|
|
Post by Admin on Feb 10, 2022 0:37:27 GMT
Figure 4 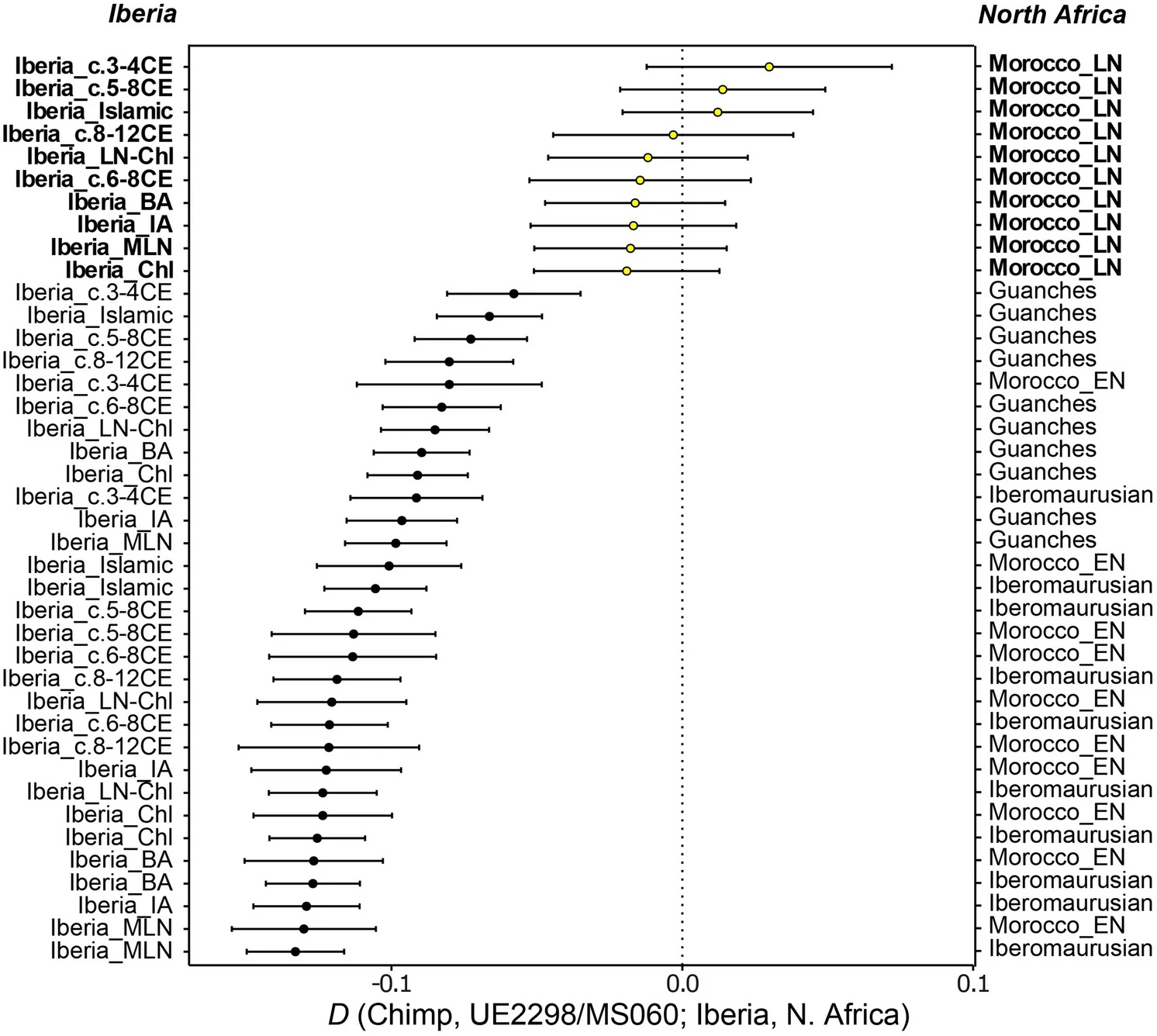 We tested different qpAdm 1-way scenarios using different proximal Iberian sources as left populations. Models using populations from Andalusia (Iberia_c.5-8CE and Iberia_c.3-4CE, which already displayed North African-related ancestry6) are accepted (p-values: 0.092 and 0.343, respectively), whereas models using populations from Catalonia, in the northeast of the Peninsula, are rejected (p-value < 0.05) (Supplementary Table S8). However, considering the genetic heterogeneity in different regions of Iberia through time, and given the complex history of population interactions in Iberia during the first millennium CE16,18, it is unlikely that UE2298/MS060 descends directly from Andalusian Visigothic populations and therefore we also explored 2-way admixture scenarios. Notably, 1-way qpAdm analysis was consistent with UE2298/MS060 descending from Islamic_Andalusia (p-value = 0.327) but not from Islamic_Valencia (p-value = 0.0005), in line with the position of UE2298/MS060 in the PCA (Fig. 3b) and highlighting regional genetic differences during this period. Alternatively, UE2298/MS060 could be modelled using 2-way combinations of distal and proximal Iberian populations (showing varied proportions of North-African related ancestry6) and either the Guanches or Morocco_LN (Table 1; Supplementary Table S9). D-statistics comparing these two North African populations indicate that UE2298/MS060 is closer to Morocco_LN (|Z|> 3) (Supplementary Table S7) than to the Guanches. Table 1 Accepted 2-way qpAdm admixture models with standard errors (SE) and p-values. Models accepted using both datasets (“mapDamage --rescale” and “soft-clipping”) are shown in italics.
Target Left populations Admixture proportions SE p-value
Pop. 1 Pop. 2 Pop. 1 Pop. 2
UE2298/MS060 Guanches Iberia_Islamic 0.102 0.898 0.066 0.148617
UE2298/MS060 Guanches Iberia_c.8-12CE 0.172 0.828 0.058 0.220096
UE2298/MS060 Guanches Iberia_c.5-8CE 0.122 0.878 0.064 0.168943
UE2298/MS060 Guanches Iberia_c.3-4CE 0.091 0.909 0.07 0.391308
UE2298/MS060 Guanches Iberia_IA 0.349 0.651 0.048 0.078756
UE2298/MS060 Morocco_LN Iberia_Islamic 0.235 0.765 0.149 0.146791
UE2298/MS060 Morocco_LN Iberia_c.8-12CE 0.308 0.692 0.13 0.099908
UE2298/MS060 Morocco_LN Iberia_c.6-8CE 0.593 0.407 0.059 0.053639
UE2298/MS060 Morocco_LN Iberia_c.5-8CE 0.18 0.82 0.17 0.091952
UE2298/MS060 Morocco_LN Iberia_c.3-4CE 0.094 0.906 0.23 0.287481
|
|
|
|
Post by Admin on Feb 11, 2022 0:18:31 GMT
Mobility in Islamic Segorbe In order to assess whether or not UE2298/MS060 was likely to have spent their childhood in the local region, we performed stable oxygen analysis on eight individuals from Plaza del Almudín. Tooth enamel carbonate data is presented in Supplementary Table S10 and plotted in Fig. 5a. The δ18OVSMOW values for the Segorbe population (excluding outlier MS075) range from 26.2 to 27.6‰ (range = 1.4‰, n = 7), with a mean of 26.8 ± 0.5‰ (1σ). The converted δ18Odw values (mean -6.0‰, excluding MS075) fit with the meteoric water values for eastern Iberian coast. The δ18OVSMOW values from both teeth sampled from UE2298/MS060 are consistent with the rest of the population and the small difference in values between the different molars (M1/M2 and M3) provide no indication of movement between early childhood and adolescence. Overall, there is no evidence that UE2298/MS060 was an immigrant in East Spain, on the basis of his oxygen values. Figure 5 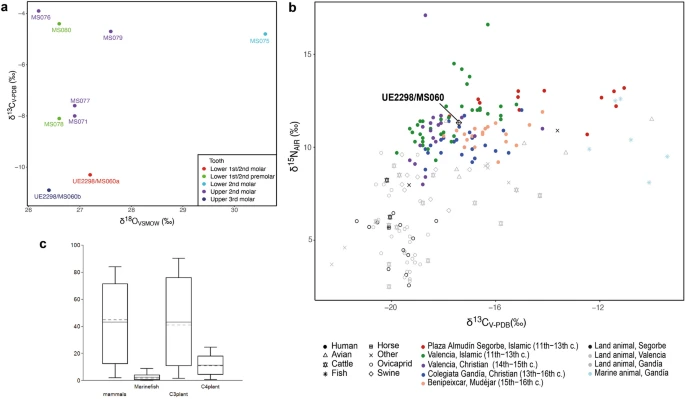 By contrast, one other individual reported here (MS075) seems to be an outlier (δ18OVSMOW = 30.6; > 1.5 times the interquartile range above quartile 3)47, and possibly a migrant from a warmer climate, with a δ18Odw value similar to Africa or the Near East48. Detailed results and discussion of oxygen analysis can be found in Supplementary Note 2. Diet patterns in Islamic Segorbe The values for δ15N and δ13C dietary isotopes in the Islamic necropolis of Plaza del Almudín range between 10.7 to 13.2‰ and from –17.8 to –11‰, respectively, for the 13 individuals studied (Fig. 5b; Supplementary Table S11). UE2298/MS060 has a δ15N value of 11.3‰ and a δ13C value of –17.4‰, showing lower δ15N and a more negative δ13C than the majority of the humans sampled from this assemblage. Application of a Bayesian mixing model (BMM), FRUITS (Food Reconstruction Using Isotopic Transferred Signals)49, supports the observation that C4 plants likely played a substantial part in the diet of some individuals and that marine fish consumption was variable (Supplementary Fig. S11). UE2298/MS060 (Fig. 5c) seems to have consumed limited amounts of C4-plants (mean: 11.4 ± 6.5% or 4.8–17.9% of the diet) and marine protein (mean: 2.4 ± 2.4% or 0–4.8% of the diet) compared to the rest of the population analysed. On the other hand, he seems to have the highest levels of mammal and C3-plant consumption amongst the analysed individuals (Supplementary Fig. S11). Individual MS075, identified as a possible migrant due to their oxygen value, displays the lowest probability (close to zero) of marine fish consumption amongst the individuals studied here (Supplementary Fig. S11), and shows signals of a mixed C3/C4 diet, which is also a possibility for Africa50. Detailed results and discussion of diet patterns inferred from individuals from the site of Plaza del Almudín can be found in Supplementary Note 2. |
|
|
|
Post by Admin on Feb 11, 2022 3:35:28 GMT
Discussion
We analysed individual UE2298/MS060 excavated from the Islamic necropolis of Plaza del Almudín, in Segorbe, dating to the eleventh century CE. The archaeologists responsible for the excavation in 1999 considered this individual unusual due to his considerable height compared with other individuals found at the same site (despite periods of disease and/or malnutrition in childhood)27, and dubbed him the “Segorbe Giant”. The subsequent anthropological analysis suggested some African morphological features and a link was postulated to the Berber-speaking populations that settled in the region in medieval times26,27.
Analysis of the uniparental markers from UE2298/MS060 fits well with this assumption, pointing to an origin in the Maghreb, most likely from a Berber group. MtDNA lineage U6a is not only connected to modern Amazigh populations30, but has also been found in Moroccan remains associated with Iberomaurusian culture, and in the Moroccan Early Neolithic site of Ifri n’Amr or Moussa2,32 (Fig. 1b). He also carries the Y-chromosome E1b1b1b1 (E–M310) lineage. E1b1b is extremely common amongst extant North Africans and has been found in ancient North African and Levantine remains2,32,33,37 (Supplementary Fig. S7). Due to low coverage, we could only assign him to a basal position within E1b1b1b1, but it is possible that he may belong to a more derived subclade. One possibility would be E1b1b1b1a (E–M81), which is the most common haplogroup amongst modern Berber males today42,53, and has been linked to Islamic remains in southern France38. Another would be its descendant E1b1b1b1a1-M183 lineage, identified in three Guanche males, in two Islamic individuals from Granada, and in an earlier sixth century CE male from the Visigoth phase of Pla de l'Horta, in Catalonia6,33.
Although he carries both uniparental markers of North African origin, autosomal evidence paints a more complex picture. The individual is positioned in the PCA mid-way between modern/ancient Iberian populations, and Late Neolithic Moroccan, Guanches and modern North African individuals (Fig. 3a), and formal tests of admixture point to high proportions of Iberian-like ancestry (Fig. 4; Supplementary Table S7).
Considering the archaeological and historical records for this period in the region of Valencia, we envisage three possible scenarios to explain the observed ancestry in UE2298/MS060. One would be to assume that this individual is a direct migrant from North Africa (whose unique genetic composition has not yet been examined using aDNA), or derives from a population that moved into Iberia but retained its genetic identity. A second scenario is that he descends from pre-Islamic Iberian genetic diversity. Finally, the third scenario is that he is the result of admixture between Iberian and North African sources.
The first scenario would imply that pre-Islamic populations in North Africa would be genetically similar to UE2298/MS060 (or possibly to other contemporary individuals found in Spain6). The nearest temporal proxy available are the Guanches (from the seventh–eleventh centuries CE), who originated in the Maghreb but have been isolated in the Canary Islands since at least the early Iron Age. D-statistics, however, suggest that UE2298/MS060 is genetically closer to Morocco_LN than to the Guanches (Supplementary Table S7). In any case, qpAdm rejects the hypothesis that UE2298/MS060 directly descends from a population resembling either the Guanches or Morocco_LN (Supplementary Table S8). Additionally, the oxygen data for UE2298/MS060 (Supplementary Note 2) is consistent with someone who grew up in the region, and points towards low mobility between early childhood and adolescence. (In contrast, another individual from the same necropolis (MS075) does look non-local (Supplementary Note 2), possibly a migrant from a warmer climate outside the Mediterranean, with oxygen values similar to those of Africa or the Near East48). Nevertheless, one should note that aDNA sampling in North Africa is sparse and limited to a few individuals from very specific sites and periods, and we cannot rule out that a population with a similar genetic composition to that of UE2298/MS060 existed in the region around this period.
Although North African-related ancestry in present-day Spain is present at low values (typically ~ 3–8%), with a slight southwest-to-northeast decline19,20, increased African-related ancestry has been present in south Spain since the third century CE6. This North African influence is captured in our qpAdm analysis, with 1-way models using pre-Islamic Andalusian populations being accepted (Supplementary Table S8). However, it is unlikely that UE2298/MS060 descends directly from Andalusian Visigothic populations and ultimately these models, despite being statistically plausible, do not fully explain the ancestry of our individual. We note that there are no data available from or around the region of Valencia between the end of the Iron Age and the Islamic period, and post-Iron-Age genetic variation in Spain was most likely very heterogeneous across locations and centuries6. This heterogeneity is confirmed by our results showing that UE2298/MS060 forms a clade with Islamic_Andalusia, but not with Islamic_Valencia (Supplementary Table S8).
The third scenario would be that the genetic variation seen in UE2298/MS060 was a result of admixture between Amazigh people who migrated from North Africa to Iberia, and the local population inhabiting the Peninsula, at some point during either the Islamic conquest, the Caliphate period, or the Berber empires. This would explain UE2298/MS060's intermediate position in the PCA and ternary plot (supervised ADMIXTURE) (Fig. 3). D-statistics support this scenario, with tests comparing Morocco Late Neolithic and Iberian populations from different periods not showing him to be significantly closer to one or the other (Fig. 4; Supplementary Table S7). We show that UE2298/MS060 can be modelled as admixture between Iberian and North African sources (either the Guanches from the Canary Islands or Late Neolithic Moroccans) (Table 1). The fact that he still carried both uniparental markers of North African origin suggests that the admixture may have happened only a few generations before his time, coinciding with the zenith of Berber power, rather than earlier during the conquest, in agreement with admixture dates inferred from modern Iberian genomes from Aragon and Catalonia20. However, we cannot rule out assortative mating, allowing these uniparental markers to be retained for longer, or the possibility that these lineages were common in some Iberian populations before the Islamic period. The date of the burial (eleventh century CE)27 fits the historical narrative of Berber settlement in the region of Sharq al-Andalus18. Considering the genetic evidence, together with the stable isotope results and the historical accounts of intermarriage between local individuals and the North African newcomers, and in agreement with recent aDNA evidence from Iberia6, this third scenario seems the most plausible to explain the ancestry patterns seen in his genome.
Nevertheless, the original source populations are difficult to pinpoint. Due to lack of sampling in North Africa for this specific period and preceding centuries, the nearest proxies available for the North African source are the Guanches33 and the Late Neolithic Moroccan population from Kelif el Boroud site2. There is high differentiation between present-day North African populations and ancient North African individuals available to date (seen in PC3; Supplementary Fig. S9), which indicates that important population dynamics occurring after the Late Neolithic and/or Iron Age shaped extant genetic structure in the region. Modern North African populations show a signal of increased Levantine-related ancestry around the seventh century CE, as a result of movements from the Near East during the Islamic expansion into North Africa17; the impact of these movements was also seen in the Levant, as shown by the study of seventh–eighth century Islamic individuals in Syria54. Therefore, the North-African source of UE2298/MS060 might have already displayed this increased Near Eastern-related ancestry. Similarly, the population of Valencia in the immediately preceding centuries has yet to be studied.
A study in modern South Americans detected North African ancestry introduced at the early stages of European colonization55. The presence of individuals in medieval Spain with a genetic background similar to that of UE2298/MS060 would explain the source of this ancestry in America, suggesting that admixture with North Africans had a wider impact on medieval Spanish genetic variation, before virtually disappearing in the following centuries.
We found no U6 in our present-day whole-mtDNA dataset from the region of Valencia (n = 54), or in a larger previously published HVS-I database (n = 123)56. This absence might be an echo of the brutality of the decree of expulsion of Moriscos (Muslims forcibly converted to Christianity), which may have effectively erased the population carrying North African-related ancestry that lived in the region in the preceding centuries. They were replaced by settlers from regions further north with little North African-related ancestry20. This is in sharp contrast with regions of the Crown of Castilla, where historical sources claim there was better integration of the Morisco identity into the general population, and where no mass deportations were recorded: the frequency of U6, M1 and L lineages are higher in these regions today (present-day central and south Spain) (Fig. 1b; Supplementary Fig. S5). This pattern is also visible at the genome-wide level20.
This study emphasises the importance of immigration during the Islamic period. In contrast to Andalusia, the region of Valencia is not geographically close to the Maghreb, and was under Islamic rule for a shorter time, but nonetheless developed strong links with the Arab–Berber world during the Islamic period57. A contemporary individual, MS075, is evidence of continued movement during Berber rule (Supplementary Note 2).
UE2298/MS060 is a single, low-coverage sample and although the results cannot be extrapolated to the population as a whole, recently published results6 show a similar trend of admixture in Islamic Spain. The heterogeneity of genomic patterns that is now being uncovered by aDNA studies emphasises the need for much more detailed, high-resolution fine-scale studies. More individuals and a wider diversity of sites across the Peninsula should be studied to explore the population dynamics during the Islamic period in more detail and assess potential fine differences between geographical regions and periods, and between urban and rural societies.
|
|
|
|
Post by Admin on Feb 23, 2022 2:25:21 GMT
Genomic Analyses of Pre-European Conquest Human Remains from the Canary Islands Reveal Close Affinity to Modern North Africans
Summary
The origins and genetic affinity of the aboriginal inhabitants of the Canary Islands, commonly known as Guanches, are poorly understood. Though radiocarbon dates on archaeological remains such as charcoal, seeds, and domestic animal bones suggest that people have inhabited the islands since the 5th century BCE [1, 2, 3], it remains unclear how many times, and by whom, the islands were first settled [4, 5]. Previously published ancient DNA analyses of uniparental genetic markers have shown that the Guanches carried common North African Y chromosome markers (E-M81, E-M78, and J-M267) and mitochondrial lineages such as U6b, in addition to common Eurasian haplogroups [6, 7, 8]. These results are in agreement with some linguistic, archaeological, and anthropological data indicating an origin from a North African Berber-like population [1, 4, 9]. However, to date there are no published Guanche autosomal genomes to help elucidate and directly test this hypothesis. To resolve this, we generated the first genome-wide sequence data and mitochondrial genomes from eleven archaeological Guanche individuals originating from Gran Canaria and Tenerife. Five of the individuals (directly radiocarbon dated to a time transect spanning the 7th–11th centuries CE) yielded sufficient autosomal genome coverage (0.21× to 3.93×) for population genomic analysis. Our results show that the Guanches were genetically similar over time and that they display the greatest genetic affinity to extant Northwest Africans, strongly supporting the hypothesis of a Berber-like origin. We also estimate that the Guanches have contributed 16%–31% autosomal ancestry to modern Canary Islanders, here represented by two individuals from Gran Canaria.
Results and Discussion
We processed twelve samples from twelve different individuals, of which eleven yielded genome-wide sequence data (Table 1). We sampled human teeth only from intact skulls. The ancient individuals, collected from caves on Tenerife and Gran Canaria (the two largest islands of the Canary archipelago) (Figure 1), were donated to the skull collection housed in the AMEU (Anatomical Museum Edinburgh University) in the 19th century (see Experimental Model and Subject Details).
Table 1. Guanche Samples – Summary Statistics
Sample Origin Molecular Sex Genome Coverage Mitochondrial Genome Coverage SNPs in HO Dataset Mitochondrial Haplotypes Y Chromosome Haplotypes C14 Radiocarbon Date Before Present C14 Radiocarbon Date, Calibrated Common Era Mitochondrial Contamination Estimate/Confidence interval
gun002 Tenerife XY 0.21 294.7 74,618 H1cf E1b1b1b1a1 E-M183 951 ± 26 1089.4 ± 65.5 3.63%/2.42%–4.85%
gun005 Gran Canaria XX 0.47 341.1 140,873 H2a NA 1082 ± 26 956 ± 61 2.41%/1.56%–3.27%
gun008 Gran Canaria XX 0.30 690.9 101,216 L3b1a NA 1116 ± 26 935.5 ± 56.5 1.65%/1.35%–1.95%
gun011 Tenerife XY 3.93 931.6 370,465 T2c1d2 E1b1b1b1a1 E-M183 1216 ± 27 791.5 ± 96.5 0.53%/0.38%–0.69%
gun012 Tenerife XY 0.54 214.3 157,104 U6b1a E1b1b1b1a1 E-M183 1421 ± 28 621 ± 39 5.97%/4.22%–7.73%
gun001 Tenerife XY 0.016 9.6 NA U6b1a NA NA NA NA
gun004 Tenerife NA 0.004 49.98 NA J1c3 NA NA NA 5.76%/3.57%–7.95%
gun006 Gran Canaria XY 0.027 12.21 NA L3b1a NA NA NA 6.60%/1.87%–11.33%
gun007 Gran Canaria XY 0.003 3.47 NA L3b1a NA NA NA NA
gun013 Tenerife NA 0.005 14.2 NA U6b1a NA NA NA 13.79%/1.24%–26.34%
gun014 Tenerife XY 0.008 4.2 NA U6b NA NA NA NA
NA, not applicable. C14 lab numbers in order of appearance in the table: Ua-55344, Ua-55345, Ua-55346, Ua-55347, Ua-55348. See also Table S1 and Figure S1B.
|
|
|
|
Post by Admin on Feb 23, 2022 20:51:25 GMT
 The average read length (DNA fragmentation) and patterns of cytosine deamination, clustering at a significantly higher frequency at fragment termini, are consistent with expectations for ancient DNA (Figure S1A) [10]. Mitochondrial contamination estimates [11] for the five individuals included in further autosomal genomic analysis range from point estimates of 0.53% to 5.97%, showing that contamination is largely negligible (Table 1). The five individuals with the highest autosomal genome coverage (0.21× to 3.93×) were directly radiocarbon dated at the Svedberg Laboratory (Uppsala University) (Figure S1B) and span approximately 400 calendar years from the 7th to the 11th centuries CE (Table 1). All individuals predate the European colonization of the Canary Islands (15th century) and one predates the Muslim conquest of the Maghreb (7th–8th centuries), events that have had significant impact on the gene pool of the Canary Islands and North Africa, respectively (Figure S1B) [6, 12]. Hence, our sample is a good representative of the pre-conquest aboriginals. Analysis of Uniparental Genetic Markers The mitochondrial genome coverage for the eleven Guanche individuals ranges from 3.4× to 931×. The number of SNPs that support each haplotype varies between 38 and 58 and are reported as deviations from the Reconstructed Sapiens Reference Sequence (RSRS) [13] (Table S1). All positions supporting the predicted haplogroup calls and mutations missing for the absolute haplogroup assignment are listed in Table S1. We found six different mitochondrial haplogroups in eleven individuals (Tables 1 and S1). The Guanches analyzed here carried mitochondrial lineages such as J1c3, H2a, U6b, L3b1a, and T2c1d2 that are common across West Eurasia and/or North Africa [14] (Tables 1 and S1) and are consistent with previous studies on ancient Guanche mitochondrial DNA [6, 7]. Two individuals from Tenerife (gun001 and gun012) and one from Gran Canaria (gun013) carried the U6b1a haplotype, which is hypothesized to be endemic to and a founder lineage of the Canary Islands (Tables 1 and S1) [15]. We also found the H1cf haplotype in one individual from Tenerife (gun002), which is defined by a mutation at position 16260T (Table S1). H1-16260 is also considered a founder lineage of the Canary Islands, for two reasons: (1) it is present in all modern Canary populations [15] and rare outside the Canary Islands (found previously in only a single Algerian individual) [7, 16]; and (2) it was found in pre-Hispanic populations from Tenerife [6], La Palma [7], El Hierro [16], and La Gomera [17]. The three males from whom haplogroup-defining Y chromosome SNPs were retrieved carried the E1b1b1b1a1 (E-M183) haplotype (a major sub-clade of the haplogroup E1b1b1b1, defined by the derived M81 marker), which is again consistent with previous analyses of ancient Guanches [8]. This haplogroup is ubiquitous across modern North African populations and particularly common in Berber-speaking populations of North Africa [8, 18]. The derived mutations supporting the Y chromosome haplogroups are listed in Data S1, sheet 1. Population Genomic Analysis of Autosomal DNA A principal component analysis (PCA) of the five samples with the highest autosomal genome coverage, performed using genome-wide autosomal SNPs overlapping with Human Origins (HO) data [19, 20], reveals close affinity to modern Northwest African populations such as Tunisians and Algerians, but with a tendency (especially for individuals from Gran Canaria) to occupy a space outside modern Northwest African variation, closer to Europeans (Figures 2 and S2). However, outgroup f3 statistics [19] suggest that the Guanches share more genetic drift with non-African test populations than with African test populations, including Northwest African populations of Berber origin (Data S1, sheet 2). This observation is inconsistent with the PCA and the uniparental genetic marker data, indicating that the outgroup f3 statistic may be misleading, possibly due to the complex history of recent sub-Saharan admixture events in North African populations [12, 21] and the sensitivity of the f3 estimator to such patterns. This issue seems to extend to other statistics based on allele frequency correlations such as the D statistic [19] since D(Outgroup, Guanches; North African, Sardinian/Anatolian farmer) consistently produces highly significant positive values of D (Z > 4), which would imply a closer relationship between Guanches and Sardinians and Anatolian farmers than between Guanches and North African populations (Data S1, sheet 3).  Figure 2. PCA of the Five Guanche Individuals Used in the Autosomal Analysis Principal component analyses performed on the Guanches and a number of populations from Europe, the Middle East, and North Africa selected from the Human Origins dataset [19, 20]. Green symbols represent individuals from Tenerife; red symbols represent individuals from Gran Canaria. See also Figure S2. In order to resolve this, we used different statistics to measure genetic similarity between Guanches and modern populations, as well as between different modern North African populations and other populations (Table S2 and Quantification and Statistical Analysis). Both outgroup f3 statistics and average pairwise differences identify Sardinians as the population sharing the most drift with both modern North Africans and Guanches (Table S2). This is in stark contrast to our expectation that North Africans would cluster with other North Africans. However, it replicates recent findings [21] showing that both ancient and modern Egyptians share more genetic drift with European populations than they do with other North African groups, while still showing evidence of strong genetic affinity to one another in other types of analyses. This behavior might be due to varying degrees of admixture from highly divergent sources (e.g., sub-Saharan populations) into different populations, which can affect the interpretation of f3 statistics [22]. Therefore, we conclude that these statistics are not suitable to identify the modern population most similar to Guanches in the sense that we intend it for this study (Quantification and Statistical Analysis). In contrast, our extended analyses using FST (fixation index) [23, 24] and f2 [25] identify different North African and Near Eastern populations as most similar to modern North African populations and Guanches, results that are consistent with the PCA and the uniparental genetic analysis (Table S2; Figures 2 and S2). This finding is in agreement with anthropological, archaeological, linguistic [1, 4, 9, 26], and uniparental genetic data [6, 7, 8] and adds to a growing body of new evidence suggesting that some ancient domesticated plants [27] and animals [28] from the Canary Islands originated from North Africa. The Guanches’ Berber-like affinity is further supported by ADMIXTURE [29] analysis (Figures 3 and S3), where Guanches largely behave like modern Berbers across all values of K. At K = 10, a Northwest African-specific ancestry component makes up the greatest amount of autosomal ancestry in the Guanche and Berber populations in the HO dataset, such as the Mozabite and Saharawi. It is also ubiquitous across other Northwest African populations with Berber ancestry, such as Algerians and Tunisians, consistent with the PCA results. This ancestry component is also represented in present-day Canary Islanders and at a low proportion in some South European populations (Figures 3 and S3). Interestingly, it is also shared by Middle Eastern populations, including some Natufians (Figure 3). Y chromosome E1b1b haplotypes (though not M183 variants) were also common in Natufians (circa 11,000 BCE) and pre-pottery Neolithic male individuals from the Levant (circa 7,000 BCE), suggesting some affinity to North Africans [30]. |
|





