|
|
Post by Admin on Oct 4, 2021 22:55:05 GMT
Note 6: Y-chromosomal & phenotypic analysis Chuanchao Wang, Alexander Peltzer The current distribution of E1b1b1 in North Africa could also be caused by the back migration from the Near East to Africa that have already been proposed by several authors (77-79). The high frequencies of haplogroup R1-M173 in Cameroon also supported the back migration from Eurasia to Africa (80). Since it’s still unclear whether E1b1b evolved in Northeast Africa or the Near East, we were deciding against attempting to conclude whether the two haplogroups provide information about different paternal origin information in our three  samples.  For our phenotypic analysis, we investigated a set of SNPs thought to be affected by selection in our samples. Only high-quality (q > 30) bases were counted. We were able to find derived alleles for the genes SLC24A5 (rs1426654), which are known to be responsible for lighter skin pigmentation in JK2888 and JK2911. Our further tests whether the genes SLC45A2 (rs16891982), LCT (rs4988235), EDAR (rs3827760) and HERC2 (rs12913832) revealed no derived alleles for both JK2888 and JK2911. For JK2134, no sufficient coverage after quality filtering was given at the specific sites, which is why the analysis revealed no further clues (see Supplementary Table 6 for details). LCT is responsible for lactase persistence in Europe (81, 82). The SNPs at SLC24A5 and SLC45A2 are responsible for lighter skin pigmentation (83). The SNP at EDAR affects tooth morphology and hair thickness (84, 85). The SNP at HERC2 is the primary determinant of light eye colour in present-day Europeans (86, 87).
Supplementary Table 6: f4-ratio based estimates of African ancestry (α) in
Egyptians.
The “std.err” is the standard error estimated using jackknifes. “Z” is the Z-score
for the estimation.
Admixed populations West Eurasian African ancestry(α) std.err Z
Egyptian French 0.17165 0.004346 39.496
AncientEgyptians French 0.092278 0.010924 8.447
Egyptian Anatolia_N 0.142236 0.004902 29.016
AncientEgyptians Anatolia_N 0.06112 0.011791 5.184
Egyptian WHG 0.208021 0.00873 23.827
AncientEgyptians WHG 0.12929 0.013958 9.263
Egyptian EHG 0.209352 0.009337 22.421
AncientEgyptians EHG 0.138128 0.01385 9.973
Egyptian SHG 0.205056 0.007641 26.835
AncientEgyptians SHG 0.134262 0.013155 10.206
Egyptian Iran_N 0.138289 0.010708 12.914
AncientEgyptians Iran_N 0.056609 0.016074 3.522
Egyptian CHG 0.149992 0.00995 15.074
AncientEgyptians CHG 0.073004 0.01561 4.677
Egyptian MA1 0.22376 0.011275 19.846
AncientEgyptians MA1 0.148986 0.017456 8.535
Supplementary Table 7: The African admixture proportions estimated using
qpAdm.
Here “p” refers to the P-value for rank=1 and “std.err” is the standard error estimated
using jackknifes.
Admixed populations West Eurasian African ancestry(α) std.err p
Egyptian French 0.161 0.004 0.096
AncientEgyptians French 0.079 0.013 0.449
Egyptian Anatolia_N 0.13 0.005 0
AncientEgyptians Anatolia_N 0.041 0.014 0.48
Egyptian WHG 0.174 0.01 0.735
AncientEgyptians WHG 0.096 0.017 0.53
Egyptian EHG 0.177 0.011 0.504
AncientEgyptians EHG 0.089 0.018 0.519
Egyptian SHG 0.178 0.008 0.721
AncientEgyptians SHG 0.104 0.016 0.795
Egyptian Iran_N 0.109 0.012 0.821
AncientEgyptians Iran_N 0.029 0.02 0.957
Egyptian CHG 0.144 0.01 0.436
AncientEgyptians CHG 0.066 0.017 0.65
Egyptian MA1 0.172 0.014 0.838
AncientEgyptians MA1 0.09 0.022 0.903
|
|
|
|
Post by Admin on Oct 5, 2021 0:21:08 GMT
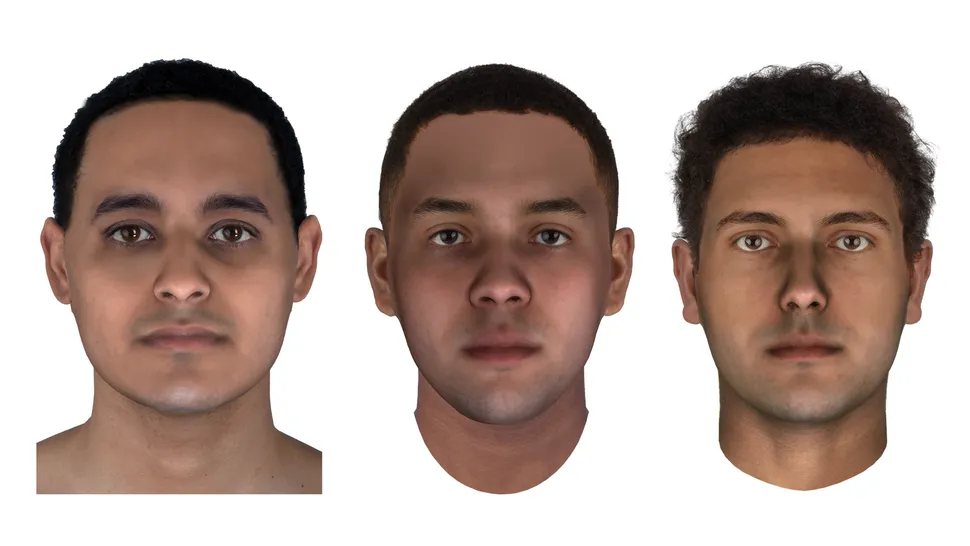 The faces of three men who lived in ancient Egypt more than 2,000 years ago have been brought back to life. Digital reconstructions depict the men at age 25, based on DNA data extracted from their mummified remains. The mummies came from Abusir el-Meleq, an ancient Egyptian city on a floodplain to the south of Cairo, and they were buried between 1380 B.C. and A.D. 425. Scientists at the Max Planck Institute for the Science of Human History in Tübingen, Germany, sequenced the mummies' DNA in 2017; it was the first successful reconstruction of an ancient Egyptian  's genome, Live Science reported at the time. "This is the first time comprehensive DNA phenotyping has been performed on human DNA of this age," Parabon representatives said in a statement. Parabon revealed the mummies' faces on Sept. 15 at the 32nd International Symposium on Human Identification in Orlando, Florida. Scientists used a phenotyping method called Snapshot to predict the men's ancestry, skin color and facial features. They found that the men had light brown skin with dark eyes and hair; overall, their genetic makeup was closer to that of modern individuals in the Mediterranean or the Middle East than it was to modern Egyptians', according to the statement. The researchers then generated 3D meshes outlining the mummies' facial features, and calculated heat maps to highlight the differences between the three individuals and refine the details of each face. Parabon's forensic artist then combined these results with Snapshot's predictions about skin, eye and hair color.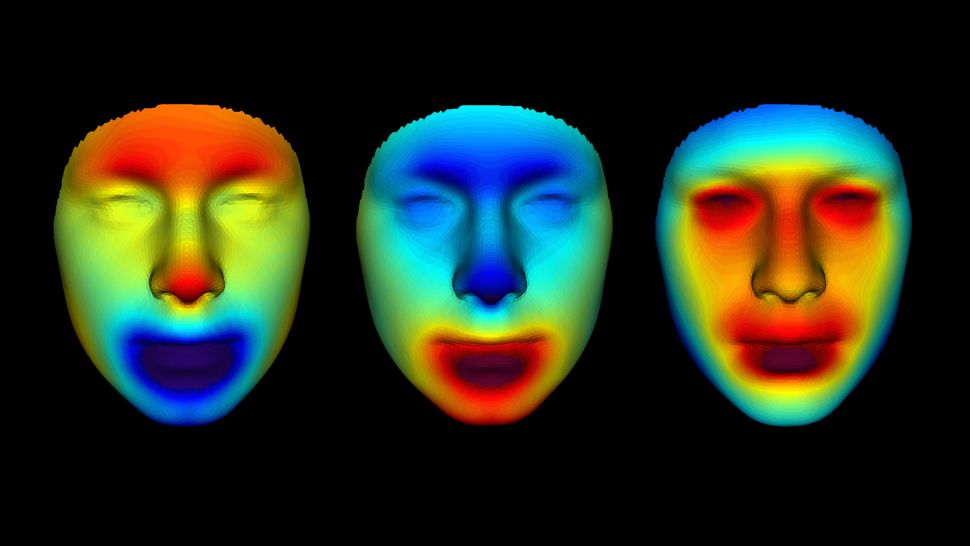  Working with ancient human DNA can be challenging for two reasons: the DNA is often highly degraded, and it's usually mixed with bacterial DNA, said Ellen Greytak, Parabon's director of bioinformatics. "Between those two factors, the amount of human DNA available to sequence can be very small," Greytak told Live Science in an email. However, because the vast majority of DNA is shared between all humans, scientists don't need the entire genome to glean a physical picture of a person. Rather, they only need to analyze certain specific spots in the genome that differ between people, known as single nucleotide polymorphisms (SNPs). Many of these SNPs code for physical differences between individuals, Greytak said. However, sometimes ancient DNA doesn't provide enough SNPs to pinpoint a given trait. In those cases, scientists can infer absent genetic data from values of other SNPs nearby, said Janet Cady, a Parabon bioinformatics scientist. Statistics that are calculated from thousands of genomes reveal how closely associated each SNP is with an absent neighbor, Cady told Live Science in an email. From there, the researchers can make a statistical prediction of what the missing SNP was. The processes used on these ancient mummies could also help scientists to recreate faces to identify modern remains, Greytak told Live Science. Of the approximately 175 cold cases that Parabon researchers have helped to solve using genetic genealogy, so far nine were analyzed using the techniques from this study, Greytak said. Originally published on Live Science. |
|
|
|
Post by Admin on Oct 5, 2021 1:46:31 GMT
Ancient Egyptian  genomes suggest an increase of Sub-Saharan African ancestry in post-Roman periods Abstract Egypt, located on the isthmus of Africa, is an ideal region to study historical population dynamics due to its geographic location and documented interactions with ancient civilizations in Africa, Asia and Europe. Particularly, in the first millennium BCE Egypt endured foreign domination leading to growing numbers of foreigners living within its borders possibly contributing genetically to the local population. Here we present 90 mitochondrial genomes as well as genome-wide data sets from three individuals obtained from Egyptian mummies. The samples recovered from Middle Egypt span around 1,300 years of ancient Egyptian history from the New Kingdom to the Roman Period. Our analyses reveal that ancient Egyptians shared more ancestry with Near Easterners than present-day Egyptians, who received additional sub-Saharan admixture in more recent times. This analysis establishes ancient Egyptian mummies as a genetic source to study ancient human history and offers the perspective of deciphering Egypt’s past at a genome-wide level. Introduction Egypt provides a privileged setting for the study of population genetics as a result of its long and involved population history. Owing to its rich natural resources and strategic location on the crossroads of continents, the country had intense, historically documented interactions with important cultural areas in Africa, Asia and Europe ranging from international trade to foreign invasion and rule. Especially from the first millennium BCE onwards, Egypt saw a growing number of foreigners living and working within its borders and was subjected to an almost continuous sequence of foreign domination by Libyans, Assyrians, Kushites, Persians, Greeks, Romans, Arabs, Turks and Brits. The movement of people, goods and ideas throughout Egypt’s long history has given rise to an intricate cultural and genetic exchange and entanglement, involving themes that resonate strongly with contemporary discourse on integration and globalization1. Until now the study of Egypt’s population history has been largely based on literary and archaeological sources and inferences drawn from genetic diversity in present-day Egyptians. Both approaches have made crucial contributions to the debate but are not without limitations. On the one hand, the interpretation of literary and archaeological sources is often complicated by selective representation and preservation and the fact that markers of foreign identity, such as, for example, Greek or Latin names and ethnics, quickly became ‘status symbols’ and were adopted by natives and foreigners alike2,3,4. On the other hand, results obtained by modern genetic studies are based on extrapolations from their modern data sets and make critical assumptions on population structure and time5. The analysis of ancient DNA provides a crucial piece in the puzzle of Egypt’s population history and can serve as an important corrective or supplement to inferences drawn from literary, archaeological and modern DNA data. Despite their potential to address research questions relating to population migrations, genetic studies of ancient Egyptian mummies and skeletal material remain rare, although research on Egyptian mummies helped to pioneer the field of ancient DNA research with the first reported retrieval of ancient human DNA6. Since then progress has been challenged by issues surrounding the authentication of the retrieved DNA and potential contaminations inherent to the direct PCR method7. Furthermore, the potential DNA preservation in Egyptian mummies was met with general scepticism: The hot Egyptian climate, the high humidity levels in many tombs and some of the chemicals used in mummification techniques, in particular sodium carbonate, all contribute to DNA degradation and are thought to render the long-term survival of DNA in Egyptian mummies improbable8. Experimental DNA decay rates in papyri have also been used to question the validity and general reliability of reported ancient Egyptian DNA results9. The recent genetic analysis of King Tutankhamun’s family10 is one of the latest controversial studies that gave rise to this extensive scholarly debate11. New data obtained with high-throughput sequencing methods have the potential to overcome the methodological and contamination issues surrounding the PCR method and could help settle the debate surrounding ancient Egyptian DNA preservation8. However, the first high-throughput sequences obtained from ancient Egyptian mummies12 were not supported by rigorous authenticity and contamination tests. Here, we provide the first reliable data set obtained from ancient Egyptians using high-throughput DNA sequencing methods and assessing the authenticity of the retrieved ancient DNA via characteristic nucleotide misincorporation patterns13,14 and statistical contamination tests15 to ensure the ancient origin of our obtained data. By directly studying ancient DNA from ancient Egyptians, we can test previous hypotheses drawn from analysing modern Egyptian DNA, such as recent admixture from populations with sub-Saharan16 and non-African ancestries17, attributed to trans-Saharan slave trade and the Islamic expansion, respectively. On a more local scale, we aim to study changes and continuities in the genetic makeup of the ancient inhabitants of the Abusir el-Meleq community (Fig. 1), since all sampled remains derive from this community in Middle Egypt and have been radiocarbon dated to the late New Kingdom to the Roman Period (cal. 1388BCE–426CE, Supplementary Data 1). In particular, we seek to determine if the inhabitants of this settlement were affected at the genetic level by foreign conquest and domination, especially during the Ptolemaic (332–30BCE) and Roman (30BCE–395CE) Periods. Figure 1: Geographic context, of the samples used in this study. 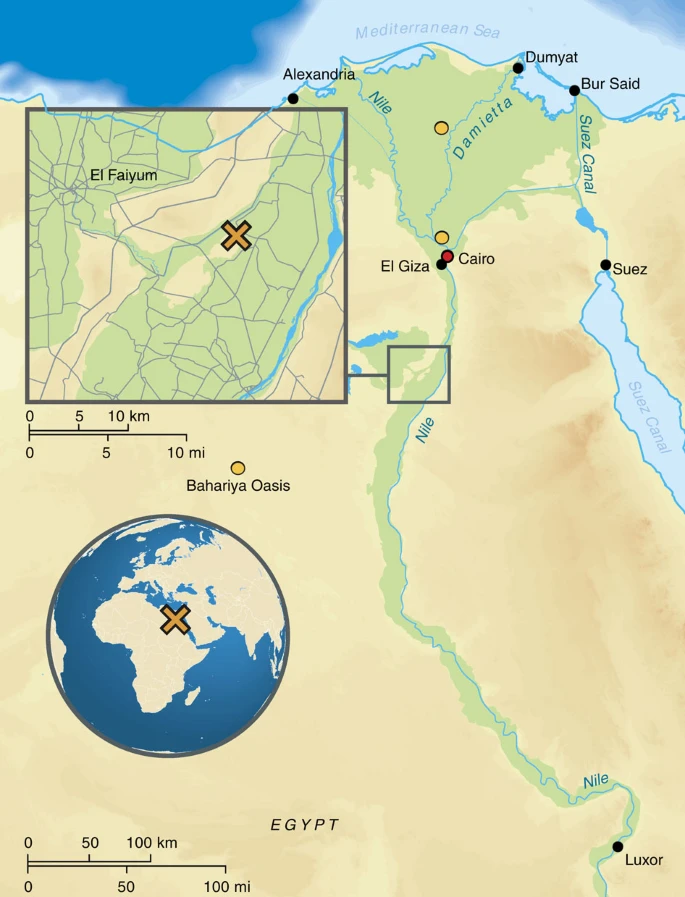 |
|
|
|
Post by Admin on Oct 5, 2021 4:41:41 GMT
Results Samples and anthropological analysis All 166 samples from 151 mummified individuals (for details of the 90 individuals included in the later analysis, see Supplementary Data 1) used in this study were taken from two anthropological collections at the University of Tübingen and the Felix von Luschan Skull Collection, which is now kept at the Museum of Prehistory of the Staatliche Museen zu Berlin, Stiftung preußischer Kulturbesitz (individuals: S3533, S3536, S3544, S3552, S3578, S3610). According to the radiocarbon dates (Supplementary Data 1, see also ref. 18), the samples can be grouped into three time periods: Pre-Ptolemaic (New Kingdom, Third Intermediate Period and Late Period), Ptolemaic and Roman Period. During their conservation in the Tübingen and Berlin collections the remains underwent different treatments: some were preserved in their original mummified state, while others were macerated for anthropological analysis or due to conservation problems19. In most cases, non-macerated  heads still have much of their soft tissue preserved. Some of the remains (individuals analysed in our study: 1543, 1547, 1565, 1577, 1611) have traces of gold leaf near the mouth and the cheekbone, which is characteristic for mummies from the Ptolemaic Period onwards20. In most cases the brain was removed and the excerebration route was highly likely transnasal, resulting in visible defects on the cribriform plate (for the individuals analysed in our study, see Supplementary Data 1). In summary, the excellent bone preservation and the more or less good soft tissue preservation made a wide-ranging analysis possible19. Recently, various studies were conducted on these remains, including a study on ancient Egyptian embalming resins, two ancient DNA studies and an anthropological examination of the macerated crania12,18,19,21. While the possibilities of a demographic reconstruction based on anthropological finds are naturally limited—due to incompleteness of the assemblage, the following anthropological observations were made on the assemblage: For a first assessment, computer tomographic scans of 30 mummies with soft tissue preservation were produced to describe sex (Supplementary Data 1), age at death (Supplementary Data 1) and the macroscopic health status; the six macerated mummies were examined directly. It is notable that most of the individuals are early and late adults, and that subadult individuals are underrepresented (Supplementary Data 1). It is possible that the sample’s demographic profile is the result of different burial treatments for adults and subadults, but it seems more likely that it is due to collection bias, with collectors favouring intact adult skulls. Almost all of the teeth show significant dentine exposure up to a total loss of the crown. This abrasion pattern is likely due to the food and food preparation itself, in particular for a cereal-rich diet containing a high proportion of coarse sandy particles. These particles act to abrade the dental tissues, allowing bacteria to penetrate the interior of the teeth. As a result, carious lesions or periapical processes appear in the analysed individuals (Supplementary Data 1)19. For the DNA analysis we sampled different tissues (bone, soft tissue, tooth), macerated and non-macerated, to test for human DNA preservation. Processing and sequencing of the samples We extracted DNA from 151 mummified human remains and prepared double-stranded Illumina libraries with dual barcodes22,23. Then we used DNA capture techniques for human mitochondrial DNA24 and for 1.24 million genomic single nucleotide polymorphisms (SNPs)25 in combination with Illumina sequencing, through which we successfully obtained complete human mitochondrial genomes from 90 samples and genome-wide SNP data from three male individuals passing quality control. |
|
|
|
Post by Admin on Oct 5, 2021 19:19:03 GMT
Comparison of the DNA preservation in different tissues We tested different tissues for DNA preservation and applied strict criteria for authenticity on the retrieved mitochondrial and nuclear DNA to establish authentic ancient Egyptian DNA. First, DNA extracts from several tissues (that is, bone, teeth, soft tissue and macerated teeth) from 151 individuals were screened for the presence of human mitochondrial DNA (mtDNA) resulting in a total of 2,157 to 982,165 quality filtered mitochondrial reads per sample, and 11- to 4,236-fold coverage. To estimate, identify and filter out potential contamination we applied the program schmutzi15 with strict criteria for contamination and kept only samples with less than 3% contamination for further analysis. For a comparison of different source material (soft tissue, bone and teeth) ten individuals (Supplementary Table 1) were sampled multiple times. Yields of preserved DNA were comparable in bone and teeth but up to ten times lower in soft tissues (Fig. 2a, Supplementary Table 1). Nucleotide misincorporation patterns characteristic for damaged ancient human DNA allowed us to assess the authenticity of the retrieved DNA13,14. The observed DNA damage patterns differed for the source materials with on average 19% damage in soft tissues and around 10% damage in bone tissue and teeth (Fig. 2b,c, Supplementary Table 1). Importantly, mtDNA haplotypes were identical for all samples from the same individuals. Our results thus suggest that DNA damage in Egyptian mummies correlates with tissue type. The protection of bone and teeth by the surrounding soft tissue or the embalmment of soft tissue may have contributed to the observed differences. Figure 2: DNA preservation and DNA damage of the samples used in this study. 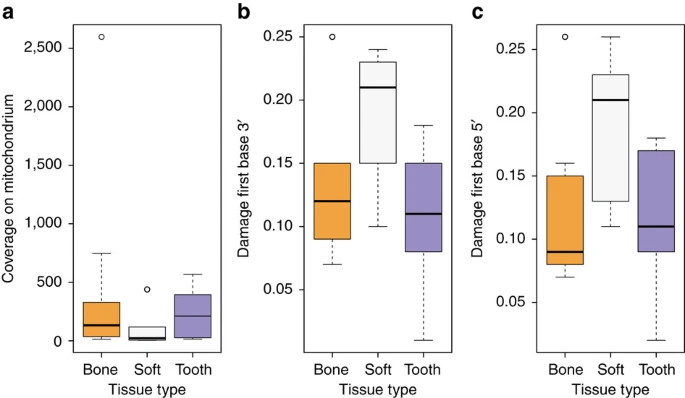 (a) coverage boxplots separated by tissue type (bone, mummified tissue, teeth), (b) boxplots showing damage of first base at the 3′ end separated by tissue type according to a, (c) damage on first base at the 5′ end of mapped reads separated by tissue type according to a and b. Generation of nuclear data In order to analyse the nuclear DNA we selected 40 samples with high mtDNA coverage and low mtDNA contamination. Using in solution enrichment for 1.2 million genome-wide SNPs26, we obtained between 3,632 and 508,360 target SNPs per sample (Supplementary Data 2). Overall, the nuclear DNA showed poor preservation compared to the mtDNA as depicted by a high mitochondrial/nuclear DNA ratio of on average around 18,000. In many samples, nuclear DNA damage was relatively low, indicating modern contamination. We sequenced two libraries per sample: one untreated library to assess DNA damage, and one library treated with enzymatic damage repair27, which was used for downstream analysis. We applied strict criteria for further analysis: we considered only male samples with at least 8% average cytosine deamination rates at the ends of the reads from the untreated library, and with at least 150 SNPs on the X chromosome covered at least twice, in order to estimate contamination levels reliably. Three out of 40 samples fulfilling these criteria had acceptable nuclear contamination rates: Two samples from the Pre-Ptolemaic Periods (New Kingdom to Late Period) had 5.3 and 0.5% nuclear contamination and yielded 132,084 and 508,360 SNPs, respectively, and one sample from the Ptolemaic Period had 7.3% contamination and yielded 201,967 SNPs. As shown below, to rule out any impact of potential contamination on our results, we analysed the three samples separately or replicated results using only the least contaminated sample. |
|
 samples.
samples.
 samples.
samples.




