Post by Admin on Nov 7, 2023 19:48:46 GMT
Genetic contributions to Kalinago skin color variation
One motivation for undertaking this work was to characterize genetic contributions to skin pigmentation in a population with primarily Native American and African genetic ancestry, so that we could focus on the effect of Native American hypopigmenting alleles without interference from European alleles. The Kalinago population described here comprises the only population we are aware of that fits this genetic ancestry profile. To control for the effects of the major European pigmentation loci, all Kalinago samples were genotyped for SLC24A5A111T and SLC45A2L374F. The phenotypic effects of these variants and OCA2NW273KV are shown in Figure 4. Each variant decreases melanin pigmentation, with homozygotes being lighter than heterozygotes. The greatest effect is seen in the OCA2NW273KV homozygotes (the albino individuals), as previously noted. The frequencies of the derived alleles of SLC24A5A111T and SLC45A2L374F in the Kalinago sample are 0.14 and 0.06, respectively.
Figure 4 with 2 supplements
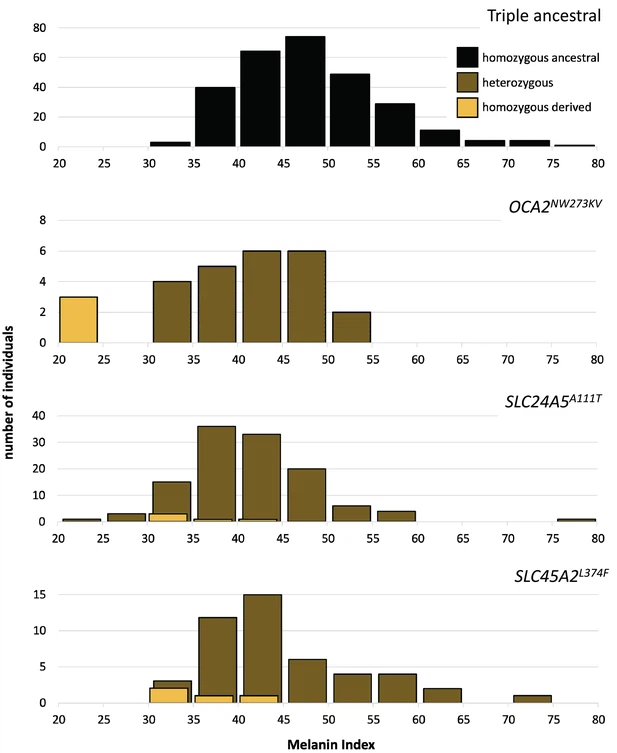
Skin color distribution of Kalinago samples according to genotype.
The ‘triple ancestral’ plot is individuals ancestral for three pigmentation loci (SLC24A5111A, SLC45A2374L, and OCA2273NW). In the other plots, heterozygosity or homozygosity is indicated for the … see more
Figure 4—source data 1
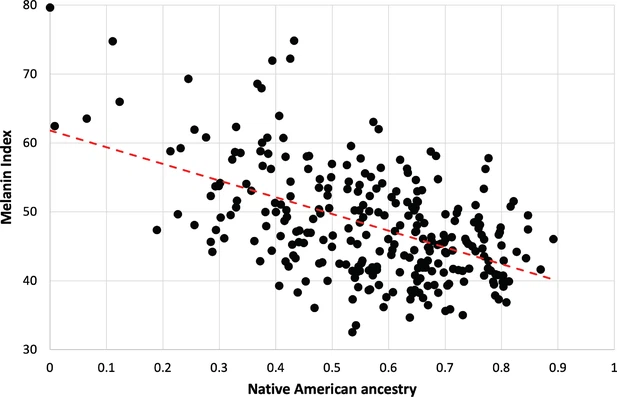
The source file contain melanin index distribution as function of community-described ancestry.
cdn.elifesciences.org/articles/77514/elife-77514-fig4-data1-v3.xlsx
Download elife-77514-fig4-data1-v3.xlsx
Figure 4—source data 2
The source data contains data of melanin indices according to genotype.
cdn.elifesciences.org/articles/77514/elife-77514-fig4-data2-v3.xlsx
Download elife-77514-fig4-data2-v3.xlsx
The markedly higher frequency of SLC24A5A111T compared to SLC45A2L374F is not explained solely by European admixture, given that most Europeans are nearly fixed for both alleles (Soejima and Koda, 2007). This deviation can be explained by the involvement of source populations that carry the SLC24A5A111T variant but not SLC45A2L374F. Although some sub-Saharan West African populations (the likeliest source of AFR genetic ancestry in the Kalinago) have negligible SLC24A5A111T frequencies, moderate frequencies are found in the Mende of Sierra Leone (MSL, allele frequency = 0.08) (Micheletti et al., 2020; Auton et al., 2015), while some West African populations such as Hausa and Mandinka who have allele frequencies of 0.11 and 0.15, respectively (Cheung et al., 2000; Rajeevan et al., 2012). Such African individuals carrying the SLC24A5A111T allele could potentially cause the observed frequencies by founder effect. In addition, the region of chromosome 5 containing SLC45A2 exhibits low European genetic ancestry (6.5%) that is consistent with low observed SLC45A2L374F frequency.
One motivation for undertaking this work was to characterize genetic contributions to skin pigmentation in a population with primarily Native American and African genetic ancestry, so that we could focus on the effect of Native American hypopigmenting alleles without interference from European alleles. The Kalinago population described here comprises the only population we are aware of that fits this genetic ancestry profile. To control for the effects of the major European pigmentation loci, all Kalinago samples were genotyped for SLC24A5A111T and SLC45A2L374F. The phenotypic effects of these variants and OCA2NW273KV are shown in Figure 4. Each variant decreases melanin pigmentation, with homozygotes being lighter than heterozygotes. The greatest effect is seen in the OCA2NW273KV homozygotes (the albino individuals), as previously noted. The frequencies of the derived alleles of SLC24A5A111T and SLC45A2L374F in the Kalinago sample are 0.14 and 0.06, respectively.
Figure 4 with 2 supplements

Skin color distribution of Kalinago samples according to genotype.
The ‘triple ancestral’ plot is individuals ancestral for three pigmentation loci (SLC24A5111A, SLC45A2374L, and OCA2273NW). In the other plots, heterozygosity or homozygosity is indicated for the … see more
Figure 4—source data 1
The source file contain melanin index distribution as function of community-described ancestry.
cdn.elifesciences.org/articles/77514/elife-77514-fig4-data1-v3.xlsx
Download elife-77514-fig4-data1-v3.xlsx
Figure 4—source data 2
The source data contains data of melanin indices according to genotype.
cdn.elifesciences.org/articles/77514/elife-77514-fig4-data2-v3.xlsx
Download elife-77514-fig4-data2-v3.xlsx
The markedly higher frequency of SLC24A5A111T compared to SLC45A2L374F is not explained solely by European admixture, given that most Europeans are nearly fixed for both alleles (Soejima and Koda, 2007). This deviation can be explained by the involvement of source populations that carry the SLC24A5A111T variant but not SLC45A2L374F. Although some sub-Saharan West African populations (the likeliest source of AFR genetic ancestry in the Kalinago) have negligible SLC24A5A111T frequencies, moderate frequencies are found in the Mende of Sierra Leone (MSL, allele frequency = 0.08) (Micheletti et al., 2020; Auton et al., 2015), while some West African populations such as Hausa and Mandinka who have allele frequencies of 0.11 and 0.15, respectively (Cheung et al., 2000; Rajeevan et al., 2012). Such African individuals carrying the SLC24A5A111T allele could potentially cause the observed frequencies by founder effect. In addition, the region of chromosome 5 containing SLC45A2 exhibits low European genetic ancestry (6.5%) that is consistent with low observed SLC45A2L374F frequency.