|
|
Post by Admin on May 28, 2023 17:09:50 GMT
There is a vast span of human existence of which little is known. Archeological ruins from the beginnings of civilization have been unearthed, and there have been occasional discoveries of a more prehistoric nature, but not much else. The impermanency of our physical existence is responsible for this void; our flesh and body fluids rapidly decompose after death. Unless preserved by extraordinary means, even skeletal remains eventually crumble and disappear. Early peoples did not practice ceremonial burial. Left to the elements, bodies soon completely decomposed: "Dust to dust" was not a mere poetic metaphor. It was a recorded observation of our transient natures.
Only in the last century have scientists and anthropologists begun using biological markers such as the blood groups in the search for humanity's imprint on our distant past. These studies have allowed a greater understanding of the movements and groupings of early peoples as they adapted to changing climates, mutating germs, and uncertain food supplies. Recent analyses, using sophisticated genetic measures, have produced the most accurate picture to date of human evolution.
The variations, strengths and weaknesses of each blood group can be seen as part of humanity’s continual process of acclimating to different environmental challenges. Most of these challenges have involved the digestive and immune systems. It is no surprise, then, that many of the distinctions between the blood groups involve basic functions of our digestive and immune systems.Evolution is usually considered in the context of millions of years, which is the time frame needed to explain the many differences between animals or other species. Yet humanity’s own life span provides ample time for the myriad number of small day-to-day refinements, representing the constant struggle between inherited traits and environmental challenges.
And, although evidence points to the fact that the individual genetic mutations that produced the ABO genes are quite ancient (1) this is trivial importance with regard to the actual demographics of the individual ABO blood groups in ancient populations. In genetics it is not the actual age of the gene that matters, it is its frequency or drift. This is computed by geneticists using a formula called the Hardy-Weinberg equation. Hardy-Weinberg posits that if the only evolutionary force acting on the population is random mating, the gene frequencies remain unchanged constant. In essence if you start off with a small number of a particular gene in a larger gene pool (such as the gene for blood group B in the gene pool for ABO blood type) and nothing other than random mating occurred, at the end of a period of time, you would still have a small number of B genes in the ABO gene pool.
So something other than random mating is responsible for the present day differences in frequency between the ABO blood groups; why for example, are there such large populations of blood group O (40-45%) and A (35-40%) versus much lower rates of groups B (4-11%) and AB (0-2%)?
First, it can be said that perhaps the mutation that produced the B gene was just not as common an occurrence as the mutation that produced the A gene. Yet, if they occurred at the same time, why would this be? Also, if the mutations are of such paramount importance, why is the distribution of the B gene so geographically limited to an area of high concentration stretching as a belt of territory from the Himalayas to the Urals?
The answer lies not in the ancient nature of the mutations that produced the A and B genes, but rather in the discreet interactions that occurred between early man and his environment that were under the influence of his ABO blood group. These included the areas and climates he chose to inhabit, each with their unique populations of microbes and foods that he chose to catch or cultivate.
As humans migrated and were forced to adapt their diets to local conditions, the new diets provoked changes in their digestive tracts and immune systems, necessary for them to first survive and later thrive in their new habitats. Different foods metabolized in a unique manner by each ABO blood group probably resulted in that blood group achieving a certain level of susceptibility (good or bad) to the endemic bacteria, viruses and parasites of the area. This probably more than any other factor was what has influenced the modern day distribution of our blood group. It is fascinating to note that virtually all the major infectious diseases that ran so rampant throughout our pre-antibiotic history have ABO blood group preferences of one group or another.(2)
This results from the fact that many microbes possess ABO "blood types" of their own. It is perhaps useful to understand that the ABO blood group antigens are not unique to humans, although humans are the only species with all four variants. They are relatively simple sugars which arte abundantly found in nature. A bacteria which for example possessed an antigen on its surface that mimicked the blood group A antigen would have a much easier time infecting a person who was group A, since that bacteria would more likely be considered "self" to the immune system of a blood group A person. Also microbes may adhere to the tissues of one ABO group in preference to another, by possessing specialized adhesion molecules for that particular blood group.(3)
The horrors of the Black Plague, which ran unchecked throughout Europe in the thirteenth and fourteenth centuries, is a perfect example. The Plague was a disease caused by bacterial infection and was almost certainly fatal to those who contracted it in the early years of its initial spread. By the fifteenth century, however, fatalities were rare, although many people continued to contract the infection. In just two generations, traits were developed in the survivors that protected them from fatal infections. Since these traits were necessary to survival, they were then passed on and retained as a form of genetic memory.
The Black Plague is especially interesting from a perspective of the ABO blood groups, since Yersinia is a bacteria with a preference for individuals of specific ABO group, in this case, group O. (4,5)
The effects of ABO blood group on survival against most forms of epidemic illness is so distinct that a modern day map of the ABO blood group distribution in Europe closely parallels the locations of major epidemics, with higher densities of blood group A and lower frequencies of blood group O in areas historically known to have had long histories of repeated pandemics.
On the other hand, in pre-urbanization days the survival advantage would have laid with blood group O as they are known to be more resistant to the flukes and worms that routinely parasitized these early humans, probably because they are the only blood group with antibodies against two other antigens, A and B.
These changes are reflected in the local success or failure of each of the blood groups, which appear to have each had a moment of pre-eminence at a critical juncture in our history. The ascent of humans to the top of the food chain (the early advantage of blood group O), the change from hunter-gathering to a highly concentrated, urban environment and agriculturally-based diet (the ascent of blood group A), and the mingling and migration of the races from the African homeland to Europe and Asia ( the opportunity for blood groups B and AB).
The Ancestral Foundation
From a purely scientific point of view, chemical analysis of the group O antigen reveals that from a structural perspective, it is the simplest blood group and it serves as the backbone for the synthesis of increasingly complex A, B and AB. These later blood groups evolved by adding other sugars onto the basic O sugar, much like a modern city might be built upon the foundations of an ancient one. Thus if the mutations that produced the A and B antigens are ancient, the gene for blood group O is infinitely older.
Another dimension testifying to the great antiquity of group O comes from the science of physical anthropology and suggests that a greater part of humanity's existence has been lived exclusively as group O.
New studies on mitochondrial DNA (mtDNA) support the theory that Homo sapiens emerged in Africa and only later infiltrated other regions. Unlike DNA, which is inherited from both parents and changes minutely with each generation, mtDNA is passed directly from mother to child. It is contained in eggs but not in sperm. Since only random mutations alter its sequence, it is a more accurate measure of the trajectory of human evolution. Extensive mtDNA studies demonstrate that humans evolved from a common ancestor. These studies also confirm the theory that the blood groups evolved as migratory mutations.
The extraordinarily high percentage of blood group O in "ancient" or otherwise isolated populations also testifies to its great age. (6) Even though the early migrations dispersed the gene for group O blood throughout the world, there are some extraordinary examples of "old" populations existing in our world today. Because of their geographic locations, these societies have remained isolated from interaction with other populations. If A, B and O had developed simultaneously, the isolated population groups would have had all of them. But these "old societies" are group O because genes for the later blood groups never had the opportunity to enter into their populations. They have remained unchanged.
The Basques are an ancient people whose origins are still a mystery. The Basque language, the only western European language not connected by Indo-European roots, appears to be related to several dialects found in small isolated populations in the valleys of the Caucasus Mountains. Although they look much like their French and Spanish neighbors, Basques possess the lowest frequency of blood group B---originally having no group B at all---and the highest frequencies of blood group O in Europe. Cattle, abundant on the European plains, and fresh water fish seem to have been the staples of their early existence, as evidenced by the extraordinary renderings of the famous cave paintings found in the Basque country.
More than fifty percent of the Basque population is Rh negative, as opposed to sixteen percent for the rest of Europe. Like the gene for group O, the genetic mechanism for the Rh negative blood type is simpler, hence undoubtedly older, than the gene for Rh positive.
Native Americans are another example of the "old peoples" existing in our world today. It has often been asserted that all full-blooded American Indians are group O, and recent studies on largely intermingled Amerindian populations show a very high (sixty-seven to eighty percent) predominance of O, indicating that their migration from Asia to Alaska was probably much earlier than previously believed.(7,8) Their high rate of blood group O suggests that the Amerindians and Eskimos are directly descended from Cro-Magnon ancestors, probably Mongolians, who migrated around 15,000 B.C. to the Americas. In contrast to the Basques, however, the Asian Amerindians must have mingled extensively with other Asian populations, picking up along the way the gene for Rh positive blood.
As with the Basques, few Native Americans are group B, so they must have migrated to the Americas late enough to pick up the Rh positive gene, but too early to pick up the gene for B. (9) This migration probably took place across the land bridge that at one time connected Siberia to Alaska. As the last Ice Age ebbed and the lands warmed and glaciers receded, the rising water levels eliminated the land bridge between Asia and American, bottling up the Native Americans and a high-O enclave and preventing for another 10,000 years any communication between the continents. Forensic studies support this theory: in Chile no B or AB have been noted either in pre-Columbian or Colonial mummies. (19)
Another theory for the extremely high incidence of blood group O in Native Americans is that O individuals seem relatively resistant to syphilis and smallpox, major killers of Native Americans that were introduced into the Old World by Columbus. (22)
Agricultural advances in the Americas were late in coming, because the new American homeland was abundantly populated with game and fish, which discouraged agriculture. Even corn, which was the staple grain, didn't appear to be domesticated until 4500 B.C., and common beans appear to be an even more recent addition, first being cultivated around 2200 B.C. So, like the Basques, meats and not grains were the primary staple of the Native American diet.
In England, Wales and Scotland there is a strong association between ABO blood group and geographical differences in the death rate (10) Studies of blood group distribution in the British Isles show a general increase of group O frequency from relatively low numbers in southern England to increasingly higher ones in northern England, Wales, Scotland, and Ireland. (11) This suggests that the Anglo-Saxons had relatively high A levels, and that O increased as the proportion of Celtic ancestry increased, although the origin of the high incidence of blood group O in the Irish may represent the remnants of Mesolithic peoples. (23) This is also the case with continental Europe, where the percentage of group O increases in northern Germans and Danes. It is also known that the Icelanders had high O frequencies, close to those frequencies found in the populations of Scotland and Ireland.
Among the Nomads of the Arabian Peninsula, and the Berbers of the Atlas Mountains, two old populations, the frequency of the blood group O gene is high. Africans, on average, have more O genes and less A genes than do Europeans. So it can be seen that the gene carried by people who are blood group O is ancient by evolutionary standards.
|
|
|
|
Post by Admin on Jul 3, 2023 17:52:35 GMT
An analysis of the blood types of one Denisovan and three Neanderthal individuals has uncovered new clues to the evolutionary history, health, and vulnerabilities of their populations. Silvana Condemi of the Centre National de la Research Scientifique (CNRS) and colleagues at Aix-Marseille University, France, present hese findings in the open-access journal PLOS ONE on July 28, 2021. 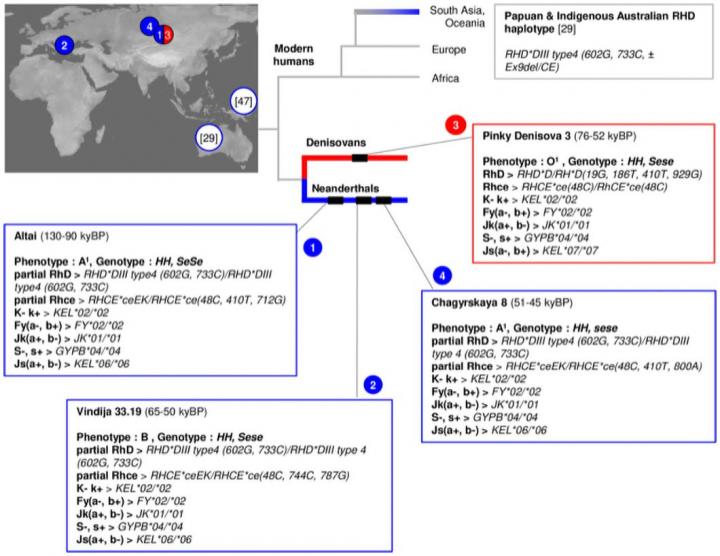 Neanderthals and Denisovans were ancient humans who lived across Eurasia, from Western Europe to Siberia, from about 300,000 to 40,000 years ago. Previous research efforts have produced full-genome DNA sequences for 15 of these ancient individuals, greatly enhancing understanding of their species. However, despite being encoded in DNA, these ancient individuals' blood types have received little attention. In the new study, Condemi and colleagues investigated the previously sequenced genomes of one Denisovan and three Neanderthal individuals (ranging from 100,000 to 40,000 years ago)in order to determine their blood types and analyze the implications. While 43 different systems exist for assigning blood types, the researchers focused on seven systems that are often used in medical settings for blood transfusions. This analysis of the four individuals' blood types revealed new information about their species. For instance, the ancient individuals had blood type alleles--different versions of the same gene--in combinations that are consistent with the idea that Neanderthals and Denisovans originated in Africa. In addition, a distinct genetic link between the Neanderthal blood types and the blood types of an Aboriginal Australian and an indigenous Papuan suggests the possibility of mating between Neanderthals and modern humans before modern humans migrated to Southeast Asia. The Neanderthal individuals also had blood type alleles associated with increased vulnerability to diseases affecting fetuses and newborns, as well as reduced variability of many alleles compared to modern humans. This pattern is in line with existing evidence that links low genetic diversity and low reproductive success with the eventual demise of Neanderthals. Overall, these findings highlight the relevance of blood types in understanding humans' evolutionary history. The authors add: "This work identifies the blood group systems in Neandertals and Denisovans in order to better understand their evolutionary history and to consolidate hypotheses concerning their dispersal in Eurasia and interbreeding with early Homo sapiens. The results of the Groups system analysis of Neandertals and Denisovans confirm their African origin as well as the weakness in their fertility and susceptibility to virus infection leading to a high infant mortality rate." ### Citation: Condemi S, Mazières S, Faux P, Costedoat C, Ruiz-Linares A, Bailly P, et al. (2021) Blood groups of Neandertals and Denisova decrypted. PLoS ONE 16(7): e0254175. doi.org/10.1371/journal.pone.0254175 |
|
|
|
Post by Admin on Jul 28, 2023 13:34:33 GMT
Blood groups of Neandertals and Denisova decrypted Abstract Blood group systems were the first phenotypic markers used in anthropology to decipher the origin of populations, their migratory movements, and their admixture. The recent emergence of new technologies based on the decoding of nucleic acids from an individual’s entire genome has relegated them to their primary application, blood transfusion. Thus, despite the finer mapping of the modern human genome in relation to Neanderthal and Denisova populations, little is known about red cell blood groups in these archaic populations. Here we analyze the available high-quality sequences of three Neanderthals and one Denisovan individuals for 7 blood group systems that are used today in transfusion (ABO including H/Se, Rh (Rhesus), Kell, Duffy, Kidd, MNS, Diego). We show that Neanderthal and Denisova were polymorphic for ABO and shared blood group alleles recurrent in modern Sub-Saharan populations. Furthermore, we found ABO-related alleles currently preventing from viral gut infection and Neanderthal RHD and RHCE alleles nowadays associated with a high risk of hemolytic disease of the fetus and newborn. Such a common blood group pattern across time and space is coherent with a Neanderthal population of low genetic diversity exposed to low reproductive success and with their inevitable demise. Lastly, we connect a Neanderthal RHD allele to two present-day Aboriginal Australian and Papuan, suggesting that a segment of archaic genome was introgressed in this gene in non-Eurasian populations. While contributing to both the origin and late evolutionary history of Neanderthal and Denisova, our results further illustrate that blood group systems are a relevant piece of the puzzle helping to decipher it. Introduction Over the last decade, technological progress has allowed generation of data from the entire genome of some fifteen extinct Neanderthal and Denisova hominins who lived 40,000 to 100,000 years ago from Western Europe to Siberia [1]. It reveals population structure, several demographic fluctuations and gene flows across hominin populations, worldwide dispersal of archaic genes by admixed Homo sapiens, and even the existence of a super-archaic ‘ghost’ population [2]. In addition, the availability of Neanderthal and Denisova DNA sequences provides a phylogenetic status and chronological depth that have significantly enhanced the understanding of gene variation in modern humans for phenotype, metabolic, and immune traits (reviewed in [3]). Red cell blood groups are powerful anthropological markers. Phenotype and genotype geographical distribution mirrors past human migrations and natural selection [4–7] and comparison with primates makes it possible to evoke their evolutionary and migration trajectory with accuracy [8, 9]. Red cell blood groups are also crucial in medicine to ensure transfusion safety, transplants, and foeto-maternal compatibility [10]. To date, the International Society of Blood Transfusion (retrieved from the ISBT website, has recorded more than 380 blood group specificities grouped into 40 systems. In transfusion, it is routine practice to scrutinize six blood groups: ABO, Rh, Kell, Duffy, Kidd and MNS (reviewed in [10]). Curiously, despite their significance and the amount of available genotypic data on modern Humans that is continuously accumulating [11], almost no attention has been paid to these major red cell blood polymorphisms in palaeogenetic studies [12]. In the present study, we analyze Neanderthal and Denisovan blood groups in order to trace back the current human diversity and to discuss health aspects and vulnerabilities of archaic populations. For that purpose, we investigated the high-quality nuclear genomes previously published from three Neanderthals one Denisovan. |
|
|
|
Post by Admin on Jul 29, 2023 10:33:22 GMT
Material and methods Selection criteria of the probands To ensure genotype calling rate, consistency across individuals and phylogenetic positioning in relation to anatomically modern humans, we did not consider contaminated, admixed, low-depth and archaic genomes with abundant uncalled positions in the loci understudy. We hence retained only high-quality genomes from one Denisovan (Denisova 3) and three Neanderthal individuals i.e., Altaï Neanderthal (Denisova 5), Vindija 33.19, and Chagyrskaya 8 [13–16] (S1 Table). These four probands are representative of the two archaic human-related species that spanned over 50,000 years of the Late Pleistocene and across approximately 5,000 km of Eurasia. Presentation of the blood groups under study We studied 7 blood group systems covering 11 genes: ABO including H system and Secretor status (ISBT 001 and 018, ABO, FUT1 and FUT2 genes), Rh (ISBT 004, RHD and RHCE genes), Kell and the covalently linked Kx protein (ISBT 006, KEL and XK genes), Duffy (ISBT 008, ACKR1 gene), Kidd (ISBT 009, SLC14A1 gene), MNS (ISBT 002, GYPB gene), and Diego and its Band 3-Memphis variant (ISBT 010, SLC4A1 gene) (S2 Table). Exploration procedure for blood group alleles For the probands and blood groups under study, we downloaded the already published [13–16] and curated *.vcf and *.bam(.bai) chromosome files available at the Genome Projects website of the Max Planck Department of Evolutionary Genetics (https://www.eva.mpg.de/genetics/genome-projects.html, S1 Table). For genotype calling filters, see the Supplementary Information of [13–16] and the readme files at cdna.eva.mpg.de/neandertal/. Briefly, the filters included a coverage filter stratified by GC content, minimum coverage of 10, Heng Li’s Mappability 35, Mapping Quality (MQ) of 25, no tandem repeats and no indels. Then, we briefly proceeded in a two-step screening of the blood group loci. First, we first gathered the genotypes at the key functional changes with depth, allele counts, quality and Phred scores probability using vcftools [17] (S2 Table, S1 File). Second, we browsed all exomes regions within the coding bounds (i.e. from the initiation ATG to the stop codons), in search for additional variation from hg19 (S3 Table, S1 File). While doing so, we paid specific attention to the following five points. Consideration of the reference sequence. We aligned with the reference blood group gene sequences used by the ISBT against the GRCh37 (hg19) with nucleotide labelling according to the sense (5’-3’) strand (S2 Table). We noticed that for six loci we studied (ABO, KEL, GYPB, RHCE, SCL4A1 and FUT1) the hg19 reference sequence opens by default onto the antisense strand (3’ → 5’) in the NCBI graphic window and the ancient genome browser, although their cDNA is conventionally the sense strand (5’ → 3’). This has two consequences: the chromosomal coordinates are decreasing as we progress throughout the coding strand (5’ → 3’) of these genes (from exon ’n’ to exon ’n+1’) and overall, nucleotides should be reversed-complemented. ABO genotype calls. We inferred the ABO alleles according to the functional approach developed by [18, 19] for pure and chimeric A-B transferase cDNAs. In conformity with this approach, we identified the ABO alleles by 4 letters corresponding to the 4 main amino acid changes in the catalytic site of the glycosyltransferase of pure A or B allele positions, preceded by the presence or not of the G in position c.261 (rs8176719) (i.e. G-AAAA meaning 4 SNPs of A allele generating A phenotype) and the deletion or not of the C at position c.1061 (rs56392308) to differentiate the A1 and A2 alleles. We achieved the ABO allele identification with the screening of all exons and collected the genotypes at 39 additional loci previously identified as responsible for various ABO alleles [20] (S2 Table; Fig A in S1 File). Special attention has been taken to FUT2 whose amino acid numbering in NBCI and hg19 should be rectified to get the correct amino-acid changes as mentioned by the ISBT. This is due to the fact that the initiation codon is the third ATG at the beginning of exon 2 (19: 49,206,247) [21]. RHD and RHCE genotype calls. For RHD and RCE, while browsing the exons in search for variation with hg19, we gathered the genotypes of the key changes of the RH*Ce, *CE, *ce, and *cE alleles. Any variation with hg19 was consolidated with The Human Rhesusbase.com [22], Erythrogene.com [11] and screenshots of the bam sequences (Figs B-D in S1 File). For any identification of a variant, we searched for it in all four archaic genomes. In addition, for any call at two key variants of our findings, namely c.733G>C (RHD) and c.712A>G (RHCE), we searched for the other polymorphisms that usually constitute the haplotypes made with them, respectively RHD*DBU, *DLX, *DV, *DVI, *DBS, *DBT, *DUC2 and *ceAR, *ceEK, *ceBI, *ce*SM, and *ceHAR. For this, we browsed both vcf and bam alignment by varying the MQ threshold (S2 Table; Fig D in S1 File). Identification of indels. Because indels could have been filtered out in the making the vcf files, all ABO, RHD and RHCE, notably the ABO c.261delG, c.1061delC, and RHCE 209bp insert have been double-checked from the specific indels vcf files (http://ftp.eva.mpg.de/neandertal/Vindija/VCF/indels/) and bam alignments using Integrative Genomics Viewer (IGV [23]) (S2 Table and Fig A in S1 File). Low-mapped variants. The screenshots of the bam alignments in simultaneously the four archaic individuals have highlighted a difference in depth and MQ between reference and alternate alleles. This is especially true for variants with very low frequency in modern humans reference panel such as rs17418085 (RHD), rs150073306 (RHD), rs1132763 (RHCE), and rs1132764 (RHCE) in the 3 Neandertals (alternate allele) in comparison with Denisova 3, homozygous for the reference alleles (Figs C and D in S1 File). Hence, these loci may suffer from reference bias, which is known to strongly reduce the depth and mapping of the reads with the alternate alleles [24, 25], and consequently, the genotype accuracy indexes at these loci. Hence, in cases where variants have been called in the released VCFs of some probands but filtered out in the others, we screenshotted the indexed alignments with hg19 using IGV [23] to manually call genotypes with allocated reads count and MQ cut-off (i.e. value above which the reads are not visualized) (S1 File). |
|
|
|
Post by Admin on Jul 31, 2023 3:17:46 GMT
Results Detailed information for the blood group systems, genotypes and phenotypes as well as for other polymorphisms identified in these archaic hominins is presented in Tables 1 and 2 and the principal information is shown in Figs 1 and 2.  Fig 1. ABO system. A, Nomenclature of the archaic ABO alleles [18]. B, Different observed genotypes and inferred phenotypes in Neandertal and Denisova. Red: polymorphisms characteristic of the ABO*A allele, blue: polymorphisms characteristic of the ABO*B allele, brown: deletion characteristic of the ABO*O01 allele.  Fig 2. Representation of the different RHD and RHCE genotypes in Neandertal and Denisova. Black font, unknown allees in modern humans; blue font, known alleles in modern humans; gray shadow, allele background found in modern humans.  Table 1. Blood group systems, genotypes and phenotypes of four Denisova and Neanderthal archaic genomes. Each occurrence of a SNP is scored by the "+" character. |
|
