than with Early Upper Palaeolithic Europeans. However, we do not detect any such a signal (Supplementary Information section 9).
Regardless of whether a population closely related to Satsurblia is responsible for the affinity of Villabruna Cluster samples to the Near East, there is evidence that a new lineage with affinities to present-day Near Easterners spread across Europe at this time. The evidence for this spread is that the genetic affinity of pre-Neolithic Europeans to Near Easterners abruptly increases with the appearance Villabruna Cluster, with no earlier European sample showing as strong an affinity despite sharing large amounts of genetic drift with the Villabruna Cluster (Figure 4b). An important direction for future work is to analyse more ancient DNA samples from southeastern Europe and western Asia in order to understand the history of the migratory events that our data show must have occurred around this time.
The Villabruna Cluster is represented by the largest number of individuals in this study. This allows us to study heterogeneity within this cluster (Supplementary Information section 13). First, we detect differences in the degree of allele sharing with members of the El Mirón Cluster, as revealed by significant statistics of the form D(Test1, Test2; El Mirón Cluster, Mbuti). Second, we detect an excess of allele sharing with east Asians in a subset of Villabruna Cluster individuals— beginning with an ~13,000-year-old individual from Switzerland—as revealed by significant statistics of the form D(Test1, Test2; Han, Mbuti) (Fig. 4b and Extended Data Fig. 3). For example, Han Chinese share more alleles with two Villabruna Cluster individuals (Loschbour and LaBrana1) than they do with Kostenki14, as reflected in significantly negative statistics of the form D(Kostenki14, Loschbour/LaBrana1; Han, Mbuti)4. This statistic was originally interpreted as evidence of Basal Eurasian ancestry in Kostenki14. However, because this statistic is consistent with zero when Han is replaced with Ust’-Ishim, these findings
cannot be driven by Basal Eurasian ancestry (as we discuss earlier), and must instead be driven by gene flow between populations related to east Asians and the ancestors of some Europeans (Supplementary Information section 8).
Nature (2016) doi:10.1038/nature17993

 that lived south of the Alpine ridge ~5,200 years before present (ybp), during the Copper Age. Despite studies that have investigated his genetic profile, the relation of the Iceman´s maternal lineage with present-day mitochondrial variation remains elusive. Studies of the Iceman have shown that his mitochondrial DNA (mtDNA) belongs to a novel lineage of haplogroup K1 (K1f) not found in extant populations. We analyzed the complete mtDNA sequences of 42 haplogroup K bearing individuals from populations of the Eastern Italian Alps – putatively in genetic continuity with the Tyrolean Iceman—and compared his mitogenome with a large dataset of worldwide K1 sequences. Our results allow a re-definition of the K1 phylogeny, and indicate that the K1f haplogroup is absent or rare in present-day populations. We suggest that mtDNA Iceman´s lineage could have disappeared during demographic events starting in Europe from ~5,000 ybp. Based on the comparison of our results with published data, we propose a scenario that could explain the apparent contrast between the phylogeographic features of maternal and paternal lineages of the Tyrolean Iceman within the context of the demographic dynamics happening in Europe from 8,000 ybp.
that lived south of the Alpine ridge ~5,200 years before present (ybp), during the Copper Age. Despite studies that have investigated his genetic profile, the relation of the Iceman´s maternal lineage with present-day mitochondrial variation remains elusive. Studies of the Iceman have shown that his mitochondrial DNA (mtDNA) belongs to a novel lineage of haplogroup K1 (K1f) not found in extant populations. We analyzed the complete mtDNA sequences of 42 haplogroup K bearing individuals from populations of the Eastern Italian Alps – putatively in genetic continuity with the Tyrolean Iceman—and compared his mitogenome with a large dataset of worldwide K1 sequences. Our results allow a re-definition of the K1 phylogeny, and indicate that the K1f haplogroup is absent or rare in present-day populations. We suggest that mtDNA Iceman´s lineage could have disappeared during demographic events starting in Europe from ~5,000 ybp. Based on the comparison of our results with published data, we propose a scenario that could explain the apparent contrast between the phylogeographic features of maternal and paternal lineages of the Tyrolean Iceman within the context of the demographic dynamics happening in Europe from 8,000 ybp.





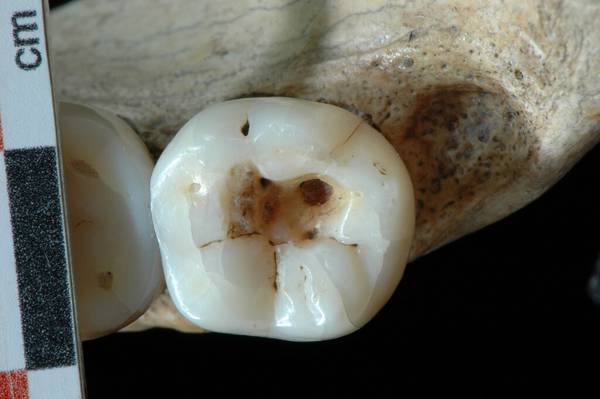



 Figure 3: Scanning Electron Microscopy (SEM) images of the striations observed within the carious cavity of the Villabruna RM3.
Figure 3: Scanning Electron Microscopy (SEM) images of the striations observed within the carious cavity of the Villabruna RM3.