Post by Admin on Mar 5, 2019 22:58:55 GMT
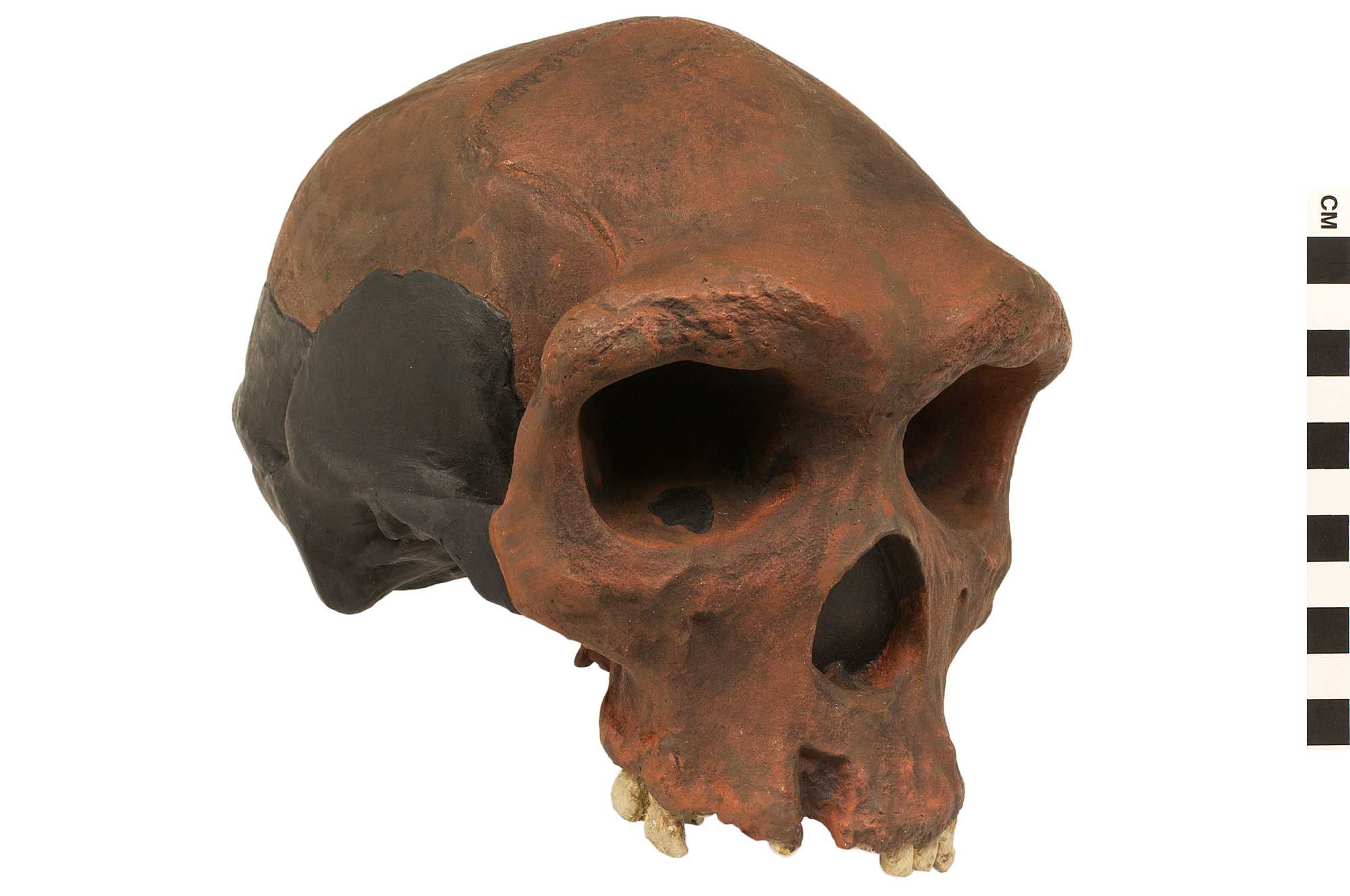
Kabwe 1
Nickname: Rhodesian Man
Site: Kabwe, Zambia Year of Discovery: 1921
Discovered by: Tom Zwiglaar
Age: Between 300,000 and 125,000 years old
Species: Homo heidelbergensis
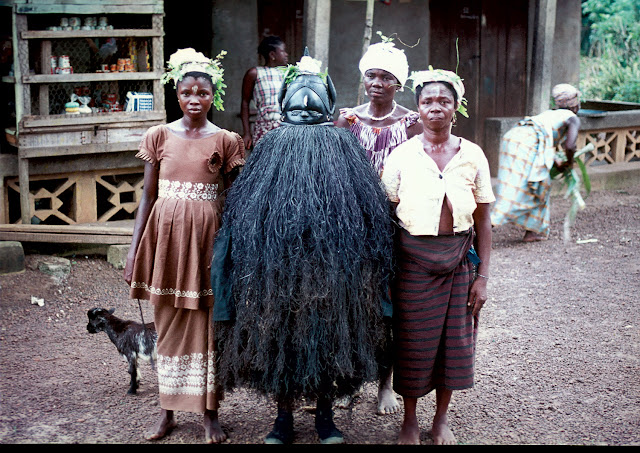
Searching for metal ore deposits in the limestone caves of Kabwe, Zambia, Swiss miner Tom Zwiglaar is credited with finding the first early human fossil ever to be discovered in Africa. When Kabwe (also known as Broken Hill) was sent to Arthur Smith Woodward, Woodward assigned the specimen to a new species: Homo rhodesiensis. Today, most scientists assign Kabwe to Homo heidelbergensis. Plaster cast of the cranium of Homo heidelbergensis, otherwise known as Rhodesian Man. Discovered in Kabwe (Broken HIll), Zambia, this species was once assigned as the type specimen for Homo rhodesiensis, although researchers now commonly attribute it to Homo heidelbergensis.

Kabwe shows features similar to H. erectus such as a low braincase profile (the area towards the back of the skull), large brow ridges, a slight widening of the midface known as the sagittal keel, and a protrusion at the back of the skull named the occipital torus. But Kabwe also resembles modern humans with a flatter, less prognathic face, and larger brain (1300 cubic centimeters). This skull is one of the oldest known to have tooth cavities. They occur in 10 of the upper teeth. The individual may have died from an infection related to dental disease or from a chronic ear infection.

Putative H. heidelbergensis specimens include African examples such as Bodo (Ethiopia) and Broken Hill (Zambia), European fossils such as Petralona (Greece) and Arago and possibly also Asian specimens such as Dali (China) and Narmada (India). Some argue that the Mauer mandible should not be used to name this larger Mid-Pleistocene group. As the quote (attributed to William Straus) goes: “While the skull is the creation of God, the jaw is the work of the devil.” These researchers agree that there are similarities between some of these Mid-Pleistocene fossils, but point out that basing the species definition on an isolated mandible is problematic. Not only is mandibular material lacking for most of the other fossils, but the mandible does not have a large number of taxonomically diagnostic features. If Mauer is excluded from the taxon likely to be the last common ancestor, the name H. rhodesiensis would take priority if the Broken Hill fossil is included. However, a study on Pleistocene and modern human mandibles argues that a suite of diagnostic features can be identified that support the grouping of these Mid-Pleistocene fossils with Mauer, and thus the name H. heidelbergensis is appropriate for the last common ancestor.

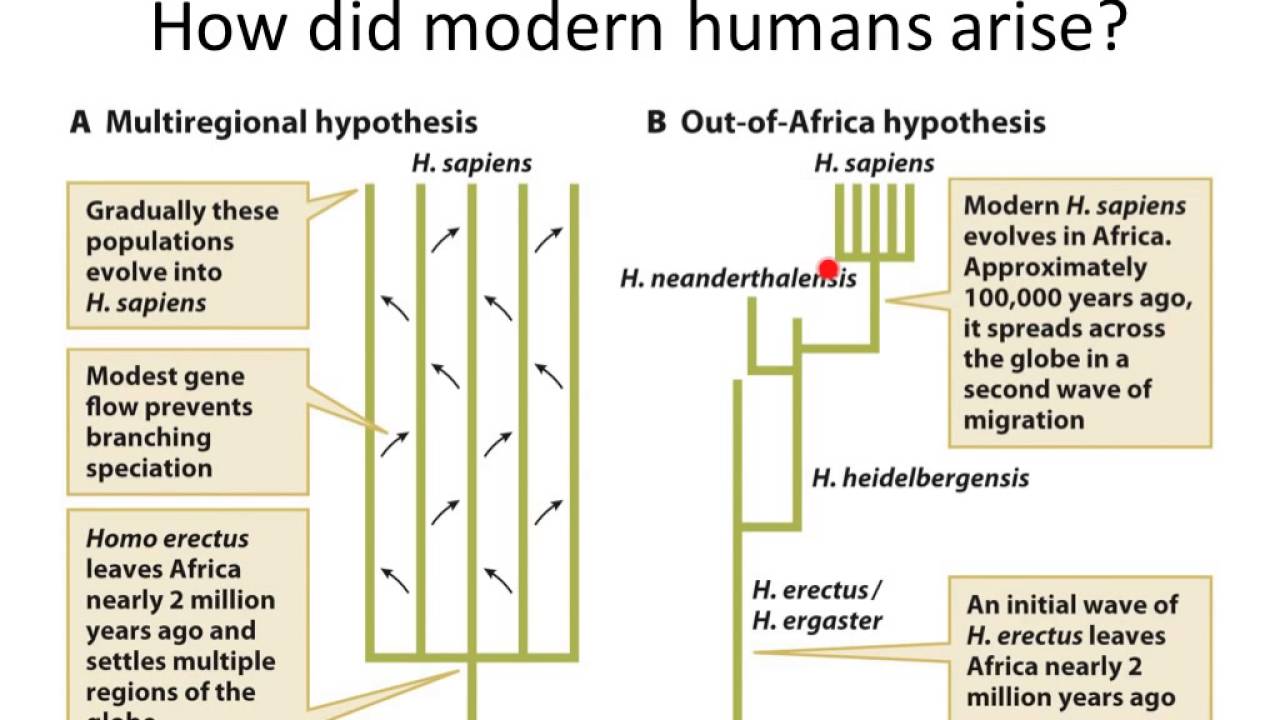
Figure 1. The two main hypotheses for the taxonomic placement of H. heidelbergensis.
(A) European H. heidelbergensis as the exclusive ancestor of Neanderthals and the related Denisovans. (B) European H. heidelbergensis as the last common ancestor of modern humans, Neanderthals and the related Denisovans. Pictures: H. antecessor Wikimedia Commons; H. heidelbergensis (Arago) in Figure 1A: Chris Stringer; all other images with kind permission of the Natural History Museum, London.
Because of the presence of some claimed Neanderthal traits, such as in the morphology of the teeth and the shape of the face, in European Mid-Pleistocene specimens, some researchers have argued that H. heidelbergensis is the ancestor of Neanderthals to the exclusion of the African fossils and H. sapiens (Figure 1A). In this scenario, similarities between African and European Mid-Pleistocene hominins are merely primitive traits inherited from H. erectus; H. sapiens is descended from a separate African Mid-Pleistocene species (possibly H. rhodesiensis), and the last common ancestor of H. sapiens and Neanderthals would lie chronologically further back, perhaps in the form of H. antecessor, thus far only recognised from the site of Gran Dolina at Atapuerca (Spain) at ∼800 ka. The hypothesis of H. heidelbergensis as uniquely ancestral to the Neanderthals is highly dependent on the inclusion of the remains from another site at Atapuerca, the controversial Sima de los Huesos (‘Pit of the bones’), which is where most of the fossils with claimed Neanderthal affinities come from. The more than 6000 fossils from that site show distinctive Neanderthal features, but have often been included in the H. heidelbergensis taxon because of their supposed great age (up to 600 ka). Further research has now suggested that the material looks too Neanderthal and is too young (∼400 ka) to represent H. heidelbergensis, making these fossils early Neanderthals instead. However, adding extra complexity, recent findings reveal the Sima people’s mitochondrial DNA to be more similar to that of the Denisovans (see below) than Neanderthals.
African H. heidelbergensis material, such as Broken Hill, shares numerous features with European fossils such as Petralona, leading many to group them together. As long as Mauer is also included, this taxon can be named H. heidelbergensis. Proponents of this wide concept of H. heidelbergensis assert that the mosaic of primitive and derived features shared by this group of fossils is unique, with few traits linking them exclusively to either modern humans or Neanderthals (Figure 1B). H. heidelbergensis is thus hypothesised to be the last common ancestor of both Neanderthals in Eurasia and H. sapiens in Africa. This scenario is probably the most popular and well supported at present.
In 2010, DNA sequenced from a fossil finger bone in Siberia showed the existence of an unexpected additional human population in the late Pleistocene. This group was christened the ‘Denisovans’ after the site of Denisova Cave. We still don’t know what the Denisovans looked like, or completely understand their position relative to other species, but genomic data suggest they are related to the Neanderthals (Figure 1A,B). However, evidence of interbreeding with recent H. sapiens in Southeast Asia shows that the Denisovans were once widespread in the region. As there are potential H. heidelbergensis fossils from Asia, it is possible they could represent the ancestors of the Denisovans. Recent advances in ancient DNA extraction and processing mean that recovering diagnostic genetic material from mid-Pleistocene fossils such as the remains from Asia is now an exciting possibility. The geographic origin of H. heidelbergensis is still unknown, but the early fossils from Asia suggest that continent is as likely a place of origin as Europe or Africa at the moment. An Asian origin for a species directly ancestral to our own would certainly shake up the current rather Afro-centric view of our evolution.
According to one theory, Neanderthals, Denisovans, and modern humans are all descended from the ancient human Homo heidelbergensis. Between 300,000 to 400,000 years ago, an ancestral group of H. heidelbergensis left Africa and then split shortly after. One branch ventured northwestward into West Asia and Europe and became the Neanderthals. The other branch moved east, becoming Denisovans. By 130,000 years ago, H. heidelbergensis in Africa had become Homo sapiens who did not begin their own exodus from Africa until about 60,000 years ago. Another branch of H. heidelbergensis could have remained in Africa until 40,000 years ago and interbred with sub-Saharan African populations until they died out. As a result, Africans derive 2-19% of their genetic ancestry from an archaic population that could possibly be H. heidelbergensis.
Homo sapiens originated from a single, common ancestor that lived in Africa 300 000 years ago. Our African origins have been demonstrated countless times by genetic analyses and fossil evidence. But what's known as the multiregional model suggests that modern humans evolved independently of each other from different pre-human populations. Asians originated from the Asian Homo erectus, Europeans from the Neanderthal man, and Africans from the African Homo heidelbergensis. The multiregional model may be partially right with regard to Europeans' Neanderthal ancestry (2-3%) and H. Heidelbergensis introgression in Africans (2-19%). While there is no genetic evidence that East Asians are descended from the Asian Homo erectus, they are up to 5% Neanderthal and 0.2% Denisovan. SE Asians and Australoids are 5-6% Denisovan and they may harbor some H. erectus ancestry, too.
Multiple present-day African populations inherited genes from an unknown archaic population that diverged
before modern humans and Neanderthals split.
While introgression from Neanderthals and Denisovans has been well-documented in modern humans outside
Africa, the contribution of archaic hominins to the genetic variation of present-day Africans remains poorly
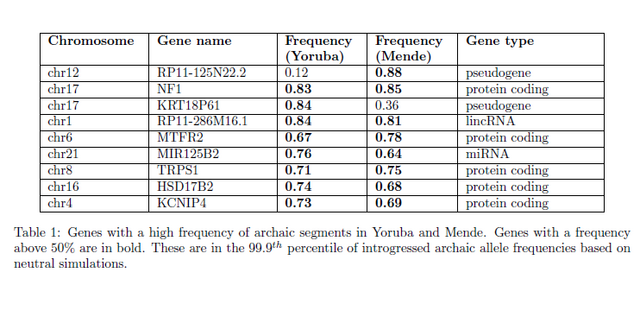
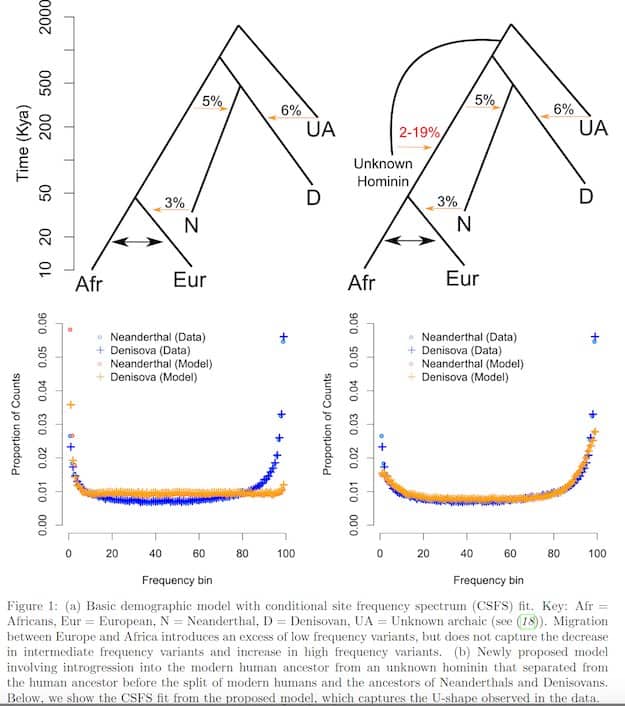
understood. Using 405 whole-genome sequences from four sub-Saharan African populations, we provide complementary lines of evidence for archaic introgression into these populations. Our analyses of site frequency spectra indicate that these populations derive 2-19% of their genetic ancestry from an archaic population that diverged prior to the split of Neanderthals and modern humans. Using a method that can identify segments of archaic ancestry without the need for reference archaic genomes, we built genome-wide maps of archaic ancestry in the Yoruba and the Mende populations that recover about 482 and 502 megabases of archaic sequence, respectively. Analyses of these maps reveal segments of archaic ancestry at high frequency in these populations that represent potential targets of adaptive introgression. Our results reveal the substantial contribution of archaic ancestry in shaping the gene pool of present-day African populations.
Admixture has been a dominant force in shaping patterns of genetic variation in human populations (1).
Comparisons of genome sequences from archaic hominins to those from present-day humans have documented multiple interbreeding events including gene fow from Neanderthals into the ancestors of all non-Africans (2),
from Denisovans into Oceanians (3) and eastern non-Africans (4, 5), as well as from early modern humans
into the Neanderthals (6). However, the sparse fossil record and the difficulty in obtaining ancient DNA
30 have made it challenging to dissect the contribution of archaic hominins to genetic diversity within Africa.
While several studies have revealed contributions from deep lineages to the ancestry of present-day Africans
(7, 8, 9, 10, 11, 12), the nature of these contributions remains poorly understood.
We leveraged whole genome sequence data from present-day African populations as well as archaic hominins
to compute statistics that are sensitive to introgression in the history of African populations. Specifically, we tabulated the distribution of the frequencies of derived alleles (where a derived allele is determined
relative to an inferred human ancestor) in African populations at single nucleotide polymorphisms (SNPs)
for which a randomly sampled allele from an archaic individual was observed to also be derived. Theory
predicts that this conditional site frequency spectrum (CSFS) is expected to be uniformly distributed when
alleles are neutrally evolving under a demographic model in which the ancestor of modern and archaic hu40
mans, assumed to be at mutation-drift equilibrium, split with no subsequent gene fow between the two
groups (13, 14). Importantly, this expectation is robust to assumptions about changes in population sizes
in the history of modern human or archaic populations. Further, we show that this expectation holds even
when there is population structure or gene fow in the history of the archaic population (SI Section S1).
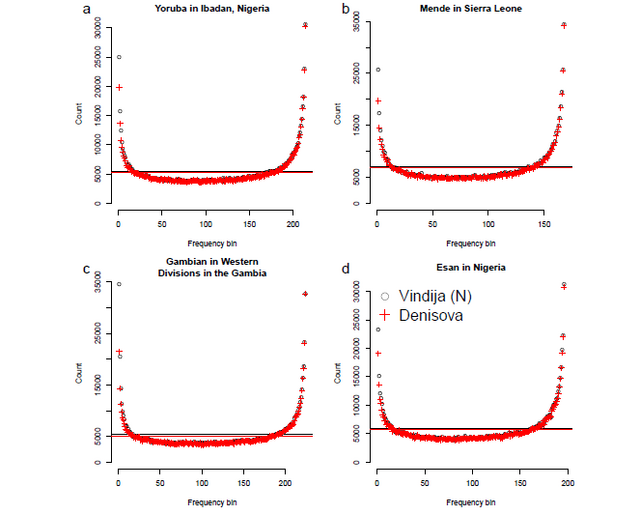
We computed CSFSYRI;N: the CSFS in the Yoruba from Ibadan (YRI) while restricting to SNPs where
a randomly sampled allele from the high-coverage Vindija Neanderthal (N) genome was observed to be
derived (15). In contrast to the uniform spectrum expected from theory, we observe that the CSFSYRI;N has
a U-shape with an elevated proportion of SNPs with low and high frequency derived alleles relative to those
at intermediate frequencies (Figure 1, Figure S4). The CSFS is nearly identical when we replace the Vindija
Neanderthal genome with the high-coverage Denisova genome (4) (Figure 1, Figure S4). We observed a
similar U-shaped CSFS in each of three additional African populations (Esan in Nigeria [ESN], Gambian in
Western Divisions in the Gambia [GWD], and Mende in Sierra Leone [MSL]) included in the 1000 Genomes
Phase 3 data set (Figure S4).