Post by Admin on Nov 20, 2015 7:49:49 GMT

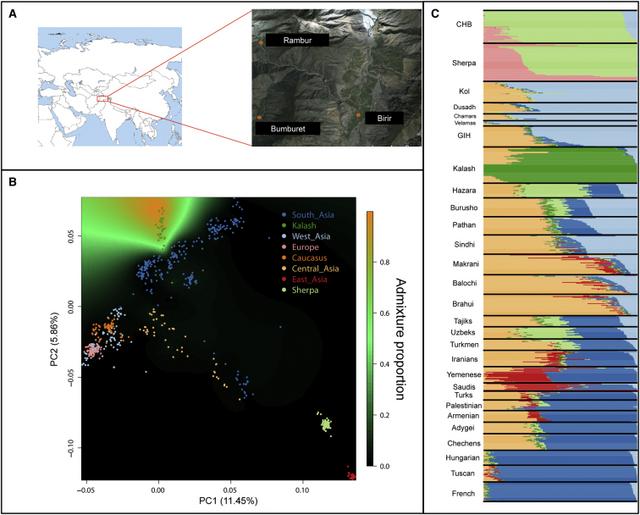
Figure 1 Population Structure and Isolation of the Kalash
(A) Geographic location of the three Pakistani villages where the Kalash samples were collected.
(B) Principal-component analysis (PCA) of Eurasian populations shows the first two components superimposed with the spatial kriging interpolation of the admixture coefficient of the Kalash genetic cluster. The proportion of admixture is indicated by color: orange represents the maximum level of admixture, and black represents the lowest. There is no gradient into the proportion of admixture with the Kalash cluster, suggesting a low level of gene flow between nearby populations and a high degree of isolation.
(C) Admixture analysis in which the lowest cross-validation error (k = 7) shows the unique Kalash cluster (dark green).
We assessed the genetic relatedness between three ancient genomes and modern human populations, including the Kalash, by computing outgroup f3 statistics. The measure circumvents potential bias from classical genetic relatedness tests, such as PCA (which has sample-size bias) and FST (which is sensitive to genetic drift that has occurred since divergence of the test populations), when using ancient genomes. According to outgroup f3 statistics, the Kalash share a high level of genetic drift with MA-1, a Paleolithic Siberian hunter-gatherer skeleton dated to ∼24,000 years ago, but not a very high level with La Braña 1, the Mesolithic European hunter-gatherer (skeletal remains dated to ∼7,000 years ago) or the European farmers represented by Ötzi, the Tyrolean Iceman dated to ∼5,300 years ago (Figure 3). Similar to Native Americans, the Kalash share a high proportion of genetic drift with MA-1. In comparison with other populations from Pakistan and India, the Kalash also share a higher proportion of genetic drift with La Braña 1 and Ötzi. The level of drift shared with La Braña 1 and Ötzi is comparable to that of other North European populations (Figure 3B). We also used TreeMix to estimate the proportion of Neandertal ancestry from the high-coverage archaic Altai Neandertal who lived ∼50,000 years ago. The jackknife estimate of Neandertal-to-Kalash gene flow was 2.4% ± 0.48%.
The Kalash represent a unique branch in the South Asian population tree and appear to be the earliest population to split from the ancestral Pakistani and Indian populations, indicating a complex scenario for population origins in the sub-continent rather than just the ancestral northern and southern Indian components identified previously.35 These Indo-European speakers were possibly the first migrants to arrive in the Indian sub-continent from northern or western Asia. This is supported by the higher level of shared genetic drift between the Kalash and the Paleolithic Siberian hunter-gatherer skeleton (MA-1) than between MA-1 and the other South Asian populations.

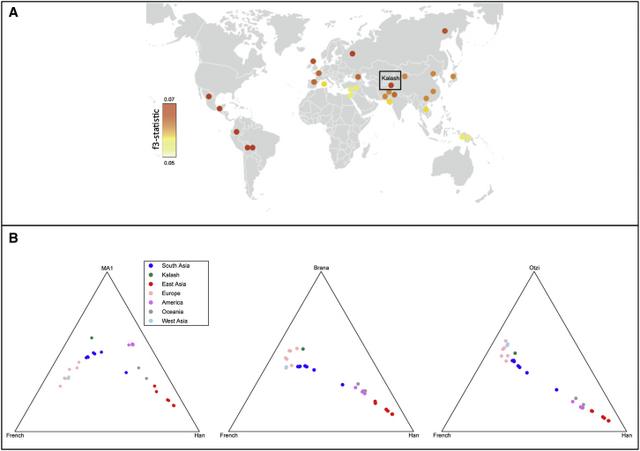
Figure 3 Shared Genetic Drift with Ancient Genomes
(A) Proportion of shared genetic drift (measured as f3 statistics) between extant world-wide HGDP-CEPH populations (including the Kalash) and the ancient Siberian hunter-gatherer (MA-1). The magnitude of the computed f3 statistics is represented by the graded heat key. The proportion of genetic drift shared between the Kalash and MA-1 is comparable to that shared between MA-1 and the Yakut, Native Americans, and northern European populations.
(B) Ternary plot of shared genetic drift with three ancient genomes: MA-1 (left), La Braña 1 (middle), and Ötzi, the Tyrolean Iceman (right). The high proportion of genetic drift shared between the Kalash and MA-1 is comparable to that shared between MA-1 and Native Americans. In comparison with other populations from South Asia, the Kalash also share a higher proportion of genetic drift with La Braña 1 and Ötzi.
Whereas the Kalash have recently been reported to have European admixture, postulated to be related to Alexander’s invasion of South Asia,6 our results show no evidence of admixture. Although several oral traditions claim that the Kalash are descendants of Alexander’s soldiers, this was not supported by Y chromosomal analysis in which the Kalash had a high proportion of Y haplogroup L3a lineages, which are characterized by having the derived allele for the PK3 Y-SNP and are not found elsewhere.7 They also have predominantly western Eurasian mitochondrial lineages and no genetic affiliation with East Asians.4

We observed that the Kalash share a substantial proportion of drift with a Paleolithic ancient Siberian hunter-gatherer, who has been suggested to represent a third northern Eurasian genetic ancestry component for present-day Europeans.36, 37 This is also supported by the shared drift observed between the Kalash and the Yamnaya, an ancient (2,000–1,800 BCE) Neolithic pastoralist culture that lived in the lower Volga and Don steppe lands of Russia and also shared ancestry with MA-1.36, 37 Thus, the Kalash could be considered a genetically drifted ancient northern Eurasian population, and this shared ancient component was probably misattributed to recent admixture with western Europeans.
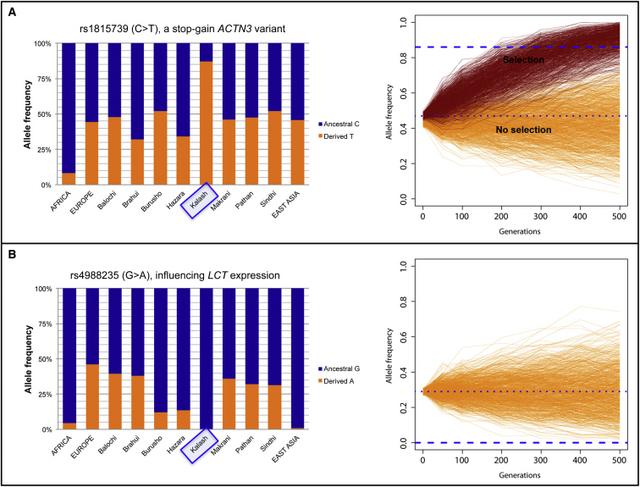
We also looked at how this long-term separation, isolation, and low effective population size affected the patterns of genetic variation in the Kalash. One striking example is the frequency of the derived allele for rs4988235, which has been linked to lactose tolerance. The Kalash, like the MA-1, are fixed for the ancestral allele for this variant, whereas their neighbors in Pakistan have been observed to have moderate frequencies of the derived allele. Although this supports their long-term isolation, it is surprising in other ways because the Kalash have no reported lactose intolerance and indeed celebrate a “milk day” during their annual spring rituals.38 This suggests that there might be additional derived lactase-persistence alleles in the LCT-MCM6 (MIM: 601806) region in this population.

Another example is the extremely high frequency (93%) of the stop-gain ACTN3 variant (rs1815739) associated with normal variation in human muscle strength and speed.39 This variant was picked up as an outlier in the PBS test for selection in the Kalash. Simulations indicated that such a high frequency of the derived allele in the Kalash can only be obtained under a scenario that includes positive selection. The variant might be relevant in cardiovascular conditioning and muscle strength related to climbing up and down high mountain passes. Although ACTN3 has not been associated with adaptation to high altitude, RYR2, another gene with an intronic outlier variant (rs2992644) in PBS, has.40
It has been postulated that South Asia, which is now a densely occupied land, was encountered by the first populations of modern humans that ventured out of Africa more than 50,000 years ago. The exact route taken by these earliest settlers is not known, although it has been suggested that they traveled via a southern coastal route.41, 42 The genetically isolated Kalash might be seen as descendants of the earliest migrants that took a route into Afghanistan and Pakistan and are most likely present-day genetically drifted representatives of these ancient northern Eurasians. A larger survey that includes populations from their ancestral homeland in Nuristan, Afghanistan, would provide more insights into their unique genetic structure and origins and help explain the complex history of the peopling of South Asia.
www.cell.com/ajhg/fulltext/S0002-9297(15)00137-8