Post by Admin on Apr 6, 2019 18:43:21 GMT
Typing of Y-chromosome polymorphisms
DNA samples were extracted from peripheral blood using the standard salting-out method.13 Initially, 16 bi-allelic Y-chromosome polymorphisms were used to identify the major haplogroups (Fig. 1A).14 , 15 Haplogroup designation follows the nomenclature proposed by the Y Chromosome Consortium.16 Subsequently, haplogroup J* defined by the M304 mutation, was resolved further after screening for the mutations M267, M172, P58, M410, M318 and M1212 using a single-base extension method.17
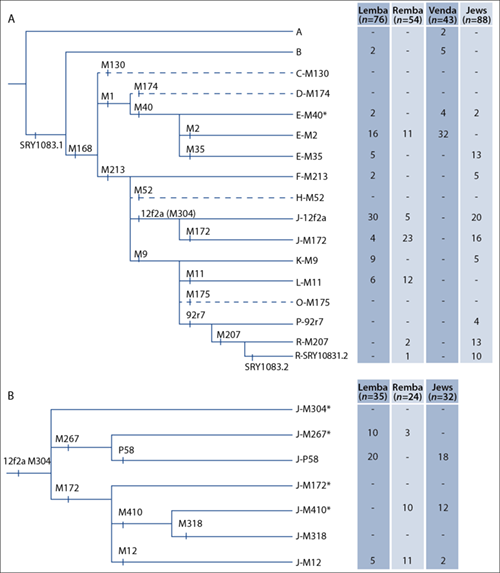
Fig. 1. (A) Phylogeny and distribution of Y chromosome haplogroups defined by the 16 bi-allelic markers used in this study (black bars) in the Lemba, Remba, Venda and South African Jewish populations. The 13 haplogroups found in the total sample are indicated with solid lines, while those haplogroups not found in the sample are shown using dashed lines. (B) Phylogeny showing the finer resolution of haplogroup J chromosomes using 6 additional single nucleotide polymorphisms and the number of individuals from a subset of Lemba, Remba and Jewish samples with these sub-haplogroups.
Haplotypes were resolved using both bi-allelic and STR polymorphisms. The original CMH was defined using 6 STR loci: DYS19, DYS388, DYS390, DYS391, DYS392 and DYS393 (in this order) that were amplified in two separate multiplex polymerase chain reactions (PCR).18 The 12 STRs (DYS19, DYS385a, DYS385b, DYS388, DYS389I, DYS389II, DYS390, DYS391, DYS392, DYS393, DYS426 and DYS439 that defined the extended CMH in this order) were genotyped in a subset of 91 individuals who were found to have haplogroup J-12f2a or J-M172 Y chromosomes. This was accomplished by using AmpFISTR Y-filer (Applied Biosystems) which amplified 17 Y-STRs: DYS19, DYS389I, DYS389II, DYS390, DYS391, DYS392, DYS393, DYS385a, DYS385b, DYS437, DYS438, DYS439, DYS456, DYS458, DYS635 and YGATA H4 (underlined loci part of the extended CMH) and the Multiplex II PCR amplification kit (Applied Biosystems) that included the two STRs DYS388 and DYS426.
A comparison of the haplogroup distribution among Remba, Lemba, Venda and Jews revealed that Remba and Lemba had some haplogroups in common with the Venda and others with Jews (Fig. 1A). Of the 11 haplogroups found in the combined Remba/Lemba sample, only four (E-M2, J-12f2a, J-M172 and L-M11) were shared between them. Although the four haplogroups together constituted a major component of the Y chromosomes in both Lemba (73.7%) and Remba (94.4%), the frequencies of these haplogroups (with the exception of haplogroup E-M2) differed in the two populations (Fig. 1A). The most common haplogroup in the Remba was J-M172, which accounted for 42.6% of their Y chromosomes, whereas J-12f2a was most common in the Lemba (39.5%). Both J haplogroups were observed in Jews at frequencies of 18.1% and 22.7%, respectively. The CMH (14-16-23-10-11-12), which was resolved on haplogroup J-12f2a was only found in the Lemba (13.2%) and Jewish (15.9%) groups, but not in the Remba (H54, Appendix 1).

With the exception of L-M11, which was only found in Lemba and Remba, all haplogroups derived from non-African ancestry in the combined Lemba/Remba sample were also found in Jews.
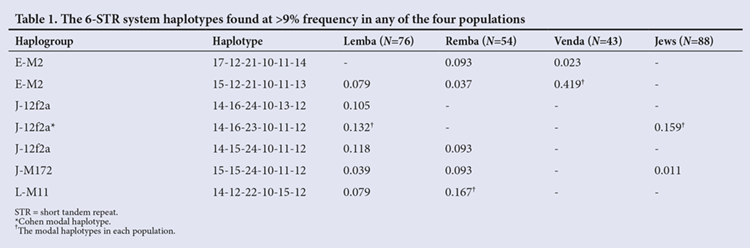
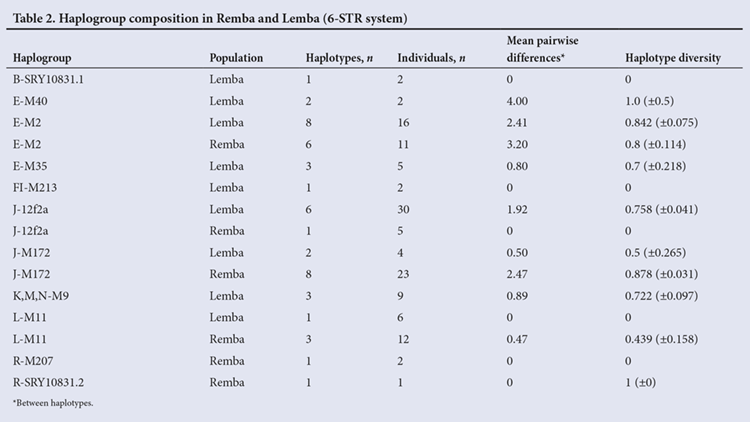
Using the exact test of differentiation, all population pairs, including the Remba v. Lemba, were found to be significantly different from each other using both haplogroup (p<0.001) and haplotype (p<0.001) frequency data. Only 5 (H26, H58, H65, H67, H84) of the 42 haplotypes found in the combined Lemba/Remba sample were shared between the two groups (Appendix 1). Altogether, these lineages accounted for 39.2% of the Y chromosomes in the Lemba/Remba and they shared predominantly higher-frequency haplotypes (Table 1).
There was little divergence, as reflected in the small mean pairwise differences and low haplotype diversity values (Table 2), within haplogroups found in the Lemba/Remba. Several haplogroups were represented only by a single haplotype, some of them at high frequency (Table 2). Other haplogroups comprised one major haplotype from which few one-step neighbours had evolved (Appendix 1).
Finer resolution of haplogroup J* and the extended CMH
Screening for the 6 binary markers within haplogroup J* in 35 Lemba, 24 Remba and 32 SA Jews resolved haplogroup J* into 4 sub-haplogroups (Fig. 1B). Haplogroup J-P58 which is associated with the CMH was found in the Lemba (57.1%) and SA Jews (56.3%). Haplogroup J-M267* was found in the Lemba (28.6%) and Remba (16.0%) but not in the SA Jews. Haplogroup J-M172 was further resolved into sub-haplogroups J-M410* in the Remba (43.5%) and SA Jews (37.5%) and J-M12* in the Lemba (14.3%), Remba (44.4%) and SA Jews (6.3%).
The 12-STR marker system resolved the 91 M304 Y chromosomes (Fig. 1B) into 46 haplotypes (Appendix 2). The extended CMH (14-13-15-16-13-30-23-10-11-12-11-12) on the J-P58 haplogroup background was only found in 1/14 Jewish individuals who were found to have the original CMH (6 STRs). None of the 10 Lemba who were found to have the original CMH had the extended CMH. The most common haplotype 14-13-19-16-13-29-23-10-11-12-11-12 found in eight Lemba differed from the extended CMH by 5 mutational steps; 4 at the DYS385b locus and 1 at the DYS389II locus. The closest haplotype in the Lemba to the extended CMH differed by 4 mutational steps (3 at the DYS385b locus and 1 at the DYS389II locus).
At this higher level of resolution at both haplogroup and haplotype level, the Lemba and Remba only shared 3 haplotypes – 1 on the J-M267* background and 2 on the J-M12 background (Appendix 2). None of the combined Lemba/Remba haplotypes were shared with the Jews.
DNA samples were extracted from peripheral blood using the standard salting-out method.13 Initially, 16 bi-allelic Y-chromosome polymorphisms were used to identify the major haplogroups (Fig. 1A).14 , 15 Haplogroup designation follows the nomenclature proposed by the Y Chromosome Consortium.16 Subsequently, haplogroup J* defined by the M304 mutation, was resolved further after screening for the mutations M267, M172, P58, M410, M318 and M1212 using a single-base extension method.17
Fig. 1. (A) Phylogeny and distribution of Y chromosome haplogroups defined by the 16 bi-allelic markers used in this study (black bars) in the Lemba, Remba, Venda and South African Jewish populations. The 13 haplogroups found in the total sample are indicated with solid lines, while those haplogroups not found in the sample are shown using dashed lines. (B) Phylogeny showing the finer resolution of haplogroup J chromosomes using 6 additional single nucleotide polymorphisms and the number of individuals from a subset of Lemba, Remba and Jewish samples with these sub-haplogroups.
Haplotypes were resolved using both bi-allelic and STR polymorphisms. The original CMH was defined using 6 STR loci: DYS19, DYS388, DYS390, DYS391, DYS392 and DYS393 (in this order) that were amplified in two separate multiplex polymerase chain reactions (PCR).18 The 12 STRs (DYS19, DYS385a, DYS385b, DYS388, DYS389I, DYS389II, DYS390, DYS391, DYS392, DYS393, DYS426 and DYS439 that defined the extended CMH in this order) were genotyped in a subset of 91 individuals who were found to have haplogroup J-12f2a or J-M172 Y chromosomes. This was accomplished by using AmpFISTR Y-filer (Applied Biosystems) which amplified 17 Y-STRs: DYS19, DYS389I, DYS389II, DYS390, DYS391, DYS392, DYS393, DYS385a, DYS385b, DYS437, DYS438, DYS439, DYS456, DYS458, DYS635 and YGATA H4 (underlined loci part of the extended CMH) and the Multiplex II PCR amplification kit (Applied Biosystems) that included the two STRs DYS388 and DYS426.
A comparison of the haplogroup distribution among Remba, Lemba, Venda and Jews revealed that Remba and Lemba had some haplogroups in common with the Venda and others with Jews (Fig. 1A). Of the 11 haplogroups found in the combined Remba/Lemba sample, only four (E-M2, J-12f2a, J-M172 and L-M11) were shared between them. Although the four haplogroups together constituted a major component of the Y chromosomes in both Lemba (73.7%) and Remba (94.4%), the frequencies of these haplogroups (with the exception of haplogroup E-M2) differed in the two populations (Fig. 1A). The most common haplogroup in the Remba was J-M172, which accounted for 42.6% of their Y chromosomes, whereas J-12f2a was most common in the Lemba (39.5%). Both J haplogroups were observed in Jews at frequencies of 18.1% and 22.7%, respectively. The CMH (14-16-23-10-11-12), which was resolved on haplogroup J-12f2a was only found in the Lemba (13.2%) and Jewish (15.9%) groups, but not in the Remba (H54, Appendix 1).
With the exception of L-M11, which was only found in Lemba and Remba, all haplogroups derived from non-African ancestry in the combined Lemba/Remba sample were also found in Jews.
Using the exact test of differentiation, all population pairs, including the Remba v. Lemba, were found to be significantly different from each other using both haplogroup (p<0.001) and haplotype (p<0.001) frequency data. Only 5 (H26, H58, H65, H67, H84) of the 42 haplotypes found in the combined Lemba/Remba sample were shared between the two groups (Appendix 1). Altogether, these lineages accounted for 39.2% of the Y chromosomes in the Lemba/Remba and they shared predominantly higher-frequency haplotypes (Table 1).
There was little divergence, as reflected in the small mean pairwise differences and low haplotype diversity values (Table 2), within haplogroups found in the Lemba/Remba. Several haplogroups were represented only by a single haplotype, some of them at high frequency (Table 2). Other haplogroups comprised one major haplotype from which few one-step neighbours had evolved (Appendix 1).
Finer resolution of haplogroup J* and the extended CMH
Screening for the 6 binary markers within haplogroup J* in 35 Lemba, 24 Remba and 32 SA Jews resolved haplogroup J* into 4 sub-haplogroups (Fig. 1B). Haplogroup J-P58 which is associated with the CMH was found in the Lemba (57.1%) and SA Jews (56.3%). Haplogroup J-M267* was found in the Lemba (28.6%) and Remba (16.0%) but not in the SA Jews. Haplogroup J-M172 was further resolved into sub-haplogroups J-M410* in the Remba (43.5%) and SA Jews (37.5%) and J-M12* in the Lemba (14.3%), Remba (44.4%) and SA Jews (6.3%).
The 12-STR marker system resolved the 91 M304 Y chromosomes (Fig. 1B) into 46 haplotypes (Appendix 2). The extended CMH (14-13-15-16-13-30-23-10-11-12-11-12) on the J-P58 haplogroup background was only found in 1/14 Jewish individuals who were found to have the original CMH (6 STRs). None of the 10 Lemba who were found to have the original CMH had the extended CMH. The most common haplotype 14-13-19-16-13-29-23-10-11-12-11-12 found in eight Lemba differed from the extended CMH by 5 mutational steps; 4 at the DYS385b locus and 1 at the DYS389II locus. The closest haplotype in the Lemba to the extended CMH differed by 4 mutational steps (3 at the DYS385b locus and 1 at the DYS389II locus).
At this higher level of resolution at both haplogroup and haplotype level, the Lemba and Remba only shared 3 haplotypes – 1 on the J-M267* background and 2 on the J-M12 background (Appendix 2). None of the combined Lemba/Remba haplotypes were shared with the Jews.