Post by Admin on Jun 23, 2023 20:20:16 GMT
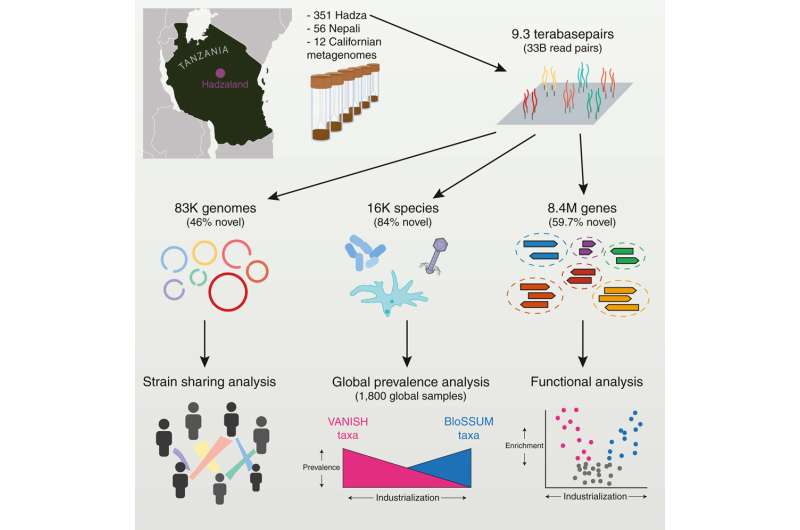
A genomic study led by researchers at Stanford University, California, has performed ultra-deep metagenomic sequencing on 351 fecal samples from the Hadza hunter-gatherers of Tanzania. In their paper, "Ultra-deep sequencing of Hadza hunter-gatherers recovers vanishing gut microbes," published in Cell, the team details their collection and identification of the most extensive set of gut microbiome sequencing data from a hunter-gatherer population.
Microbiome composition can vary significantly worldwide, depending on local diet and environmental contact. Microbiome studies are heavily biased toward Western industrialized populations with low microbiome diversity. The low diversity is thought to be caused by intake of highly processed foods, more frequent antibiotic use, cesarean section rates, sanitation levels and reduced physical contact with animals and soil.
This causal association is supported by looking at non-industrialized populations, including hunter-gatherers, with extremely high microbiome diversity. It can also be observed in the altered microbiome of immigrants from non-industrialized areas to industrialized Western cultures as a transition toward lower microbiome diversity.
Previous analysis of ancient human coprolites shows that ancient microbiomes more closely resemble the high diversity of the modern non-industrialized microbiome than the industrialized one.
These human-associated microbial lineages have been passed across hominid generations over deep evolutionary time, and knowing that gut microbiota plays essential roles in human health, suggests that the now missing aspects of the industrialized microbiome mean a loss of functions that these microbes provided.
The Hadza people in the current study live in the central Rift Valley of Tanzania in camps of approximately five to 30 people. Individuals move between camp locations about every four months. They primarily drink from water springs and streams and have a diet that includes foraged tubers, berries, honey, and hunted animals.
Metagenomic sequencing was conducted on stool samples collected from 167 Hadza individuals. Over half (59.7%) of the 8.4 million protein families found in Hadza gut microbes are absent from the Unified Human Gastrointestinal Protein database. Bacterial and viral species found in the microbiomes increase the global catalog of known microbiota by around 24%.
For a local comparison, the team compared the findings to adult Californians and found that while the average Hadza adult gut microbiome contains 730 species, there were only 277 species in the Californian gut microbiome.
The authors state, "An important challenge is to characterize the impact of these microbes on human physiology and determine in which contexts the absence or presence of species and functions are beneficial or detrimental to human health."
It will be fascinating to know the impacts of the lack of diversity on Western metabolisms or what diseases, if any, the Hadza populations are avoiding by having this robust diversity. It is unlikely that the missing microbiota from the industrialized populations was there without a reason, though it could simply be that the reasons no longer apply.
Thanks to the current effort, future researchers have many new mysteries to solve and an ultra-deep sequenced data set to guide their questioning.
More information: Matthew M. Carter et al, Ultra-deep sequencing of Hadza hunter-gatherers recovers vanishing gut microbes, Cell (2023). DOI: 10.1016/j.cell.2023.05.046
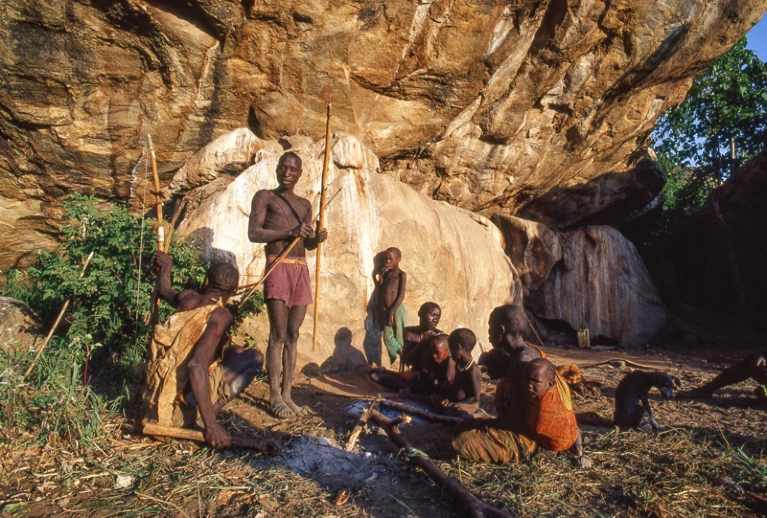

The human gut is teeming with trillions of microbes, but most studies of this vast community have focused on people living in urban regions. Now, a team of researchers has sequenced gut microbiomes from Hadza people — members of a hunter-gatherer society in northern Tanzania — and compared them with those from people in Nepal and California1. The study has found not only that the Hadza tend to have more gut microorganisms than people in the other groups, but that a Western lifestyle seems to diminish the diversity of gut populations.
The Hadza had an average of 730 species of gut microbe per person. The average Californian gut microbiome contained just 277 species, and the Nepali microbiomes fell in between. People with a farming-based lifestyle had an average of 436 microbe species, whereas those who live by foraging had an average of 317.
The team also found species in the Hadza microbiomes that were not present in the Californian samples, such as the corkscrew-shaped bacterium Treponema succinifaciens. Only some of the Nepali microbiomes contained this microbe, suggesting that the bacterium is dying out as societies become more industrialized.
Redressing the balance
Previous research has found that human gut microbiomes vary across regions and lifestyles, but there is a lack of data from non-industrialized populations, says study co-author Justin Sonnenburg, a microbiologist at Stanford University in California. “Part of the sequencing effort was to help fill that gap and provide more data for regions of the world that are under-represented,” he says.
Although it is well known that the microbiomes of people living non-industrial lifestyles are more diverse than those of people in industrialized societies, the findings show that the difference is more pronounced than previously thought, says study co-author Matthew Carter, also a microbiologist at Stanford.
“The data greatly expand our picture of the human microbiome,” says Andrew Moeller, an evolutionary biologist at Cornell University in Ithaca, New York. “I am sure there are untold stories that remain hidden in the sequences.”
The researchers sequenced microbiomes from fecal samples collected from 167 Hadza people — including infants and mothers — between 2013 and 2014. For comparison, the team also generated sequences from stool samples collected from four groups of people in Nepal in 2016, and samples from Californian participants in a 2021 study2 that explored how diet affects the microbiome.
Diversity dwindles
From these samples, Sonnenburg and his team sequenced more than 90,000 genomes from microbes found in the human gut, including bacteria, viruses that infect bacteria, and single-celled organisms from groups called archaea and eukaryotes. Some 44% of these microbial genomes had not yet been recorded in large catalogues such as the Unified Human Gastrointestinal Genome database. Among the genome sequences recovered from the Hadza samples, more than 1,000 were from bacterial or archaeal species that are new to science.
Furthermore, gut-microbe species commonly found in industrialized populations often contained genes associated with responding to oxidative damage. The team suspects chronic inflammation in the gut could trigger such damage, creating a selective pressure for those genes, says study co-author Matthew Olm, a microbiologist at Stanford. “If you have a state of chronic inflammation, it would make sense that your gut microbiome has to adapt,” he says. These genes were not detected in the Hadza microbiomes.
Samuel Forster, a microbiologist at the Hudson Institute of Medical Research in Melbourne, Australia, says that studying non-Western populations will help to build a more complete picture of the human gut microbiome and how it differs across lifestyles and regions. This could help researchers to track which species are disappearing in industrialized populations and how that affects human health, says Forster. “We have an opportunity to understand the full complement of microbes we carry,” he says. “It’s effectively avoiding an extinction event by understanding them now, before they’re lost.”
References
Carter, M. M. et al. Cell doi.org/10.1016/j.cell.2023.05.046 (2023).
www.cell.com/cell/fulltext/S0092-8674(23)00597-4
