Post by Admin on Feb 26, 2024 21:38:37 GMT
An uproar broke out on social media this week after Nature published a paper about a massive U.S. health research effort to capture the genetic diversity of people across the country. Critics said a key figure, which depicts patterns of relatedness among nearly 250,000 study volunteers whose genomes were sequenced, could mislead some readers into thinking the data support the idea that humans fall into distinct races.
The flap highlights the challenge of describing human ancestry data, some scientists say. The leader of the challenged All of Us study, funded by the National Institutes of Health, acknowledged in a statement that “many excellent points have been raised” about how researchers communicated their results. The paper’s authors have no immediate plans to revise the figure, but discussions with Nature are ongoing, an All of Us spokesperson said. “The feedback highlights how quickly this field of research is evolving, as well as its complexity,” geneticist and All of Us CEO Josh Denny said in the statement.
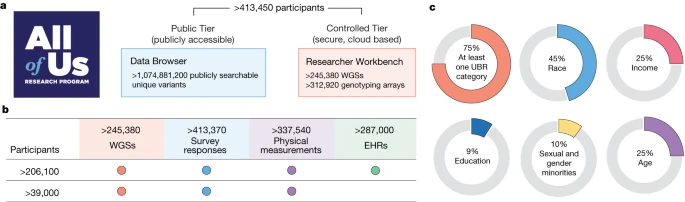
The study, which aims to eventually recruit 1 million volunteers across the United States, was designed to address concerns that existing genomic data sets are primarily composed of data from people of European descent. All of Us, however, has prioritized recruiting Black people, Latinos, and others with normally underrepresented backgrounds. The Nature paper, one of several from the study published this week, identified more than 1 billion DNA differences, or variants, among the nearly 250,000 genomes, noting that about one-quarter of those variants are novel and some could yield fresh insights into diseases.
Many researchers noted the value of the data set for expanding genomic research to include a greater diversity of people. However, several prominent geneticists quickly expressed concern that the way the All of Us team depicted the diversity in its data set was overly simplistic. The authors had used an algorithm called uniform manifold approximation and projection (UMAP) to summarize the variation and visually represent genetic relationships among participants who described themselves as white, Black, Asian, or a member of another racial group. This resulted in a graph consisting of several blobs of different colors (see the figure here).
The problem, critics said, is that UMAP creates blobs that look distinct while masking the inherent messiness in the data. “The fact that they are distinct is an artefact/feature of UMAP,” Ewan Birney, director of the European Bioinformatics Institute, wrote in a long thread on the social media platform X (formerly Twitter) describing how UMAP takes complex genomic data and summarizes them in 2D. “Almost certainly, some of the people in the other big blobs are some sort of cousin to the main blob.”
Birney acknowledged there’s no “easy way to represent this data in 2D” but also expressed concern that “it can easily be read as ‘race is pretty real, and associated with genetics’ which is … *not* a good interpretation.” Stanford University geneticist Jonathan Pritchard expressed a similar concern. “I’m not a UMAP hater in all settings, but I think it’s misleading and potentially harmful for this specific problem,” he wrote on X, adding that it could be “misinterpreted by the public.”
The paper’s corresponding author, geneticist Alexander Bick of Vanderbilt University Medical Center, acknowledges that the figure could have been labeled more clearly. But he points out that the three other major human genome papers published in the past few years, from the UK Biobank, a database called gnomAD, and the Mexican Biobank, also use the UMAP algorithm, which “is frankly why we selected it.” Trying to depict complex genomics data in 2D is “really challenging,” he says.
Bick also counters arguments by some critics that the All of Us paper authors disregarded a recent National Academies of Sciences, Engineering, and Medicine report on the appropriate use of population labels in genetics studies. He notes that the report came out after the Nature paper was first submitted, but that he and his co-authors incorporated its advice on several matters, such as not including race and ethnicity in the same figure.
Outspoken geneticist and former eLife Editor-in-Chief Michael Eisen called on X for a retraction of the Nature paper, warning that it “features a scientifically invalid representation of genetic diversity and race that is going to feature in racist literature for decades.”
When asked about the concerns, a Nature spokesperson said: “We are aware of the discussions that are taking place and are in contact with the authors.”
Geneticist Daniel MacArthur of the Garvan Institute of Medical Research tried to find a middle ground in the discussion. “All Of Us is one of the most thoughtfully inclusive programs in the history of human genetics, and will have enormous impact on reducing inequity in genomic medicine,” he posted on X. But, he added, the lesson of the UMAP flap is to “be careful with ancestry labels; they matter.”
www.nature.com/articles/s41586-023-06957-x
The flap highlights the challenge of describing human ancestry data, some scientists say. The leader of the challenged All of Us study, funded by the National Institutes of Health, acknowledged in a statement that “many excellent points have been raised” about how researchers communicated their results. The paper’s authors have no immediate plans to revise the figure, but discussions with Nature are ongoing, an All of Us spokesperson said. “The feedback highlights how quickly this field of research is evolving, as well as its complexity,” geneticist and All of Us CEO Josh Denny said in the statement.
The study, which aims to eventually recruit 1 million volunteers across the United States, was designed to address concerns that existing genomic data sets are primarily composed of data from people of European descent. All of Us, however, has prioritized recruiting Black people, Latinos, and others with normally underrepresented backgrounds. The Nature paper, one of several from the study published this week, identified more than 1 billion DNA differences, or variants, among the nearly 250,000 genomes, noting that about one-quarter of those variants are novel and some could yield fresh insights into diseases.
Many researchers noted the value of the data set for expanding genomic research to include a greater diversity of people. However, several prominent geneticists quickly expressed concern that the way the All of Us team depicted the diversity in its data set was overly simplistic. The authors had used an algorithm called uniform manifold approximation and projection (UMAP) to summarize the variation and visually represent genetic relationships among participants who described themselves as white, Black, Asian, or a member of another racial group. This resulted in a graph consisting of several blobs of different colors (see the figure here).
The problem, critics said, is that UMAP creates blobs that look distinct while masking the inherent messiness in the data. “The fact that they are distinct is an artefact/feature of UMAP,” Ewan Birney, director of the European Bioinformatics Institute, wrote in a long thread on the social media platform X (formerly Twitter) describing how UMAP takes complex genomic data and summarizes them in 2D. “Almost certainly, some of the people in the other big blobs are some sort of cousin to the main blob.”
Birney acknowledged there’s no “easy way to represent this data in 2D” but also expressed concern that “it can easily be read as ‘race is pretty real, and associated with genetics’ which is … *not* a good interpretation.” Stanford University geneticist Jonathan Pritchard expressed a similar concern. “I’m not a UMAP hater in all settings, but I think it’s misleading and potentially harmful for this specific problem,” he wrote on X, adding that it could be “misinterpreted by the public.”
The paper’s corresponding author, geneticist Alexander Bick of Vanderbilt University Medical Center, acknowledges that the figure could have been labeled more clearly. But he points out that the three other major human genome papers published in the past few years, from the UK Biobank, a database called gnomAD, and the Mexican Biobank, also use the UMAP algorithm, which “is frankly why we selected it.” Trying to depict complex genomics data in 2D is “really challenging,” he says.
Bick also counters arguments by some critics that the All of Us paper authors disregarded a recent National Academies of Sciences, Engineering, and Medicine report on the appropriate use of population labels in genetics studies. He notes that the report came out after the Nature paper was first submitted, but that he and his co-authors incorporated its advice on several matters, such as not including race and ethnicity in the same figure.
Outspoken geneticist and former eLife Editor-in-Chief Michael Eisen called on X for a retraction of the Nature paper, warning that it “features a scientifically invalid representation of genetic diversity and race that is going to feature in racist literature for decades.”
When asked about the concerns, a Nature spokesperson said: “We are aware of the discussions that are taking place and are in contact with the authors.”
Geneticist Daniel MacArthur of the Garvan Institute of Medical Research tried to find a middle ground in the discussion. “All Of Us is one of the most thoughtfully inclusive programs in the history of human genetics, and will have enormous impact on reducing inequity in genomic medicine,” he posted on X. But, he added, the lesson of the UMAP flap is to “be careful with ancestry labels; they matter.”
www.nature.com/articles/s41586-023-06957-x