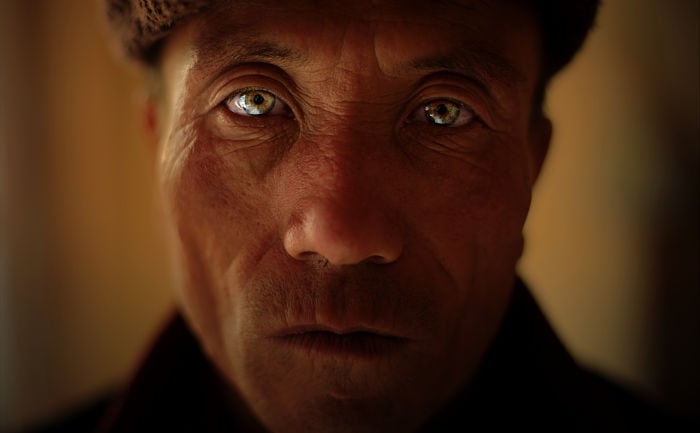
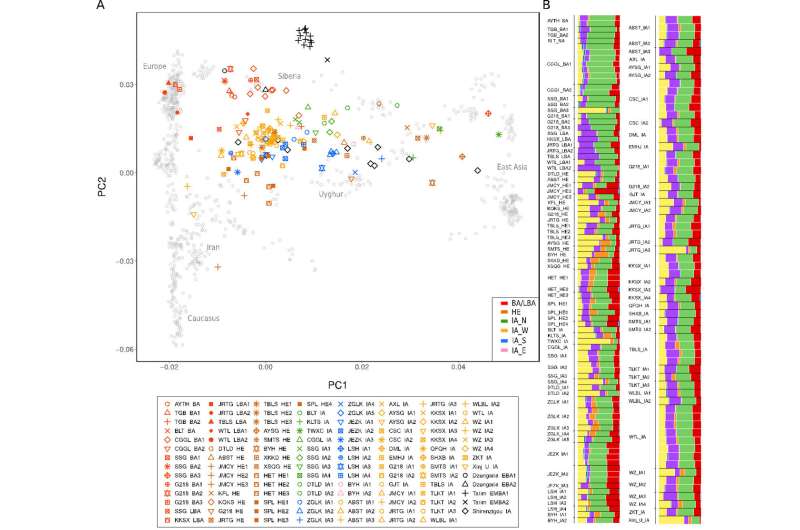
These Koreans who are naturally born with light eyes may be endowed with the R1a haplogroup commonly found in Russia. Figure 2C shows that the overall frequency of R1a and P*(xR1a) is 2-3% in Korea and there are genetic ties between the Uyghurs in western China, who are known for Caucasian looks, and ethnic Koreans. Moreover, the origins of the Yemaek tribes in Korea may be traced back to the Altai region, where Russia, China and Mongolia come together, and the ancient Korean tribes weren't indigenous to the Korean peninsula. The Songhua River basin in northern China was once a home to the Yemaek tribes, where they interacted with the nomadic steppe herders, who developed a sophisticated kurgan culture.
13% of Mongolians also belong to R1a and this Mongolian girl looks half Russian. Recent genetic evidence suggests that a group of ancient Russians called Yamna steppe herders from the Pontic-Caspian steppe historically migrated to Central Asia, leaving their genetic footprints on Central Asian populations.

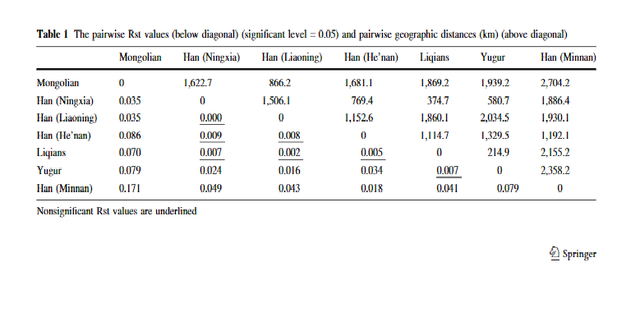
Approximately 1000 males from 27 East Asian populations were typed with 61 Y-chromosomal markers, and we first describe the basic properties of this data set. The 45 binary markers identified 31 haplogroups (including paragroups) in the sample, while the 15 STRs defined 730 different haplotypes (Figure 1, Table 1; see also supplemental Table 1 at
www.genetics.org/supplemental/). Population diversities ranged from 0.60 to 0.94 for binary markers and from 0.84 to 1.00 for STRs (Table 2). There was considerable variation in the distribution of lineages between populations, but this did not correspond to the major ethnic distinction in the area, which is between the Han Chinese (>80% of the combined populations of China, Mongolia, Korea, and Japan) and the other populations. AMOVA analysis showed that only 1.8 and 0.5% of variation lay between Han and non-Han populations using binary and STR markers, respectively, and neither of these values was significantly greater than zero. There were, however, major geographical differences. Figure 2 shows that, despite the overall predominance of haplogroup O (56%), specific haplogroups were concentrated in each geographical region: C and N in the north; P and J in the west; O2b in the east; and O1*, O2*, and O3d in the south. We therefore wished to identify the most important elements of the geographical pattern in an objective way.
We based the subsequent analyses on the STR data unless otherwise indicated because of the problems in interpreting data from preascertained binary markers. SAMOVA analysis (Dupanloup et al. 2002) identifies, for a prespecified number of groups of populations, the geographical groups that are most differentiated from one another. Application of this method to the East Asian Y-STR data set using two or three groups distinguished small numbers of unusual populations, a finding that is readily understood from the high frequencies of the “star cluster” (Zerjal et al. 2003) and “Manchu cluster” (Xue et al. 2005) lineages in some northern populations, and reflects extreme expansions of individual patrilines within historical times. The use of four groups provided the most informative subdivision, with a cluster of six southern populations distinguished in addition to some of the northern ones (Figure 3A). This pattern corresponds well to the north–south distinction seen with classical markers and shows that, in this respect, the Y-chromosomal variation is typical of that on other chromosomes. The division of the sample into more groups led to further subdivisions in the south (e.g., Figure 3B). Spatial autocorrelation analysis (Bertorelle and Barbujani 1995), based on the binary marker variation, produced correlograms that indicated significant clinal patterns or long-distance differentiation (not shown). The north–south haplogroup structure is therefore a continuum rather than a sharp bipartite division. To understand it further, we have explored the characteristics of the populations in more detail, concentrating on the 22 non-Han populations because of the spread of the Han during historical times (Wen et al. 2004).
A simple property of a population is the variation it contains, and this can be expressed in a number of ways. A widely used measure, diversity, is so high when 15 STRs are used that the differences between populations are small (Table 2) and difficult to interpret. Reducing the number of STRs to an arbitrary four or three (Table 2, supplemental Figure 1 at
www.genetics.org/supplemental/) produces a wider range of diversity values, and these are notably higher in the north than in the south. An alternative measure of variation within a population, average squared distance (ASD), shows a similar pattern. BATWING analysis allows demographic parameters of the populations to be explored. Using a model where the population size remains constant for a period and then begins to expand exponentially, we estimated, for each population, posterior values of (1) the effective population size during the constant period, Nposterior; (2) the time at which growth began; (3) the rate of growth per generation, α; and (4) the time to the most recent common ancestor (TMRCA) of the population (Table 2). We again noted substantial variation with latitude. Median Nposterior was higher in the north, the expansion began earlier, the rate of growth was slower, and the TMRCA was longer. Although all of these variables correlated significantly with latitude when examined individually in regression analyses (Table 3), the highest was with expansion time (adjusted R2 = 0.68), compared with 0.40 for the next highest, ASD. Unsurprisingly, a stepwise multiple regression analysis identified expansion time as the best predictor of north–south distance, and only α increased this significantly to reach an adjusted R2-value of 0.75. Thus earlier expansion time in the north and, to a lesser extent, more rapid expansion in the south, account best for the observed north–south differences. We display the expansion times as a contour plot in Figure 4, where the consistent difference between north and south is apparent. Figure 4 suggests, however, that the highest correlation of population expansion may not be with distance due north–south, but with distance along an axis tilted slightly northwest–southeast, and further examination showed that a tilt of ∼10° in fact gave the highest R2-value (0.71 compared with 0.69).

Figure 2.— Geographical distributions of Y-chromosomal haplogroups. (A) Populations sampled. (B–F) Haplogroup frequencies: circle area is proportional to sample size and sector area to haplogroup frequency. (B–E) Haplogroups are sorted into those showing predominantly northern (B), western (C), southern (D), and eastern (E) distributions. (F) The overall frequency of the most common haplogroup, O.
The distribution of Y-chromosomal haplogroups in East Asia has been extensively documented (e.g., Jin and Su 2000; Karafet et al. 2001; Deng et al. 2004), but these observations have raised questions about the relationship of northern and southern populations that remain unanswered. Su et al. (1999) typed 19 binary markers, 12 of which were chosen because they were already known to be variable in East Asia, and found higher diversity in the south than in the north and that the northern lineages were a subset of the southern ones, leading them to suggest that the northern populations were derived from the south by northward migrations. In contrast, Karafet et al. (2001) used a larger set of 52 binary markers ascertained mainly because of their variation in worldwide populations and discovered higher diversity (mean pairwise differences) in the north and that the northern lineages were not a subset of the southern ones. They concluded that a contribution to the northern populations from Central Asia was likely. The use of preascertained binary markers introduces a bias into estimates of diversity, but STRs are essentially free of this bias because they are variable in all populations. In our samples, STR diversity and ASD measurements were higher in the north than in the south (Table 2), a finding that is not easily reconciled with a largely or exclusively southern origin for the northern populations. It has been suggested that some populations, such as Hui, Uygurs, and Mongolians, have recent admixture with Central Asia and so reliance on them may give a false impression (Shi et al. 2005), but our findings are common to most populations from the north (Table 2).
Our most striking observation was the demographic contrast between north and south, which was explained largely by the variation in the start of population expansion (Tables 2 and 3; Figure 4). Despite the simplified demographic model and wide confidence intervals in the BATWING estimates (Table 2), the median values exhibit a simple and striking pattern: all of the northern estimates lie between 22 and 34 KYA, while all of the southern estimates are between 12 and 18 KYA. These suggest that the northern populations started to expand before the LGM (∼18–21 calendar KYA), while the southern populations started to expand after it. These time estimates are calibrated against historical events (Zhivotovsky et al. 2004) and so do not depend on the assumption of a particular male generation time, but nevertheless are uncertain, and so any interpretation based on them must be regarded with caution. Importantly, however, they are affected little by extensive admixture (Figure 5) and in such a case reflect the earlier expansion time. While extreme northern latitudes were inhospitable to early humans, Siberia has an extensive Upper Paleolithic archaeological record (Kuzmin and Orlova 1998) and a highly productive environment stretched across Asia. This showed an abundance of large animals and has been called the “Mammoth Steppe” (Guthrie 1990). Expansion times calculated in the same way for the Central Asian populations described by Zerjal et al. (2002), excluding those showing recent severe bottlenecks, lay between 24 (13–45) and 36 (16–74) KYA, like those of the northern populations from East Asia. We therefore propose that this cold but rich environment allowed the demographic expansion of populations who learned to exploit the profuse animal resources, and these people contributed in sufficient numbers to the ancestry of the northern populations we have tested to leave a signature in their paternal lineages. In contrast, this environment did not extend to the southern region, and the populations based there expanded only after the end of the LGM as the climate became warmer and more stable. The large-scale use of underground tubers is thought to have begun in the south as early as 15 KYA (Tong 2004), and it is notable that population expansion was subsequently more rapid there. The survival of this distinct demographic signature provides further evidence for the genetic differentiation between north and south and lack of extensive gene flow, leading to a genetic boundary seen initially in classical marker studies (Xiao et al. 2000).
GENETICS April 1, 2006 vol. 172 no. 4, 2431-2439; DOI: 10.1534/genetics.105.054270
academic.oup.com/genetics/article/172/4/2431/6061643