|
|
Post by Admin on May 7, 2023 18:28:50 GMT
Figure 3 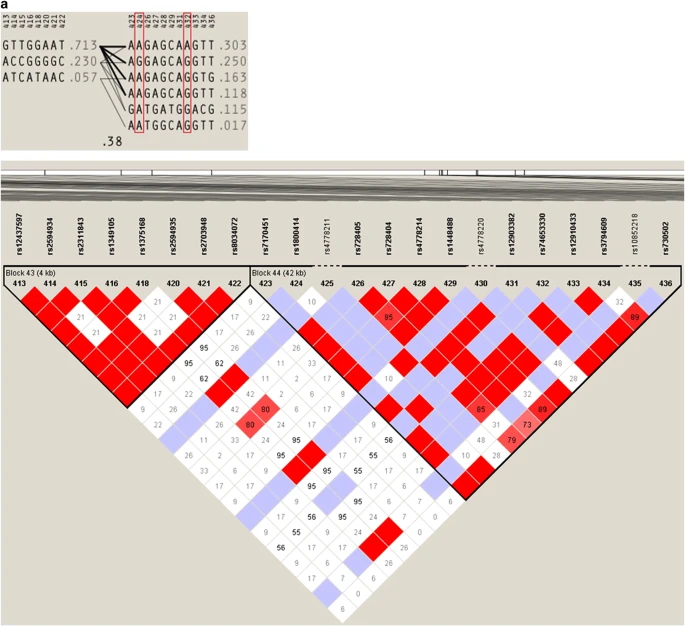 Haplotype block structure and pattern of LD of the OCA2 region including markers rs1800414 (marker 424) and rs74653330 (marker 432). (a) Northern East Asia; (b) in the rest of Asia. Recent ancient DNA studies, which have characterized dense genomic data in Eurasian individuals spanning a broad archaeological period (e.g., from hunter gatherers to individuals living in the Bronze Age), have provided important information about the temporal distribution of genetic markers associated with pigmentation variation in Europe and have strengthened the case for selection operating in pigmentation-related genes in this region.20–22 Similar studies in East Asia have the potential to clarify the major events that have shaped the interesting distribution of the two non-synonymous variants of the OCA2 gene in this vast area. In this respect, it will be important to consider not only potential selective effects but also the major population movements that have taken place in this region during the past 15,000 years. |
|
|
|
Post by Admin on May 8, 2023 18:07:37 GMT
Native American Ancestry and Pigmentation Allele Contributions to Skin Color in a Caribbean Population Abstract Interest in the genetic basis of variation in skin pigmentation in Native American populations led us to seek indigenous populations of the Western Hemisphere with African and minimal European admixture to study the effect of Native American ancestry on skin color. Admixture analysis from DNA collected from 458 individuals in the Kalinago territory of the Commonwealth of Dominica showed shared ancestry with East Asians at K=3 and 55% Native American, 32% African, and 11% European ancestry at K=6, the highest Native American ancestry of Caribbean populations. Skin pigmentation was 20 to 80 melanin units, averaging 46. Three albino individuals were homozygous for multi-nucleotide polymorphism OCA2NW273KV of African origin, whose population allele frequency was 0.03 and single allele effect size was −8 melanin units. Hypopigmenting allele frequencies for SLC24A5A111T and SLC45A2L374F were 0.14 and 0.05, whose single allele effect sizes were −6 and −3, respectively. Skin color plots of individuals lacking known hypopigmenting alleles suggests that Native American Ancestry reduced pigmentation by more than 20 melanin units (low and high estimates 21.8 and 28.5). Shared ancestry with East Asians at K=3 suggests potential sharing of one or more pigmentation alleles. Introduction Skin pigmentation is a highly heritable polygenic trait influenced by health and environment (Barsh, 2003). European variants that lighten pigmentation include non-ancestral coding polymorphisms in SLC24A5 (Basu Mallick et al., 2013; Lamason et al., 2005; Soejima and Koda, 2007) and SLC45A2 (Lucotte and Yuasa, 2013; Soejima and Koda, 2007; Yuasa et al., 2006) that are nearly fixed. However, the genetic basis for lighter skin pigmentation in Native American and East Asian populations, who carry the same ancestral alleles as Africans at these loci, has yet to be established. Dark skin was the ancestral phenotype for anatomically modern humans (Jablonski and Chaplin, 2000; Lamason et al., 2005), whose evolution towards lighter skin at higher latitudes occurred independently in eastern and western Eurasia (Adhikari et al., 2019; Basu Mallick et al., 2013; Norton et al., 2007; Yang et al., 2016), potentially driven by a UV-dependent photoactivation step in the formation of vitamin D (Engelsen, 2010; Hanel and Carlberg, 2020; Holick, 1981; Loomis, 1967). Native Americans share common ancestry with East Asians (Derenko et al., 2010; Tamm et al., 2007), diverging before 15 kya (Gravel et al., 2013; Moreno-Mayar et al., 2018; Reich et al., 2012), but the extent to which these populations share pigmentation variants remains to be determined. High European admixture is characteristic of most Native American populations (Brown et al., 2017; Gravel et al., 2013; Klimentidis et al., 2009; Reich et al., 2012), complicating the characterization of pigmentation variants specific to Native Americans. Prior to European contact, the Caribbean islands were inhabited by populations who migrated from the northern coast of South America (Benn-Torres et al., 2008; HARVEY et al., 1969; Honychurch, 2012; “Island Caribs,” 2016; Torres et al., 2015, 2013). During the Colonial period, large numbers of Africans were introduced into the Caribbean as slave labor (Honychurch, 2012; Torres et al., 2013). As a consequence African and European admixture, and high mortality among the indigenous populations, Native American ancestry now contributes only a minor portion of the ancestry of most Caribbean islanders (1000 Genomes Project Consortium, 2010; The 1000 Genomes Project Consortium, 2015; Torres et al., 2015, 2013). The islands of Dominica and St. Vincent were the last colonized by Europeans, in the late 1700s (Honychurch, 2012, 1998; Rogoziński, 2000). In 1903, the British granted 15 km2 (3,700 acres) on the eastern coast of Dominica as a reservation for the Kalinago, who were then called “Carib”. When Dominica gained Independence in 1978, legal rights and a degree of protection from assimilation were gained by the inhabitants of the Carib Reserve (redesignated Kalinago Territory in 2015) (Honychurch, 2012). The Kalinago, numbering about 3,000 living within the Territory (“Kalinago Territory,” 2021), consider themselves to be of primarily Native American and African ancestry. Early in our genetic and phenotypic survey of the Kalinago, we noted one individual with albinism, and discovered that two others were known to reside in the Territory. Oculocutaneous albinism (OCA) is a recessive trait characterized by visual system abnormalities and hypopigmentation of skin, hair, and eyes (Gargiulo et al., 2011; Gronskov et al., 2007; Grønskov et al., 2014; Hong et al., 2006; Vogel et al., 2008) that is caused by mutations in any of several autosomal pigmentation genes (Carrasco et al., 2009; Edwards et al., 2010; Gao et al., 2017; Grønskov et al., 2013; Kausar et al., 2013; King et al., 2003; Spritz et al., 1995; Stevens et al., 1997, 1995; Vogel et al., 2008; Woolf, 2005; Yi et al., 2003). Here, we report on ancestry, distribution of measured skin color, identification of an albinism allele, and the hypopigmenting effects of this allele and the European SLC24A5A111T and SLC45A2L374 alleles in a sample representing 15% of the population (Figure S1). www.biorxiv.org/content/10.1101/2021.11.24.469305v3.full |
|
|
|
Post by Admin on Jul 8, 2023 21:28:28 GMT
The type of pigment that gives hair its color is called melanin. There are two types of melanin: eumelanin, which gives hair a brown or black color, and pheomelanin, which gives hair a red or blonde color. In general, the more eumelanin a person has, the darker their hair will be. The more pheomelanin they have, the lighter their hair will be. Children tend to have less melanin overall, and therefore often have lighter hair. As they grow older and their bodies produce more melanin, their hair may become darker. Additionally, some children may have a different distribution of melanin in their hair, which can change over time, leading to a change in hair color as well. Hair color is determined by the amount of a pigment called melanin in hair. An abundance of one type of melanin, called eumelanin, gives people black or brown hair. An abundance of another pigment, called pheomelanin, gives people red hair. The type and amount of melanin determines hair color Hair color Type and amount of melanin Black Large amount of eumelanin Brown Moderate amount of eumelanin Blond Very little eumelanin Red Mostly pheomelanin with a little eumelanin The type and amount of melanin in hair is determined by many genes, although little is known about most of them. The best-studied hair-color gene in humans is called MC1R. This gene provides instructions for making a protein called the melanocortin 1 receptor, which is involved in the pathway that produces melanin. The melanocortin 1 receptor controls which type of melanin is produced by melanocytes. When the receptor is turned on (activated), it triggers a series of chemical reactions inside melanocytes that stimulate these cells to make eumelanin. If the receptor is not activated or is blocked, melanocytes make pheomelanin instead of eumelanin. Many other genes also help to regulate this process. Most people have two functioning copies of the MC1R gene, one inherited from each parent. These individuals have black or brown hair, because of the high amount of eumelanin. It is estimated that more than 90 percent of people in the world have brown or black hair. Some people have variations in one copy of the MC1R gene in each cell that causes the gene to be turned off (deactivated). This type of genetic change is described as loss-of-function. For these individuals, eumelanin production is lower, while pheomelanin production is higher, so they have strawberry blond, auburn, or red hair. In an even smaller percentage of people, both copies of the MC1R gene in each cell have loss-of-function changes, and the melanin-production pathway produces only the pheomelanin pigment. The hair of these individuals is almost always very red. Even when the melanin-production pathway is making eumelanin, changes in other genes can reduce the amount of eumelanin produced. These changes lead to blond hair. Hair color ranges across a wide spectrum of hues, from flaxen blond to coal black. Many genes other than MC1R play a role in determining shades of hair color by controlling levels of eumelanin and pheomelanin. Some of these genes, including ASIP, DTNBP1, GPR143, HPS3, KITLG, MLPH, MYO5A, MYO7A, OCA2, SLC45A2, SLC24A5, TYRP1, TYR, ERCC6, GNAS, HERC2, IRF4, OBSCN, SLC24A4, TPCN2, and MITF, are involved in the production of melanin in hair. Some of these genes are associated with gene transcription (which is the first step in protein production), DNA repair, the transport of substances (such as calcium) across cell membranes, or the structure of hair follicles. Several of these genes contribute to eye and skin color, but the exact role they play in determining hair color is unknown. Hair color may change over time. Particularly in people of European descent, light hair color may darken as individuals grow older. For example, blond-haired children often have darker hair by the time they are teenagers. Researchers speculate that certain hair-pigment proteins are activated as children grow older, perhaps in response to hormonal changes that occur near puberty. Almost everyone’s hair will begin to turn gray as they age, although when it happens and to what extent is variable. Gray hair is partly hereditary and may vary by ethnic origin; it is also somewhat dependent on external factors such as stress. Hair becomes gray when the hair follicle loses its ability to make melanin, but exactly why that occurs is not clear. Scientific journal articles for further reading Branicki W, Liu F, van Duijn K, Draus-Barini J, Pośpiech E, Walsh S, Kupiec T, Wojas-Pelc A, Kayser M. Model-based prediction of human hair color using DNA variants. Hum Genet. 2011 Apr;129(4):443-54. doi: 10.1007/s00439-010-0939-8. Epub 2011 Jan 4. PubMed: 21197618. Free full-text available from PubMed Central: PMC3057002. www.ncbi.nlm.nih.gov/pmc/articles/PMC3057002/Commo S, Gaillard O, Thibaut S, Bernard BA. Absence of TRP-2 in melanogenic melanocytes of human hair. Pigment Cell Res. 2004 Oct;17(5):488-97. PubMed: 15357835. Han J, Kraft P, Nan H, Guo Q, Chen C, Qureshi A, Hankinson SE, Hu FB, Duffy DL, Zhao ZZ, Martin NG, Montgomery GW, Hayward NK, Thomas G, Hoover RN, Chanock S, Hunter DJ. A genome-wide association study identifies novel alleles associated with hair color and skin pigmentation. PLoS Genet. 2008 May 16;4(5):e1000074. doi: 10.1371/journal.pgen.1000074. PubMed: 18483556. Free full-text available from PubMed Central: PMC2367449. Ito S, Wakamatsu K. Diversity of human hair pigmentation as studied by chemical analysis of eumelanin and pheomelanin. J Eur Acad Dermatol Venereol. 2011 Dec;25(12):1369-80. doi: 10.1111/j.1468-3083.2011.04278.x. PubMed: 22077870. Liu F, Wen B, Kayser M. Colorful DNA polymorphisms in humans. Semin Cell Dev Biol. 2013 Jun-Jul;24(6-7):562-75. doi: 10.1016/j.semcdb.2013.03.013. Epub 2013 Apr 12. PubMed: 23587773. Schaffer JV, Bolognia JL. The melanocortin-1 receptor: red hair and beyond. Arch Dermatol. 2001 Nov;137(11):1477-85. PubMed: 11708951. Siewierska-Górska A, Sitek A, Żądzińska E, Bartosz G, Strapagiel D. Association of five SNPs with human hair colour in the Polish population. Homo. 2017 Mar;68(2):134-144. doi: 10.1016/j.jchb.2017.02.002. Epub 2017 Feb 4. PubMed: 28242083. Sturm RA. Molecular genetics of human pigmentation diversity. Hum Mol Genet. 2009 Apr 15;18(R1):R9-17. doi: 10.1093/hmg/ddp003. PubMed: 19297406. Tobin DJ. Human hair pigmentation--biological aspects. Int J Cosmet Sci. 2008 Aug;30(4):233-57. doi: 10.1111/j.1468-2494.2008.00456.x. PubMed: 18713071. |
|
|
|
Post by Admin on Jul 9, 2023 3:19:12 GMT
Model-based prediction of human hair color using DNA variants Wojciech Branicki,1,2 Fan Liu,3 Kate van Duijn,3 Jolanta Draus-Barini,1 Ewelina Pośpiech,1 Susan Walsh,3 Tomasz Kupiec,1 Anna Wojas-Pelc,4 and Manfred Kayser3 Abstract Predicting complex human phenotypes from genotypes is the central concept of widely advocated personalized medicine, but so far has rarely led to high accuracies limiting practical applications. One notable exception, although less relevant for medical but important for forensic purposes, is human eye color, for which it has been recently demonstrated that highly accurate prediction is feasible from a small number of DNA variants. Here, we demonstrate that human hair color is predictable from DNA variants with similarly high accuracies. We analyzed in Polish Europeans with single-observer hair color grading 45 single nucleotide polymorphisms (SNPs) from 12 genes previously associated with human hair color variation. We found that a model based on a subset of 13 single or compound genetic markers from 11 genes predicted red hair color with over 0.9, black hair color with almost 0.9, as well as blond, and brown hair color with over 0.8 prevalence-adjusted accuracy expressed by the area under the receiver characteristic operating curves (AUC). The identified genetic predictors also differentiate reasonably well between similar hair colors, such as between red and blond-red, as well as between blond and dark-blond, highlighting the value of the identified DNA variants for accurate hair color prediction. Electronic supplementary material The online version of this article (doi:10.1007/s00439-010-0939-8) contains supplementary material, which is available to authorized users. Introduction The concept of personalized medicine assumes that prediction of phenotypes based on genome information can enable better prognosis, prevention and medical care which can be tailored individually (Brand et al. 2008; Janssens and van Duijn 2008). However, practical application of genome-based information to medicine requires the disease risk to be predicted with high accuracy, while knowledge on genetics of common complex diseases is still insufficient to allow their accurate prediction solely from DNA data (Alaerts and Del-Favero 2009; Chung et al. 2010; Ku et al. 2010; McCarthy and Zeggini 2009). Another potential application for prediction of phenotypes from genotypes is forensic science. Knowledge gained on externally visible characteristics (EVC) from genotype data obtained by examination of crime scene samples may be used for investigative intelligence purposes, especially in suspect-less cases (Kayser and Schneider 2009). The idea is based on using DNA-predicted EVC information to encircle a perpetrator in a larger population of unknown suspects. Such approach could also be useful in cases pertaining identification of human remains by extending anthropological findings on physical appearance of an identified individual. However, the genetic understanding of human appearance is still in its infancy. One notable exception is eye (iris) color, where previous candidate gene studies and especially recent genome-wide association studies (GWAS) revealed 15 genes involved (Eiberg et al. 2008; Frudakis et al. 2003; Graf et al. 2005; Han et al. 2008; Kanetsky et al. 2002; Kayser et al. 2008; Liu et al. 2010; Rebbeck et al. 2002; Sulem et al. 2007). One of them, HERC2, harbors genetic variation most strongly associated with human eye color variation (Eiberg et al. 2008; Kayser et al. 2008; Liu et al. 2010; Sturm et al. 2008). Moreover, a recent systematic study investigating the predictive value of eye color associated single nucleotide polymorphisms (SNPs) (Liu et al. 2009) found that a model with 15 SNPs from 8 genes predicts categorized blue and brown eye color with high accuracies with a subset of only 6 SNPs covering most of the predictive information. The IrisPlex system employing these six SNPs and a prediction model based on data from thousands of Europeans was recently developed and validated for DNA prediction of human eye color in forensic applications (Walsh et al. 2010a, b). Furthermore, a recent GWAS on quantitative eye color explained about 50% of continuous eye color variation by genetic factors (Liu et al. 2010). The recent progress on DNA prediction of human eye color raises expectations for the DNA prediction of other human pigmentation traits, such as hair color. Inheritance of one particular hair color in humans i.e., red hair, has already been explained to a significant degree. Valverde et al. (1995) have found that red hair color is mainly associated with polymorphisms in the MC1R gene. This information has been confirmed since in many other studies performed on various population samples (Box et al. 1997; Han et al. 2008; Harding et al. 2000; Flanagan et al. 2000; Kanetsky et al. 2004; Pastorino et al. 2004; Rana et al. 1999; Sulem et al. 2007). MC1R SNPs are fairly indicative for red hair and thus have already been implemented in forensic science (Branicki et al. 2007; Grimes et al. 2001), but the practical application of red hair color prediction (without the ability to predict other hair colors) strongly depends on the population it is applied to, given the strong differences in red hair color frequency between populations. Additional data on red hair color inheritance came from a recent GWAS in Icelanders, which revealed two SNPs in the ASIP gene, representing a MC1R antagonist, as significantly associated with red hair color (Sulem et al. 2008). Moreover, a position in the 3′-UTR of the ASIP gene was previously associated with dark hair color in European populations (Kanetsky et al. 2002; Voisey et al. 2006) using a candidate gene approach. The candidate gene approach also delivered two non-synonymous SNPs in SLC45A2 (MATP) with association to dark hair color in another study with several confirmation studies (Branicki et al. 2008a; Fernandez et al. 2008; Graf et al. 2005). Via several large GWASs various SNPs in/nearby genes in addition to MC1R, ASIP and SLC45A2 have been found with association to human hair color variation, such as OCA2, HERC2, SLC24A4, KITLG, TYR, TPCN2, TYRP1, IRF4, EXOC2, KIF26A, and OBSCN (Han et al. 2008; Sulem et al. 2007, 2008). Furthermore, two recently published studies tested for hair color association of SNPs from a large number of candidate genes, not only confirmed some previously known hair color genes, such as KITLG, OCA2, MC1R, TYRP1, TYR, SLC45A2, HERC2, ASIP, but additionally reported association with quantitative measures of hair color of SNPs in additional genes, such as SLC24A5, MYO5A, MYO7A, MLPH, GPR143, DCT, HPS3, GNAS, PRKARIA, ERCC6, and DTNBP1 in one or both studies (Mengel-From et al. 2009; Valenzuela et al. 2010). In the present study, we tested in Polish Europeans with single-observer hair color phenotype data the predictive power of 45 SNPs from 12 genes previously implicated with replicated evidence in human hair color variation. www.ncbi.nlm.nih.gov/pmc/articles/PMC3057002/ |
|
|
|
Post by Admin on Jul 19, 2023 19:42:42 GMT
The common occurrence of blond hair among the dark-skinned indigenous people of the Solomon Islands is due to a homegrown genetic variant distinct from the gene that leads to blond hair in Europeans, according to a new study from the Stanford University School of Medicine. “This is one of the most beautiful examples to date of the mapping of a simple genetic trait in humans,” said David Reich, PhD, a professor of genetics at Harvard University, who was not involved in the study. The study identifying the gene responsible for blond hair in the Solomon Islands, a nation in the South Pacific, represents a rare case of simple genetics determining human appearance, and shows the importance of including understudied populations in gene mapping studies, said co-senior author Carlos D. Bustamante, PhD, professor of genetics at Stanford. The findings were published May 4 in Science. “Since most studies in human genetics only include participants of European descent, we may be getting a very biased view of which genes and mutations influence the traits we investigate. Here, we sought to test whether one of the most striking human traits, blond hair, had the same — or different — genetic underpinning in different human populations,” Bustamante said. Globally, blond hair is rare, occurring with substantial frequency only in northern Europe and in Oceania, which includes the Solomon Islands and its neighbors. “Its frequency is between 5 and 10 percent across the Solomon Islands, which is about the same as where I’m from,” said co-first author Eimear Kenny, PhD, who was born in Ireland. Many assumed the blond hair of Melanesia was the result of gene flow — a trait passed on by European explorers, traders and others who visited in the preceding centuries. The islanders themselves give several possible explanations for its presence, said co-senior author Sean Myles, PhD, a former Stanford postdoctoral scholar who is now an assistant professor at the Nova Scotia Agricultural College. They generally chalked it up to sun exposure, or a diet rich in fish, he said. After researchers at UCSF generated genetic data from the samples, Kenny, a postdoctoral scholar in Bustamante’s lab, began the analysis in September 2010, the week she started at Stanford. “Within a week we had our initial result. It was such a striking signal pointing to a single gene — a result you could hang your hat on. That rarely happens in science,” she said. “It was one of the best experiences of my career.” In terms of genetic studies, the analysis was straightforward, said Kenny. But gathering the data, accomplished in 2009 by Myles and co-first author Nicholas Timpson, PhD, was more difficult. Much of the Solomon Islands is undeveloped, without roads, electricity or telephones. It’s also one of the most linguistically diverse nations in the world, with dozens of languages spoken. It was a return trip for Myles who had been there in 2004 as a graduate student with Max Planck Institute molecular anthropologist Mark Stoneking, PhD, (also a co-author of the study) to investigate whether the language variations correlated with genetic variations. While there, Myles was fascinated by the ubiquity of blond hair, which was especially common among children.  Stanford researchers say they were surprised that the evidence pointed so strongly to a single gene being responsible for the blond hair among Solomon Islanders. Additional photos are available here. “They have this very dark skin and bright blond hair. It was mind-blowing,” said Myles. “As a geneticist on the beach watching the kids playing, you count up the frequency of kids with blond hair, and say, ‘Wow, it’s 5 to 10 percent.’” A grant from the Wenner-Gren Foundation for Anthropological Research gave Myles, who at that time was doing a stint as a postdoctoral researcher at Cornell University, his chance to study the genetics of the Solomon Islanders’ hair color. Myles worked with Bustamante, who was also at Cornell, to design the study. Then back in the islands, Myles and Timpson went village to village explaining what they wanted to do and asking for permission to gather data, Myles speaking in Solomon Islands pidgin, the most widely understood language. When the local chief gave the OK, the researchers recruited participants and assessed hair and skin color using a light reflectance meter, took blood pressure readings and measured heights and weights. They asked the villagers to spit into small tubes to provide saliva to be used for DNA extraction. In the span of a month they collected more than 1,000 samples. While the islands fit many people’s notion of a tropical paradise, they lack amenities Westerners take for granted. For instance, simply finding a level spot for the scale to weigh study participants was a challenge. Then in 2010 Bustamante joined Stanford’s faculty and, with funding from the Department of Genetics, the team looked for genes underlying this striking phenotype. Soon after, Kenny joined the lab and started the analysis, selecting 43 blond- and 42 dark-haired Solomon Islanders from the opposite 10 percent extremes of the hair pigmentation range. She used these in a genome-wide association study, a method to reveal differences in the frequency of genetic variants between two groups, that usually requires thousands of samples. Because the vast majority of human physical characteristics analyzed to date have many genetic and environmental factors, Kenny expected an inconclusive result that would require much further study. Instead, she immediately saw a single strong signal on chromosome 9, which accounted for 50 percent of the variance in the Solomon Islanders’ hair color. The team went on to identify the gene responsible, TYRP1, which encodes tyrosinase-related protein 1, an enzyme previously recognized as influencing pigmentation in mice and humans. Further research revealed that the particular variant responsible for blond hair in the Solomon Islands is absent in the genomes of Europeans. “So the human characteristic of blond hair arose independently in equatorial Oceania. That’s quite unexpected and fascinating,” Kenny said. The finding underscores the importance of genetic studies on isolated populations, said Bustamante. “If we’re going to be designing the next generation of medical treatments using genetic information and we don’t have a really broad spectrum of populations included, you could disproportionately benefit some populations and harm others.” Bustamante is seeking funding to analyze the rest of the data gathered. “For instance, the genetics of skin pigmentation might be different there too — not the same as in Europe or Africa or India. We just don’t know.” Additional co-authors were Stanford postdoctoral scholars Martin Sikora, PhD, and Andres Moreno Estrada, PhD; Stanford research assistant Muh-Ching Yee, PhD; and researchers from UCSF including professor of bioengineering & therapeutic sciences and medicine, Esteban González Burchard, MD. Nicholas Timpson is currently a lecturer at the University of Bristol, U.K. In addition to the Wenner-Gren Foundation, the research was funded by the MRC Centre for Causal Analyses in Translational Epidemiology, the National Human Genome Research Institute, the National Heart, Lung, and Blood Institute and the Max Planck Society. Abstract Blond hair is a rare human phenotype found almost exclusively in Europe and Oceania. Here, we identify a cystine-to-arginine change at a highly conserved residue in tyrosinase-related protein 1 (TYRP1) as the single source of blond hair in Solomon Islanders. This missense mutation is predicted to impact catalytic activity of the protein and causes blond hair through a recessive mode of inheritance. The novel mutation is at a frequency of 26% in the Solomon Islands but is absent outside of Oceania and represents the largest genetic effect on a visible human phenotype reported to date. Our findings demonstrate that alleles of large effect reach appreciable frequencies in geographically isolated populations and underscore the importance of extending medical genomics to humans worldwide. www.ncbi.nlm.nih.gov/pmc/articles/PMC3481182/ |
|
