|
|
Post by Admin on Jul 26, 2018 18:33:39 GMT
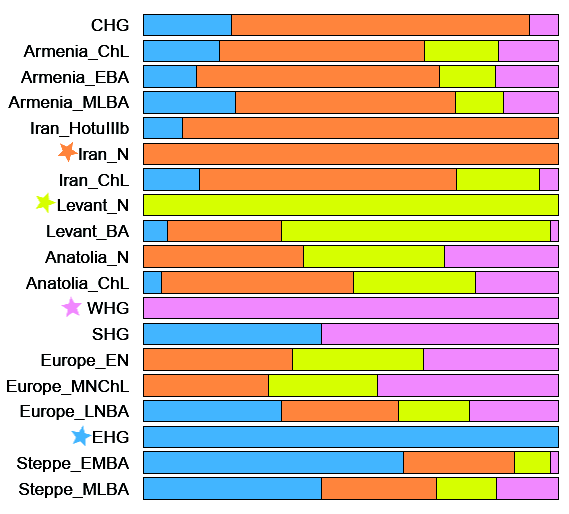 Population Genomic Analysis We investigated the genome-wide relationships of our samples to modern and other ancient humans by performing principal-component analysis (PCA) defined by a subset of Eurasian populations from the Human Origins dataset [8, 9] (Figure 1B). Our Romanian genomes were projected onto the first two PCA axes, together with a broad panel of ancient individuals (see Data S1 for sample details), augmented by two newly sequenced Spanish Mesolithic hunter-gatherer genomes (Chan_Meso, an Early Mesolithic dated at 9,131 ± 124 cal BP (calibrated age before present), and Canes1_Meso, a Late Mesolithic dated at 7,115 ± 130 cal BP) to provide better coverage for this important period (Table 1 and STAR Methods, Method Details). The three Romanian Mesolithic genomes clustered together with other Mesolithic samples, including the two new Spanish ones, and close to modern Northern European populations (Figure 1B). The Romanian Eneolithic genome GB1_Eneo, on the other hand, was placed in a different region of the plot, located in a unique position between European Mesolithic hunter-gatherers and Early Neolithic farmers on PC2. We confirmed the intermediate nature of this genome by estimating ancestral components using the clustering algorithm ADMIXTURE. Whereas the Romanian Mesolithic hunter-gatherers had a single major ancestral component shared with other WHGs, the Romanian Eneolithic sample was characterized by a mixture between this WHG component and a component maximized in Neolithic farmers (Figures 1C and S3B). Other admixed European Neolithic and Eneolithic/Chalcolithic farmers had at most 20%–30% WHG ancestry in ADMIXTURE analysis, and the Romanian Eneolithic is the only individual who is genetically predominantly Mesolithic (61.7%, 95% confidence interval [CI] 59.9%–63.4%) with a minority contribution from the Neolithic. We note that Gok2, a Swedish Neolithic sample, was originally estimated to have 77% hunter-gatherer ancestry [5], but in our analysis it has a much lower percentage (27.2%, 95% CI 25.1%–29.4%), in line with other recent analyses [23]. Although GB1_Eneo is chronologically close to the beginning of the Bronze Age, we did not find the green component (Figure 1C) characteristic of individuals from the Yamnaya culture, showing that the large hunter-gatherer component in this Eneolithic individual is unlikely to be due to admixture with incoming steppe pastoralists (Table 2). Table 2 Key D Statistics of the Form D(A,B; X,Y) A B X Y D Z Score Loci The Romanian Samples Form a Clade with Each Other (|Z| < 3; the Most Positive and Negative Statistics Are Shown for Each Comparison) Mbuti Natufian SC1_Mes oSC2_Meso 0.03 1.635 26,749 Mbuti ElMiron SC1_Meso SC2_Meso −0.0281 −1.921 44,549 Mbuti GoyetQ116-1 SC1_Meso OC1_Meso 0.0272 1.896 41,197 Mbuti SC2_Meso SC1_Meso OC1_Meso −0.0283 −2.036 57,452 Mbuti Kostenki14 OC1_Meso SC2_Meso 0 .0231.964 79,553 Mbuti Armenia_EBA OC1_Meso SC2_Meso 0.0188 −2.192 77,923 There Is Anatolian-Farmer-Related Admixture in the Romanian Eneolithic Sample as Compared to the Romanian Mesolithic Samples We formalized these inferences by computing outgroup f3 in the form f3(ancient1, ancient2, Mbuti), thus estimating the amount of shared drift between pairs of ancient samples with respect to an African outgroup. Our three Romanian Mesolithic samples share the most drift with each other (Figure 2), followed by other WHGs, including our new Spanish samples. The genetic affinity among the Romanian Mesolithic samples was such that they form a clade to the exclusion of other ancient samples, as shown by D statistics of the form D(Romanian Mesolithic 1, Romanian Mesolithic 2, other_ancient, Mbuti) (Table 2). Interestingly, this was not the case for the Spanish Mesolithic samples, as Chan is somewhat divergent from Canes1 and La Braña (Figure S3A and Table 2), highlighting the genetic diversity of European Mesolithic hunter-gatherer lineages. The Romanian Eneolithic individual, on the other hand, once again showed a mix of affinities. Based on outgroup f3, the genomes that shared the most drift with this Eneolithic sample are WHGs, in line with the large amount of that ancestry detected in ADMIXTURE. However, its affinity to Neolithic samples is also relatively high compared to the Romanian Mesolithic samples (Data S2). This conclusion is supported by D statistics of the form D(GB1_Eneo, Romanian HG, Anatolian Neolithic, Mbuti), which indicate some Near Eastern ancestry (Table 2). Our three Romanian hunter-gatherer samples are not direct representatives of the hunter-gatherer component in GB1_Eneo (Table 2); however, this might be due simply to the geographic distance between the sites, especially given the observed heterogeneity among Spanish Mesolithic hunter-gatherers. |
|
|
|
Post by Admin on Jul 27, 2018 18:29:55 GMT
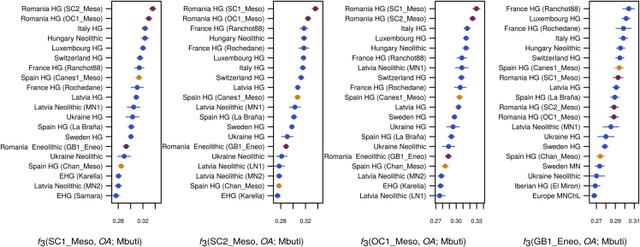 Figure 2 Outgroup f3 Statistics Analysis of runs of homozygosity (ROH) showed that the Eneolithic sample had a profile intermediate between Early Neolithic farmers and hunter-gatherers, consistent with the sample’s mixed origins (Figure S4A). Finally, we also tested the proportion of Neanderthal ancestry in each sample, which was consistent with the age of the sample (Figure S4B) [24]. Discussion The genetic analysis of the Eneolithic individual from Gura Baciului provides support for a scenario of complex interactions between hunter-gatherers and farmers in the Lower Danube Basin. At this stage, we cannot discern at what point during or after the Neolithic transition the observed hunter-farmer admixture occurred. Stable isotopic data from Lepenski Vir suggested early contacts between Neolithic immigrants and Mesolithic communities, whereby the first incomers changed the subsistence practices of the local fisher-hunter-gatherers, but not necessarily other cultural traits such as funerary practices [11, 28]. On the other hand, a resurgence of Mesolithic ancestry in the Late Neolithic has already been noted in other parts of Europe [6, 24], even though the relative contribution of this ancestry was limited compared to the levels we report for Romania. The fact that our Romanian hunter-gatherers are not direct representatives of the hunter-gatherer ancestry in GB1_Eneo (Table 2) provides some support for this latter scenario, but the geographic distance between the Mesolithic and Neolithic/Eneolithic sites might also explain this lack of a direct link. Although our results provide evidence for admixture between these communities, we do not know whether it resulted from the incoming farmers establishing their own communities into which the hunter-gatherers mixed or from small numbers of farmers joining existing hunter-gatherer communities. Our analysis of uniparental markers provides a caveat on their use to infer the dynamics of interactions among different populations. Although two of our Romanian Mesolithic samples belong to mtDNA haplogroups (U5a and U5b), which are typical of that group, another Mesolithic individual of similar age and geographical origin (OC1_Meso) shares the same haplogroup (K1) with the much later Eneolithic individual GB1_Eneo and four previously described Middle/Late Neolithic and Eneolithic samples from Romania [29]. K1 has mostly been found in early farmers from Europe [6, 17, 18, 30, 31, 32] and has been hypothesized to have its origin in the Near East [1, 9, 18]. Although the latter hypothesis is not invalidated by our data, the occurrence of this haplogroup in a Mesolithic sample from Romania suggests that it entered the European gene pool clearly ahead of the Neolithic transition and should therefore not be used as a marker for tracking farmers of Anatolian origin. Similarly, all three Romanian Mesolithic individuals belonged to the Y chromosome haplogroup R1. This haplogroup is thought to have originated shortly after the Late Glacial Maximum (LGM) [33], but its frequency is found to increase greatly in Central Europe during the Late Neolithic and Bronze Age, leading to the hypothesis that these haplotypes expanded from the East only after ∼4.5 kya [6]. Its presence in all of our Romanian Mesolithic individuals and in older European human remains [7, 34] suggests that this haplotype was already found at high frequencies in certain regions of Europe well before then, even though this observation does not negate that the changes in frequency during the Bronze Age might reflect migration by steppe pastoralists. The genome-wide data that we obtained from the Romanian Mesolithic and Eneolithic individuals suggest that the interaction between farmers and hunter-gatherers was much more complex than the simple picture of mostly demic replacement of hunter-gatherers by farmers that has been suggested by previous studies of ancient genomes from Western and Central Europe. The limited number of samples in this study prevents a formal investigation of the demographic processes that led to the observed patterns in D and f statistics. However, when our findings are considered together with the observation of an Early Neolithic individual from Hungary (KO1), whose ancestral component was completely derived from WHG [32], as well as the most recent finding that the Neolithic transition in the Baltic and Ukraine occurred with little or no input from farmers of Anatolian ancestry [7], a picture emerges in which the further north and east we go from the Carpathian Basin, the lesser the role of a demic diffusion in the spread of the Neolithic traits, thus implying an increase in importance of cultural exchange. Interestingly, a similar gradient has been suggested for East Asia, with high levels of genetic continuity with hunter-gatherer populations in the northern parts of the region and admixture between this lineage and incoming farmers further south [35]. We do not know what determined this cline. We could speculate, though, that the Neolithic package, which was developed in the Near East, might have been somewhat unsuitable for the new climates and environments encountered in the northeast of Europe, leading to a progressive mixing with local hunter-gatherers and acquisition of local knowledge, with the eventual breakdown of demic diffusion and the spread of isolated Neolithic traits by cultural diffusion. Another scenario is that hunter-gatherers were in higher densities in these eastern regions and hence that interactions between hunter-gatherers and farmers were more frequent than in regions further west. Curr Biol. 2017 Jun 19; 27(12): 1801–1810.e10. |
|
|
|
Post by Admin on May 5, 2020 21:19:38 GMT
Tracing the genetic origin of Europe's first farmers reveals insights into their social organization Anna Szécsényi-Nagy, Guido Brandt, Wolfgang Haak, Victoria Keerl, János Jakucs, Sabine Möller-Rieker, Kitti Köhler, Balázs Gusztáv Mende, Krisztián Oross, Tibor Marton, Anett Osztás, Viktória Kiss, Marc Fecher, György Pálfi, Erika Molnár, … See all authors Published:22 April 2015https://doi.org/10.1098/rspb.2015.0339 Abstract Farming was established in Central Europe by the Linearbandkeramik culture (LBK), a well-investigated archaeological horizon, which emerged in the Carpathian Basin, in today's Hungary. However, the genetic background of the LBK genesis is yet unclear. Here we present 9 Y chromosomal and 84 mitochondrial DNA profiles from Mesolithic, Neolithic Starčevo and LBK sites (seventh/sixth millennia BC) from the Carpathian Basin and southeastern Europe. We detect genetic continuity of both maternal and paternal elements during the initial spread of agriculture, and confirm the substantial genetic impact of early southeastern European and Carpathian Basin farming cultures on Central European populations of the sixth–fourth millennia BC. Comprehensive Y chromosomal and mitochondrial DNA population genetic analyses demonstrate a clear affinity of the early farmers to the modern Near East and Caucasus, tracing the expansion from that region through southeastern Europe and the Carpathian Basin into Central Europe. However, our results also reveal contrasting patterns for male and female genetic diversity in the European Neolithic, suggesting a system of patrilineal descent and patrilocal residential rules among the early farmers. 1. Introduction Agriculture was first established in the Near Eastern Fertile Crescent after 10 000 BC; and expanded from the Levant and Anatolia to southeastern Europe [1]. Archaeological research has described the subsequent spread of Neolithic farming into (and throughout) Central and southwestern Europe along two major and largely contemporaneous routes. On the Continental route, the Carpathian Basin connected southeastern Europe to the Central European loess plains, while the Mediterranean route bridged the eastern and western Mediterranean coasts, introducing farming to the Iberian Peninsula in the west [2–6]. The Early Neolithic Starčevo culture (STA) has played a major role in the Neolithization of southeastern Europe. The STA expanded from present-day Serbia to the western part of the Carpathian Basin, encompassing the regions of today's northern Croatia and southwestern Hungary (ca 6000–5400 BC) [7,8] (figure 1). The earliest Linearbandkeramik culture (LBK) emerged in the mid-sixth millennium BC in western Hungary or Transdanubia (hence called ‘LBK in Transdanubia’, or LBKT) [9], marking the beginning of sedentary life in Central Europe. The earliest LBKT coexisted with the STA in Transdanubia for approximately 100–150 years [10]. Archaeological research described an interaction zone between indigenous hunter–gatherer groups and farmers at the northernmost extent of the STA in Transdanubia, which might have given rise to the LBKT [10,11]. After its formative phase in western Hungary, the LBK spread rapidly to Central Europe, reaching central Germany around 5500 BC [2,12]. In the following 500 years, the LBK continually expanded, eventually covering a vast geographical area from the Paris Basin to Ukraine in its latest phase [2,13], while persisting in Transdanubia until approximately 4900 BC (figure 1).  Despite the well-established archaeological relations between the STA and LBKT in the Carpathian Basin and the LBK in Central Europe, their genetic relationship is yet unclear. Traditionally, scholars have explained the Neolithic transition either as an expansion of early farmers from the Near East, who brought new ideas as well as new genes (demic diffusion) [14–17], or as an adoption of farming technologies by indigenous hunter–gatherer populations with little or no genetic influence (cultural diffusion) [18–21]. These two contrasting models have been merged into complex integrationist approaches, considering small-scale population movements on regional levels [1,2,10,22]. Inferences drawn from present-day genetic data have yielded contradictory results about the Neolithic impact on the genetic diversity of modern Europeans, showing a disparity between mitochondrial DNA (mtDNA) and Y chromosomal patterns. Several Y chromosome studies supported the Neolithic demic diffusion model [17,23,24], while most mtDNA and some Y chromosomal studies have proposed a continuity of Upper Palaeolithic lineages [20,21,25,26]. The contrasting mtDNA and Y chromosomal evidence has been explained by differences in evolutionary scenarios, such as sex-biased migration [27]. Recent ancient DNA (aDNA) studies have provided direct insights into the mtDNA and nuclear genomic diversity of hunter–gatherers in Europe [28–35] and the Central European LBK [33,36–40], describing genetic discontinuity between local foragers and early farmers [28,31,38,40]. Comparative analyses with present-day populations have revealed Near Eastern affinities of the mitochondrial LBK ancestry, supporting the demic diffusion model and population replacement at the beginning of the Neolithic period [38,39]. Ancient genomic studies have described the early farmers as genetically most similar to extant populations of southern Europe [31,33,40,41]. Y chromosome data from prehistoric Europeans is still scarce but relevant for our study (electronic supplementary material, dataset S20), including hunter–gatherers [33–35,40,42], and early farmers from Germany [38,40], eastern Hungary [41], Austria [43] and southwest Europe [40,44,45]. The postulated Near Eastern origin of LBK farmers has so far only been inferred from modern-day population data, and while the first ancient mtDNA data from early Near Eastern farmers has been reported recently [46], the genetic diversity in the vast territory from the Fertile Crescent to Central Europe remains largely unexplored. Consequently, our aims were to (i) study the genetic diversity of the early farming Carpathian Basin cultures from both the mtDNA and Y chromosome perspectives, (ii) examine whether men and women had different demographic histories, (iii) investigate the contribution of the STA to the genetic variability of the LBKT and LBK, (iv) reveal the potential genetic origins of the first farmers in Eurasia and (v) assess the role of the Continental route in the European Neolithic dispersal. In this study, we present 84 mtDNA and 9 Y chromosomal DNA data from Mesolithic (6200–6000 BC) and Neolithic specimens of the STA and LBKT from western Hungary and Croatia. Spanning a time transect of the Hungarian Neolithic in Transdanubia over approximately 900 years (ca 5800–4900 BC) allowed us to gain detailed insight into the spread of farming from the Near East. |
|
|
|
Post by Admin on May 6, 2020 19:30:48 GMT
2. Results (a) Mitochondrial DNA. Using well-established aDNA methods (electronic supplementary material, material and methods), we genotyped mtDNA variability by sequencing the hyper-variable segment I and II (HVS-I/II) and 22 single nucleotide polymorphisms (SNPs) in the coding region of the mitochondrial genome [38]. Overall, we investigated 109 skeletons from one Mesolithic, six STA and eight LBKT sites from western Hungary and Croatia (electronic supplementary material, figure S8, datasets S1 and S2). We genotyped endogenous HVS-I sequences of 84 individuals (1 hunter–gatherer, 44 STA and 39 LBKT), yielding a success rate of 76% (electronic supplementary material, dataset S3). We also sequenced parts of HVS-II from 25 individuals with identical HVS-I motifs to increase the phylogenetic resolution and detect potential intra-site maternal kinship. The analysis of haplogroup defining coding region SNPs resulted in reproducible profiles for 96 individuals, with a success rate of 86% (electronic supplementary material, datasets S3 and S4). The haplotype of the Mesolithic skeleton from the Croatian Island Korčula could be assigned to mtDNA haplogroup U5b2a5 (electronic supplementary material, dataset S3). Sub-haplogroup U5b has been shown to be common in hunter–gatherer communities across Europe [28–30,32,33,47,48]. Contrary to the low mtDNA diversity observed in Central/North European hunter–gatherers [28–30], we identify a higher variability in early farming communities of the Carpathian Basin including haplogroups N1a, T1, T2, J, K, H, HV, V, W, X, U2, U3, U4 and U5a (electronic supplementary material, table S1). Previous studies described haplogroups N1a, T2, J, K, HV, V, W and X as being characteristic for the Central European LBK and suggested these as the mitochondrial ‘Neolithic package’ that had reached Central Europe in the sixth millennium BC [38,39]. Interestingly, most of these eight haplogroups show comparable frequencies between the STA, LBKT and LBK, and represent the majority of mtDNA variation in each culture (STA = 86.36%, LBKT = 61.54%, LBK = 79.63%) with similar haplotype diversity (STA = 0.97674, LBKT = 0.95277, LBK = 0.95483). By contrast, hunter–gatherer haplogroups are rare in the STA and both LBK groups (electronic supplementary material, table S1). Haplogroup H was not included in the Neolithic package, because it has also been found in pre-agricultural context in Iberia [48]. However, the low resolution of HVS-I does not allow to distinguish between H lineages of Neolithic or pre-Neolithic origins in Transdanubia and would require whole mitochondrial genome analyses. To evaluate whether the haplogroup and haplotype composition of the STA, LBKT, LBK [36–39] and hunter–gatherers from Central/North Europe [28–30,41] differ significantly from each other, we performed a haplogroup-based Fisher's exact test and a sequence-based genetic distance analysis. We also used a test of population continuity (TPC) [39] to elucidate whether the observed differences can be best explained by genetic drift or by other factors such as migration. These analyses reveal that the mtDNA composition of the Early Neolithic cultures is significantly different from that of the hunter–gatherers, both on the haplogroup (p = 0.0001) and haplotype level (Fst = 0.1764–0.187, p = 0.0000; electronic supplementary material, table S1), indicating genetic discontinuity of maternal lineages at the advent of farming in the Carpathian Basin. The TPC shows that, independent of the tested effective population size, the transition from foraging to farming cannot be explained by genetic drift alone (p < 0.000001; electronic supplementary material, dataset S13). More importantly, non-significant differences between the STA and the LBK groups from Transdanubia and Central Europe (haplogroup composition p = 0.0638–0.5518); haplotype composition (Fst = –0.00521–0.01345, p = 0.46581–0.62012) and TPC (p > 0.177; electronic supplementary material, table S1, dataset S13) support a rather homogeneous mtDNA signature of early farming communities from both regions and population continuity during the Neolithic period. We combined our Neolithic samples from the Carpathian Basin with 533 published mtDNA sequences from Upper Palaeolithic and Mesolithic [28–30,32,47,48], Neolithic [36–39,41,44,45,47–52] and Early Bronze Age [39] sites across Europe (electronic supplementary material, dataset S6) and analysed principal components (PCA), multidimensional scaling (MDS), molecular variance (AMOVA) and shared haplotypes to compare the mtDNA variability of the STA and LBKT in a broader geographical and chronological context (electronic supplementary material). The PCA and MDS show that the mtDNA composition of the STA and LBKT is strikingly similar to the LBK [36–39], eastern Hungarian Neolithic dataset [36,41,52] and to subsequent populations of the fifth/fourth millennia BC in Central Europe [39] (figure 2; electronic supplementary material, figures S1–S3, datasets S7–S10). This is predominately based on a high number of lineages attributed to the ‘Neolithic package’ and low frequencies of hunter–gatherers lineages, which clearly distinguish the cluster of farmers not only from hunter–gatherers of Central/North [28–30,41] and southwestern Europe [32,47,48], but also from Neolithic Iberian populations [44,45,47,48,50] and Central European cultures of the third/second millennia BC [39,49,51]. To exclude biases induced by potential maternal kinship within the prehistoric datasets, we also performed PCA and MDS with reduced datasets, excluding haplotypes with identical HVS-I and II sequences from the same site. The reduced and complete datasets fall closely next to each other, indicating a negligibly small bias of maternal kinship.  Figure 2. We used AMOVA to evaluate whether the observed affinities of STA and LBKT with the LBK and fifth/fourth millennia BC cultures from Central Europe are the result of a shared population structure. We pooled HVS-I sequences from the STA and LBKT and nine sixth–second millennia BC populations from Central Europe [36–39,49,51] into different groups, and tested 82 different constellations to identify the one with the highest among-group variance and simultaneously with low variation within the groups (electronic supplementary material, dataset S11). The highest among-group variance was observed when STA and LBKT was grouped with the Central European LBK and with all fifth/fourth millennia BC cultures, while the third/second millennia BC cultures were grouped separately (among-group variation = 3.50%, Fst = 0.03501, p = 0.00396; within-group variation = 0.20%, Fst = 0.00203, p = 0.31139). These results suggest a common genetic structure of the sixth–fourth millennia BC populations. We used shared haplotype analysis [53] and modified this approach by accounting for the temporal succession of cultures (termed ancestral shared haplotype analysis or ASHA; figure 3; electronic supplementary material, dataset S12). The results show that ancestral hunter–gatherer lineages were rare in the STA (2.27%), LBKT (0%) and LBK (1.85%), as well as in fifth/fourth millennia BC cultures (0%), and became more common in Central Europe during the third/second millennia BC (2.86–11.76%) [39]. By contrast, we identified a high proportion of ancestral STA lineages in all subsequent periods (LBKT = 61.54%, LBK = 55.56%, fifth/fourth millennia BC = 36.84–63.64%, third/second millennia BC = 36.17–43.18%). The subsequent LBKT reveals a smaller distinctive influence on its successors, as only 12.96% of the LBK, 0–10.53% of the fifth/fourth millennia BC and 0–3.19% of the third/second millennia BC cultures can be traced back to ancestral lineages first observed in the LBKT. The number of new ‘ancestral’ lineages is even lower in the LBK of Central Europe, with no effect on the third/second millennia BC populations. This highlights the importance of the STA in the formation of the mtDNA diversity in early farmers of Central Europe.  Figure 3. Results of the ancestral shared haplotype analysis. The bar plot shows the proportions of ancestral mtDNA lineages associated to hunter–gatherer, STA, LBKT, LBK and other subsequent populations in the Central/North European hunter–gatherer ... To identify affinities of our Neolithic datasets with present-day populations, we further collated 67 996 published HVS-I sequences from Eurasian populations, analysed principal components and mapped genetic distances (electronic supplementary material). The PCA shows that the frequencies of N1a, T1, T2, K, J and HV, and the absence of Asian and African lineages in the Carpathian Basin populations result in a clustering of the STA with populations of the Near East and the Caucasus, while the LBKT falls between the latter and populations from South and southeast Europe (Greeks, Bulgarians and Italians), which is explained by a higher frequency of haplogroup H in the LBKT (electronic supplementary material, figure S4, dataset S14). However, the dominant frequencies of haplogroup N1a, T2, and K in the STA and LBKT result in a differentiation from all present-day populations along the third component. Sequence-based genetic distance maps are largely consistent with PCA and reveal the greatest similarities of the STA to populations of the Near East (Iraq, Syria) and the Caucasus (Azerbaijan, Georgia, Armenia) as well as some European populations, such as Italy, Austria, Romania and Macedonia (electronic supplementary material, figure S5a, dataset S15). The LBKT displays affinities that are overall similar to the STA, which includes populations from Azerbaijan, Syria and Iraq (electronic supplementary material, figure S5b, dataset S15). We also observe similarities to present-day Europeans, such as the populations of Great Britain, Portugal, Romania, Crete and Russia. These similarity peaks are probably explained by elevated frequencies of shared lineages due to shared genetic drift in modern-day populations. |
|
|
|
Post by Admin on May 7, 2020 6:07:27 GMT
b) Y chromosomal DNA
We analysed 33 Y-haplogroup defining SNPs located on the non-recombining part of the Y chromosome (NRY), using multiplex [38] and singleplex PCR. We successfully generated unambiguous NRY SNP profiles for nine male individuals (STA = 7, LBKT = 2; electronic supplementary material, datasets S3 and S5). Three STA individuals belong to the NRY haplogroup F* (M89) and two specimens can be assigned to the haplogroup G2a2b (S126), and one each to G2a (P15) and I2a1 (P37.2). The two investigated LBKT samples carry haplogroups G2a2b (S126) and I1 (M253). Furthermore, incomplete SNP profiles of eight specimens potentially belong to the same haplogroups—STA: three G2a2b (S126), two G2a (P15) and one I (M170); LBKT: one G2a2b (S126) and one F* (M89).
G2a2b and F* are rare in present-day Europe. Frequencies of haplogroup G and its subgroups increase slightly towards the Near East and reach the highest values (70%) in populations of the west Caucasus [54–56], while G2a2b and its sub-clusters have been detected in modern-day Sardinia and Caucasus [56,57]. Haplogroup F* shows a diffuse dissemination pattern in Eurasia, which is based on its rarity and insufficient sub-haplogroup resolution in most population genetic studies. One STA F* haplogroup has been further analysed, and could be assigned to haplogroup H2 (L281) [40], which has been found in the extant Sardinian population [57]. Haplogroups I1 and I2a1 are most frequent in present-day populations of Europe, with the highest frequencies in Scandinavian [58–60] and southeast European populations, respectively [58].
We used PCA and genetic distance maps to identify affinities of the Carpathian Basin samples with 49 516 NRY SNP profiles from present-day Eurasian and African populations (electronic supplementary material). Owing to the small sample size and similarities in Y chromosome composition, we pooled ancient STA, Alföld LBK [41], LBKT and LBK data [38,40].
The elevated haplogroup G frequency in populations of the west Caucasus results in a clustering with the STA–LBK group on the second principal component. However, the predominant frequencies of haplogroups G and F* lead to a clear separation of the STA–LBK group from all present-day populations along the third principal component (electronic supplementary material, figure S6, dataset S16).
Similarly, the Y chromosome distance map shows the greatest similarities to populations of the west and south Caucasus, and Sardinians (electronic supplementary material, figure S7, dataset S17), which can be explained by the high frequency of haplogroup G/G2a [55,61] in these populations. This might reflect genetic drift, caused by isolation and small effective population size after a direct gene flow from the Near East, which led to a fixation of this haplogroup [54]. Intriguingly, populations of the northeast Caucasus show greater distances to the STA–LBK samples due to lower abundance of haplogroup G/G2a [55]. Genomic data from early farmers have been shown to be the most similar to modern-day Sardinians [33,40,41], which could be explained by the geographical isolation of Sardinia, having led to a conservation of the Neolithic genetic signature. Interestingly, our mtDNA population genetic analyses did not highlight the Neolithic–Sardinian affinity, but was observed on our genetic distance map. We caution that the haplogroup resolution of most modern comparative population datasets does not allow further differentiation between the sub-clusters of haplogroups C, F or G, and therefore does not provide sufficient Y-subgroup information at a smaller geographical scale.
3. Discussion
This study provides the first in-depth population survey of early farming cultures from the western Carpathian Basin and demonstrates its essential role in the genesis of the first farming communities of Central Europe. Population genetic analyses (Fisher's exact test, PCA, MDS, AMOVA, TPC) reveal similar mtDNA haplogroup composition and haplotype diversity in early farming populations from the Carpathian Basin and the populations of the Central European LBK, indicating a shared maternal genetic ancestry (figure 2; electronic supplementary material, table S1, figures S1–S3, dataset S11, S13).
The ASHA shows that about 55% of the LBK lineages ascribed to characteristic ‘Neolithic package’ haplogroups could be traced back to the STA and LBK in Transdanubia (figure 3; electronic supplementary material, dataset S12). It is therefore likely that this mtDNA signature was also present in ancient populations preceding the STA, in accordance with the archaeological record, which suggests cultural links to seventh/sixth millennia BC farming groups of the Aegean and the southern Balkans [5]. Interestingly, the STA mtDNA signature was still dominant in Neolithic cultures of the fifth/fourth millennia BC in Central Europe (figure 3; electronic supplementary material, dataset S12), attesting a direct and enduring genetic legacy of the STA/LBKT farmers in the Central European Neolithic, with minimal or no additional maternal genetic influence from outside for the subsequent 2500 years. Congruently with our results, recent genomic data also support the genetic homogeneity of early European farmers across large geographical distances [40].
Importantly, our comparative analyses with modern population data (PCA and genetic distance maps) emphasizes that both the mtDNA and NRY variability observed in the Carpathian Basin are most likely to have originated in the Near East with connections to the Caucasus (electronic supplementary material, figures S4–S7), in accordance with previous mtDNA studies of the Central European LBK [38,39]. The continuation of lineages through space and time suggests a scenario in which the genetic makeup of early farmers originated in the Near Eastern Fertile Crescent, from where it spread to Central Europe via the western Carpathian Basin, a region which acted as a natural corridor and a transient zone of adaptation to cooler climatic conditions along the Continental route. The shared Near Eastern affinities of the STA, LBKT and LBK, and the genetic continuity in the maternal and paternal gene pools, are consistent with the archaeological record, which describes the genesis of the early LBK (LBKT) from STA communities, followed by a rapid dispersal of the early LBK culture from Transdanubia towards the northwestern part of Central Europe [3,9,13]. We caution that use of modern-day data from the Near East could potentially lead to a biased interpretation. However, we still consider it the best available proxy of the prehistoric Near Eastern mtDNA variability, until aDNA datasets become available.
Recent studies using ancient genomic data of Central and North European early farmers have shown an affinity to South European rather than Near Eastern populations [33,40,41]. Of note, the proportion of western hunter–gatherer ancestry is higher in all modern-day Europeans and the early farmer ancestry being comparably best preserved in, for example, Sardinians [33,40]. These results are not necessarily contradictory but show that uniparental markers are more conservative, preserving a Near Eastern signature more consistently. Ancient genomic data from the Pre-Neolithic and Early Neolithic Near East will be needed to confirm these assumptions.
The very low frequencies of hunter–gatherer lineages (0–2.27%), in the STA, LBKT and LBK sample sets (figure 3) indicate that the arrival of agriculture in the Carpathian Basin and Central Europe was accompanied by a strong reduction of the currently known Mesolithic mtDNA substratum, resulting in a distinct and contrasting mtDNA haplogroup composition, and significant differences between hunter–gatherers and early farmers (figures (figures22 and and3;3; electronic supplementary material, figures S1–S3, table S1, datasets S7–S10, S12 and S13). This scenario is consistent with coalescent-based simulations that have revealed genetic discontinuity between Central European hunter–gatherers and LBK communities [28,38]. The detection of haplogroup U5b in the Mesolithic individual from Croatia matches previous observations, which describe sub-haplogroups of U as characteristic mtDNA substratum in Mesolithic Europe [28,39]. Residual Neolithic hunter–gatherer isolates, as reported from Central Europe by Bollongino et al. [30], probably also existed during the Early Neolithic of the Carpathian Basin, as shown by the hunter–gatherer ancestry of the sixth millennium BC human remains from an Early Neolithic Körös settlement pit in eastern Hungary [41]. According to the low proportion of hunter–gatherer mtDNA lineages in the LBK gene pool, we assume that admixture between hunter–gatherers and colonizing LBK farmers was also low in Central Europe.
Considering the relative size and speed of the LBK expansion, we have to presume a substantial population growth during the earliest LBKT, which might have resulted in a population pressure and led to emigration from Transdanubia [62]. Although such a radical population size increase was not palpable from the Early Neolithic archaeological records [7], recent extensive archaeological excavations have revealed large-scale LBKT settlements in western Hungary [9,63,64], suggesting larger source communities for expansion than previously assumed.
The discontinuity between hunter–gatherer and farmer ancestry is also visible in our Y chromosome results. Y chromosome study of modern-day Europeans has suggested a post-LGM expansion from a Franco-Cantabrian refugium for clade I1, and southeast European refugium for I2a1 based on high divergence time estimates [58]. I2a has indeed been found in Mesolithic and Neolithic Central and North European hunter–gatherers [33,34,40,41], as well as in Neolithic remains of southwestern Europe [44,45]. Haplogroup I2a (and possibly I1) might represent a pre-farming legacy of the NRY variation in Europe, alongside the recently described pre-Neolithic C (M130) haplogroups in Russia and Spain [35,42]. Y chromosome haplogroups from STA and LBKT samples, such as haplogroups G2a2b and F*, have also been reported from the Central European LBK [38], and support a homogeneity of paternal lineages among early farmers. Genetic studies on modern-day populations have discussed haplogroup G [25,65] and its subgroup G2a as potential representatives of the spread of farming from the Near East to Europe [26]. This scenario has recently been supported by Neolithic data from northern Spain [44] and southern France [45], which attested a pivotal role for G2a in the Neolithic expansion on the Mediterranean route. Furthermore, G2a has also been reported from four LBK individuals (G2a2a) [40] and the Tyrolean Iceman (G2a2a1b (L91)) [43]. Taken together, these findings suggest that sub-haplogroups of G2a were frequent in Neolithic populations of the sixth–fourth millennia BC across Europe. Thus, if we take Y chromosomal haplogroup I2a (and possibly I1) as proxy for a Mesolithic paternal genetic substratum in Europe, we observe a similar pattern to the changeover in the mitochondrial DNA variability, in which NRY G lineages were predominant among Neolithic farmers across Europe and I lineages became rare [38,40,43–45,49,51]. While the newly discovered NRY C haplogroup suggests additional Y lineages in eastern Hungary [41], the eastern Hungarian NRY data are still too few to estimate, whether there was paternal genetic difference between the populations of western and eastern Carpathian Basin.
The most characteristic mtDNA haplogroup of early farmers from the Carpathian Basin and Central Europe is N1a. N1a has previously been discussed as a potential marker of the spread of farming [36]. The presence of N1a in early farmers from the Carpathian Basin (6.82–10.26%) and Central Europe (12.04%; electronic supplementary material, table S1) lends further support to its pivotal role as a marker for the Continental route of the Neolithic expansion. On the other hand, mtDNA N1a and NRY G2a haplogroups are rare in present-day European populations, which is also reflected in the separation of the sixth millennium BC cultures from all present-day populations along the third principal component on the PCA plots (electronic supplementary material, figures S4 and S6). Intriguingly, R1a and R1b, which represent the most frequent European Y chromosome haplogroups today, are absent in early farmers of Central Europe (electronic supplementary material, dataset S20), but have been reported from cultures arriving in Europe during the third/second millennia BC [40,49,51]. These findings nonetheless indicate further demographic events after the Early/Middle Neolithic period that shaped modern-day mtDNA and NRY variability. However, we caution that the Y chromosome record is still very limited. Recent evidence from ancient mtDNA and genome-wide SNP analyses has stressed the role of further migrations from the eastern steppes reaching Central Europe during the third millennium BC [40], while three Bronze and Iron Age individuals from eastern Hungary suggest that both eastern and western influences reached the Carpathian Basin in post-Neolithic times [41].
Interestingly, recent model-based statistical analyses of contemporary NRY and mtDNA data, testing a series of population scenarios for the Neolithic transition, have revealed a shared admixture history for men and women, but not the same demographic history [66]. This study has shown that females had a larger effective population size, probably based on differential effects of social and cultural practices including increasing sedentism alongside a shift to monogamy and patrilocality in early farmer communities. It is therefore important to interpret our new uniparental genetic data in the light of those findings. Considering the entire set of published ancient NRY records available for Europe thus far, the low paternal diversity at the onset of the Neolithic is quite remarkable: G2a is the prevailing haplogroup (65.5%) in the sixth–fourth millennia BC Neolithic dataset (electronic supplementary material, datasets S5 and S20) [38,40,44,45]. The limited variation in NRY haplogroups in contrast to the high mtDNA haplogroup diversity suggests a smaller effective population size for males than females. One plausible explanation for this phenomenon is patrilocality (where women move to their husband's birth place after marriage), while other possibilities include polygyny or male-biased adult mortality. A patrilocal residential rule was probably linked to a system of descent along the father's line (patrilineality) in early farming communities. Ethnographic studies have suggested a change of residential rules at the advent of Neolithization, showing different trends in residential rules among modern foragers and non-foragers [67]. Increasing sedentism promotes territorial defence and control of resources, favouring men in the inheritance of land and property, which consequently leads to patrilocal residence [67]. However, residence patterns have to be temporarily flexible in expanding populations, allowing some of the sons to settle in new territories following population pressure and/or natural limitation of resources (e.g. after the carrying capacity of a particular region has been reached) [66]. Patrilocality has also been suggested in recent bioarchaeological studies, for example by investigating aDNA evidence from Neolithic site Treilles [45], or stable isotopes from LBK communities in Central Europe [68].
We note that patrilocality does not contradict the demic diffusion model, and it appears that both phenomena have left a discernible mark on the European Neolithic genetic diversity. While patrilocality and lineality might have caused high mtDNA and low NRY within population diversity, the demic diffusion model best explains the mtDNA and NRY affinity of the early farmers to the modern-day Near East, and the observed genetic homogeneity within southeastern and Central Europe. Importantly, local processes of sex-biased migration are unlikely to have an effect on genetic variation at broader spatial scales. Observations from many sites in Europe therefore argue for a common set of cultural and social practices across larger distances for early farming cultures in Europe. We caution that the observed differences in genetic diversity between males and females could also be influenced by resolution biases, resulting from the different sets of studied mtDNA and NRY markers. However, examining sex-specific dynamics of prehistoric societies emerges as an important area of ancient genetic research to be able to address underlying parameters such as migration rate, level of exogamy and distances of marriage-related dispersals.
The novel ancient data from early farming Neolithic populations of the Carpathian Basin help to fill the geographical gap on the Continental route of the Neolithic expansion from the Near Eastern Fertile Crescent to Central Europe. The joint analyses of mitochondrial and Y chromosomal DNA data support a demic diffusion of early farmer men and women through western Hungary, and demonstrate the paramount importance of this region as a prehistoric corridor of the migration. We point out that archaeological cultures of the Carpathian Basin provided the genetic basis of the first Central European farmers that affected subsequent prehistoric cultures for a long period of time. Considering our results in the light of newly available ancient genomic data, we observe a remarkable consistency of the different marker systems. Importantly, the new NRY data complement the sporadic European Y chromosomal dataset, and lend further support to patrilocal residential rules and patrilineal social system of the first farmers, underlining the role of demographic factors, which, depending strongly on cultural practices, notably shaped prehistoric and extant genetic diversity.
|
|





