|
|
Post by Admin on Aug 5, 2021 22:20:50 GMT
The legacy of lost father tongues and the spread of agriculture
We previously took issue with the hypothesis that the founding dispersals of language families coincided with the spread of agriculture in a volume edited by Bellwood and Renfrew, two staunch proponents of this very hypothesis (van Driem 2002). Quite to the contrary, the spread of Indo-European furnishes the most obvious counter-example. Likewise, Sumerian, Elamite, Akkadian, Hurrian, Hattic and other languages which have left no surviving linguistic descendants were the tongues spoken by early agricultural civilisations, which therefore bear witness to the permeability of linguistic boundaries for the dissemination of agriculture.
The Neolithic and Bronze Age of Asia Minor and Mesopotamia are characterised by a very long period of incursive population movements into, rather than out of, Anatolia and the Fertile Crescent, driven or lured by the relative affluence of urban centres supported by agricultural surplus. Not just Indo-Europeans such as Hittites and Mitanni were drawn in by the good life. Gutaeans, Amorites, Kassites and other peoples likewise came to settle in the Fertile Crescent and Anatolia. Toponymical evidence and details about the cults of certain deities have been used to argue that even the Sumerians originally migrated from an earlier northern homeland to lower Mesopotamia, where they adopted agriculture from a resident population, at which time the Sumerians borrowed agricultural terms such as agar ‘field’, apin ‘seeder plough’ and apsin ‘furrow’ from a substrate language (Landsberger 1943, 1944, 1945, 1974; cf. Rubio 1999).
Similarly, it is likely that the Trans-Himalayans who introduced their pre-Sinitic language to the Yellow River basin first came as migrants in search of the good life to the affluent agriculture societies on the North China plain. It is therefore not inconceivable that the Yellow River basin, where Setaria and Panicum began to be domesticated about nine millennia ago, could have been the original Altaic homeland. This primary homeland may have been abandoned after an ancient Trans-Himalayan population introduced themselves and their Proto-Sinitic language to the early inhabitants of the North China plain, after which the secondary homeland of Trans-Eurasian may have moved north towards the Amur river basin.
The Father Tongue correlation can likewise not explain everything. If the paternal lineage C2 (M217) is correlated with Altaic linguistic affinity, as appears to be the case for Turkic, Mongolic and Tungusic, then Japanese is no Father Tongue, and neither is Korean. This Y-chromosomal haplogroup accounts for 11% of Korean paternal lineages, and the frequency of the lineage is even more reduced in Japan. Yet this molecular marker may still be a tracer for the introduction of Altaic language to the archipelago, where the paternal lineage has persisted, albeit in a frequency of just 6%.
On the other hand, the Y-chromosomal haplogroup D1a2 (M55) appears to be correlated with the ancient linguistic phylum of which Ainu is the surviving remnant. Therefore, Ainu is a father tongue, and the ancient paternal lineage D1a2 (M55) also remains robustly present elsewhere in Japan. If the bearers of the paternal lineage O2b introduced Yayoi culture and wet rice cultivation to the Japanese archipelago, then agriculture and superior metallurgy appear to have contributed to the fecundity of this paternal lineage, a veritable agricultural spread but without language.
Overly simple approaches that turn up correlations where direct causation is lacking or unlikely have come to enjoy perennial popularity (e.g. Nettle 1998; Gorenflo et al. 2012; Axelsen and Manrubia 2014; Greenhill et al. 2018; Hua et al. 2019). The Farmer Language Dispersal model is no doubt superior to such approaches in seeking to understand the dispersal of language families in terms of the spread of a subsistence strategy that has changed the face of our planet. However, enthused scholars oblivious to the faulty nature of the reasoning underlying the Farmer Language Dispersal model are inclined to seek corroboration and reinterpret evidence in ways that ‘fit’ that model. Similarly, the false assumption that any widely observed phenomenon, such as the Father Tongue correlation must therefore be universal, a presumption which we have repeatedly disavowed in print, would likewise prevent us from discerning the contours of a more complex picture of the past and render us unable to find the right fit for the plentiful pieces of the puzzle of prehistory.
|
|
|
|
Post by Admin on Jan 31, 2023 18:15:30 GMT
A genetic history of migration, diversification, and admixture in Asia
Melinda A. Yang
Department of Biology, University of Richmond, Richmond, VA, USA
Academic Editor(s): Guido Barbujani, Lounès Chikhi
Received: Jun 9, 2021 | Accepted: Sep 14, 2021 | Published: Jan 6, 2022
Abstract
L.L. Cavalli-Sforza spearheaded early efforts to study the genetic history of humans, recognizing the importance of sampling diverse populations worldwide. He supported research on human evolutionary genetics in Asia, with research on human dispersal into Asia and genetic distances between present-day East Asians in the late 20th century. Since then, great strides have been made in understanding the genetic history of humans in Asia, through large-scale genomic sequencing of present-day humans and targeted sequencing of DNA from ancient humans. In this review, I survey the genetic prehistory of humans in Asia, based on research using sequence data from humans who lived in Asia as early as 45,000 years ago. Genetic studies comparing present-day Australasians and Asians show that they likely derived from a single dispersal out of Africa, rapidly differentiating into three main lineages: one that persists partially in South Asia, one that is primarily found today in Australasia, and one that is widely represented across Siberia, East Asia, and Southeast Asia. Studies of ancient DNA from human remains in Asia dating from as far back as 45,000 years has greatly increased our understanding of the population dynamics leading to the current Asian populations.
Based on “Jin L, Underhill PA, Doctor V, Davis RW, Shen P, Cavalli-Sforza LL, Oefner PJ. Distribution of haplotypes from a chromosome 21 region distinguishes multiple prehistoric human migrations. Proc Natl Acad Sci U S A. 1999;96(7):3796-3800”.
Keywords
human migration, admixture, genetic structure, Australasians, Asians
|
|
|
|
Post by Admin on Feb 1, 2023 19:34:06 GMT
Introduction In the mid to late 20th century, L.L. Cavalli-Sforza helped to initiate the field of ‘genetical demography’ of humans [1], which sought to use genetic patterns to elucidate our past population history. He recognized that quantitative analysis of genetic data could add a unique dimension to human history and was a major advocate for developing a diverse set of human genetic samples for public research, as exemplified by the Human Genome Diversity Project [2]. His collaborations with researchers as varied as linguists and archaeologists proved that he was also a strong proponent for interdisciplinary research [3,4]. Cavalli-Sforza recognized the importance of studying the genetic history of humans in Asia to understand the dispersal of modern humans. One early example of this comes from a study conducted by him and his student Li Jin, among others, in 1999 [5]. They examined the distribution of a contiguous segment (i.e., a haplotype) of chromosome 21 across present-day populations worldwide. They found that after African populations, Oceanians showed the next highest level of haplotype diversity, and that the haplotype patterns of Oceanians were different from those found in East Asians. Based on these results, they concluded that there had been at least three distinct migrations, one to Oceania, one to Asia (and the Americas), and one to Europe. This early study demonstrated the importance of Asia in understanding human genetic history. Asia is the largest inhabited continent in the world, home to nearly 60% of all humans with high ethnic diversity. While most of Cavalli-Sforza’s studies focused on the genetic history of humans in Europe [3,4], he also contributed to and supported research on human genetics in Asia. He trained and worked with East Asian scholars to examine the association of genetic distances and surnames [6,7] in the 1980s-1990s, and he spoke highly of human genetic research in Asia, as demonstrated by a commentary he published on the earliest large-scale human genetic project from the region: the Chinese Human Genome Diversity Project [8]. Major contributions from Asian researchers in the growing field of human evolutionary genetics led to efforts such as the Human Genome Organization (HUGO) Pan-Asian SNP (single nucleotide polymorphism) Consortium [9,10], increasing our understanding of modern human genetic diversity in Asia. Such programs were the precursors of the broad genomic sequencing efforts seen today, such as the GenomeAsia 100K (GA100K) project, which has sequenced DNA from 1,739 individuals from 64 Asian countries [11]. In many ways, the GA100K Project is a direct product of the vision Cavalli-Sforza had of cataloging worldwide human genetic variations. In the three decades since Cavalli-Sforza’s initial call for the HGDP, the great improvements in sequencing techniques have allowed the creation of vast repositories of genome-wide data from diverse human populations, and thousands of present-day human genomes [9,11–16] have been sequenced and analyzed. In addition, improved techniques for sequencing small amounts of DNA, preventing contamination, and compensating for post-mortem DNA damage [17–20] have increased access to DNA data from ancient humans, both archaic (Neanderthals and Denisovans) and modern (H. sapiens), allowing an unprecedented exploration of human genetic history [21–23]. The aim of this review is to understand the modern human dispersal in Asia over the past 45,000 years by examining genomic data from ancient and present-day modern humans. Our understanding of modern human expansion into Asia has greatly increased in scale and resolution because of the successful sequencing of ancient human DNA (Figure 1A). I will focus on findings that have increased our understanding of (1) the initial migration of modern humans into Asia, (2) the movement and interaction of humans in Upper Paleolithic Asia, and (3) the rapid dispersal of humans within Asia in the past 10,000 years. 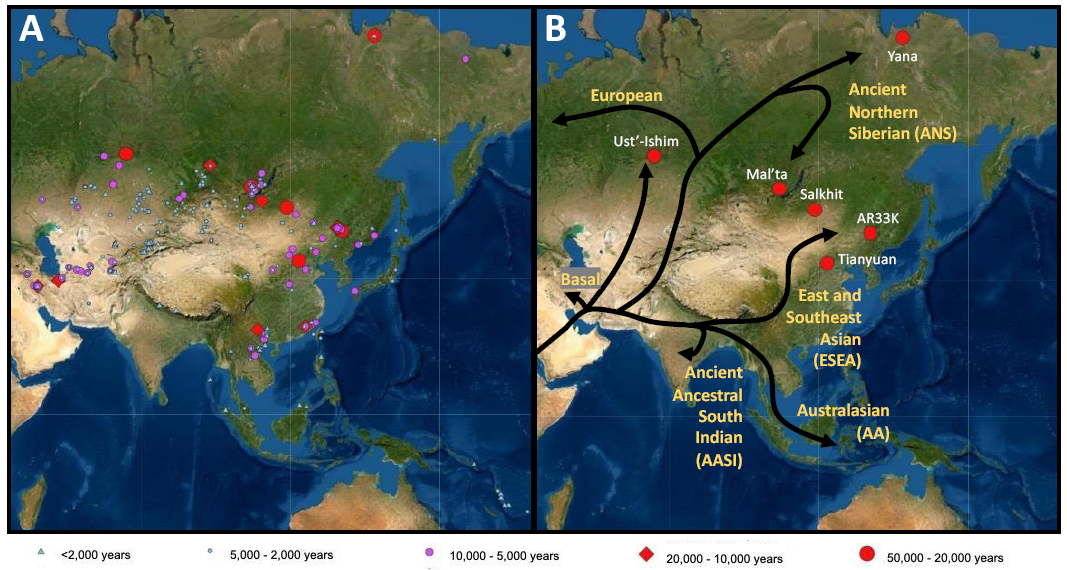 Figure 1 Maps indicating the location of ancient DNA samples from Asia and the patterns of genetic differentiation in Asia in the Early Upper Paleolithic. (A) Location of ancient individuals from Asia and Australasia who have been sequenced to date. Symbols refer to the age of individuals sequenced from that archaeological site, with the key found at the bottom. Red symbols date to 50,000–10,000 years ago (Upper Paleolithic), purple and blue symbols are younger than 10,000 years. (B) Differentiation after dispersal out of Africa in the Early Upper Paleolithic (45,000–20,000 years ago), with labeled lineages or ancestries in yellow next to the associated branch. The tree diagram shows divergence patterns and is not meant to depict migration routes from the branches or geographic origins of ancestral populations from the internal nodes. The branches predominantly associated with present-day Asian populations include the Ancient Ancestral South Indian (AASI) lineage, Australasian (AA) lineage, and East and Southeast Asian (ESEA) lineage. White labels refer to specific archaeological sites dating to the Early Upper Paleolithic. |
|
|
|
Post by Admin on Feb 3, 2023 19:51:49 GMT
Initial migration of modern humans into Asia and Australasia
An out-of-Africa model of human dispersal has been well supported by genetic studies, as discussed in [21,24,25], though later sequencing of archaic humans has demonstrated a small but notable contribution from archaic humans into early modern humans [26–28]. Later genetic studies also established that separation from African populations likely occurred 65,000–45,000 years ago [15,29]. Beginning at the turn of the 21st century, genetic studies began to address human dispersal out of Africa, such as the number of dispersals and the shape of human expansion in Eurasia and beyond.
Two models of dispersal have been proposed regarding migration into Asia and Australasia (the region consisting of Australia, New Zealand, and neighboring South Pacific islands). The single dispersal model describes a single migration out of Africa into Eurasia, with present-day Australasians deriving from an early offshoot of the Asian lineage. The multiple dispersal model posits several dispersals out of Africa by modern humans, with Australasians deriving from an earlier dispersal separate from the one contributing to mainland Asians and Europeans. Jin, Cavalli-Sforza, and others found high levels of genetic diversity in present-day populations from Oceania, and they used this finding to argue for a distinct out of Africa migration into Australasia [5], consistent with a multiple dispersal model. A phylogenetic analysis including 55,000 genome-wide single nucleotide polymorphisms (SNPs) across Asia by the HUGO Pan-Asian SNP Consortium found that East Asians and Australasians shared a closer genetic relationship to each other than to present-day Europeans [9], which they argued did not support a multiple dispersal model, but rather a single dispersal model.
In 2011, one of the first ancient DNA studies on a modern human was published, generating genome-wide data from a 100-year-old aboriginal Australian [30]. Comparison of the aboriginal Australian DNA to sequences from present-day Europeans (French) and Asians (Han Chinese) using a test of relative genetic similarity known as the D-statistic (bold text indicates methods or software described in Box 1) showed an excess of shared alleles between Europeans and Asians relative to Aboriginal Australians, supporting a multiple dispersal model. A complicating factor, however, was that Denisovans, early archaic hominins from Siberia [27,31,32] and the Tibetan Plateau [33,34], contributed up to 5% of their ancestry to populations living in Southeast Asia and Australasia island nations [27,32,35]. Because much smaller proportions of Denisovan ancestry were found in mainland Asian populations (0.05%-2%), the deep divergences observed genetically were potentially explained by archaic admixture rather than an earlier dispersal of modern humans [11,35–38].
Several studies have examined the relationship of Australasian populations to other modern humans after accounting for archaic ancestry from Neanderthals and Denisovans. After sequencing Denisovan DNA to 30-fold coverage, Meyer et al. estimated a maximum likelihood tree allowing admixture (Treemix, Box 1) and found that Papuans and East Asians were grouped together relative to Europeans [32]. Mallick et al. found that a well-fitting admixture graph (qpGraph, Box 1) grouped Papuans, Australians, and the Andamanese Onge with East Asians, with additional Denisovan admixture into Papuans and Australians [15]. Andamanese islanders such as the Onge do not show high Denisovan admixture, so Mondal et al. [39] compared whole-genome sequences from ten Andamanese individuals to other present-day populations. Using both Treemix and D-statistic analyses, they found that the Andamanese shared a close genetic relationship to mainland Indians, East Asians, and other Australasians [39]. Malaspinas et al. [40] examined the likelihood of a single dispersal model versus a multiple dispersal model by comparing the observed joint frequency spectrum to the expected joint frequency spectrum using fastsimcoal2 (Box 1). They found that accounting for Denisovan admixture led to better support for the single dispersal model, while excluding Denisovan admixture led to better support for the multiple dispersal model [40]. Collectively, these studies showed that after accounting for archaic admixture from Neanderthals and Denisovans, Australasians consistently grouped closely with mainland Asian populations, supporting a single dispersal model.
While a closer relationship between Australasians and Asians than Europeans was broadly supported, some scholars still argued for a small but notable contribution to Australasians from an earlier dispersal of modern humans out of Africa. Pagani et al. used MSMC split time estimates (Box 1) to argue that Papuans separated from African populations earlier than mainland Eurasian populations [56]. Assuming 30 years per generation and a mutation rate of 1.25 x 10-8 per generation per site, they estimated a 90,000-year split time between Papuans and Yorubans and a 75,000-year split time between mainland Eurasians and Yorubans. Using fineSTRUCTURE (Box 1), they examined haplotypes in Papuans associated with a deeper divergence and found that 2% of the haplotypes in Papuan genomes could not be explained by Denisovan admixture or shared origins with mainland Eurasians, and thus were best explained as originating from an earlier dispersal of modern humans out of Africa [56]. Though an earlier dispersal may be partially represented in the genomes of Australasians, the main pattern observed in their genomes indicates a shared evolutionary history with populations widespread today in much of the eastern regions of Asia.
|
|
|
|
Post by Admin on Feb 5, 2023 21:16:31 GMT
Defining Asian Ancestries
In the rest of this review, I examine genetic findings related to past humans from Asia, often using the term ‘ancestry’ to communicate key interpretations of genetic patterns. Ancestry is a useful concept for conveying relationships in evolutionary genetics, but imprecise application of the term can result in racial categorizations that unintentionally bolster misleading claims of the biological reality of race [57]. Mathieson and Scally [58] recently published a nuanced exploration of how the term was used and misconstrued in human evolutionary genetics, a portion of which is summarized here to clearly define how the term ancestry is used in this review.
As described by Mathieson and Scally [58], a technical definition of genetic ancestry refers to the set of paths across an individual’s ancestors through which a DNA segment was inherited. For a population, the set of genetic ancestries across all individuals is highly complex, so for practical reasons, researchers focus on summarizing the general demographic relationships observed between populations. This concept of ‘population ancestry’ then assumes that a population can be represented as a mixture of different source populations, denoted here as ancestries, dependent on the pattern of variation across the genome.
My objective is to articulate key human ancestries of Asia that have been described in the literature to date, inferred from findings of high genetic similarity among ancient and present-day populations. When a described ancestry has only been associated with current populations or is only known through partial representation in an ancient individual or present-day population, I use the term lineage rather than ancestry to indicate descent from a hypothetical ancestor that has yet to be sampled. As Mathieson and Scally note, all source populations, or ancestries, are constructs that (1) may be only distantly related to the associated sampled individuals; (2) may not actually be represented in the available samples (‘ghost populations’); and (3) create discrete sources for a population when discrete categories are not actually applicable [58]. The ancestries and lineages referred to throughout this review (first appearance bold italic, with description in Box 2) are theoretical constructs that have been associated with samples from different past or present populations, and they cannot be described as physical ancestors of any present-day individuals.
Box 2 Major lineages and ancestries described in the text.
Lineages found in Asia and Australasia that contributed to ancestries that would shape present-day Asians and Australasians
Ancient Ancestral South Indian (AASI) lineage—this lineage refers to an ancestral population that primarily contributed to humans living in South Asia, particularly southern India. Partially represented in 5,000–1,500-year-old individuals from in or near the Indus Periphery and present-day Indians [59,60].
Australasian (AA) lineage—this lineage refers to the ancestral population that primarily contributed to human populations in Australasia, or the region consisting of Australia, New Zealand, and neighboring islands in the South Pacific Ocean. Represented primarily by present-day Australasians, e.g. Papuans and Aboriginal Australians.
East and Southeast Asian (ESEA) lineage—this lineage refers to an ancestral population that primarily contributed to humans living in mainland East and Southeast Asia. Represented primarily by present-day East and Southeast Asians, e.g. Han and Kinh.
Ancestries in the ESEA lineage
Amur ancestry—ancestry associated with populations in the Amur River region, Mongolia, and Siberia, with the oldest individual sampled to date represented by a 14,000-year-old individual from the Amur River region, i.e. Amur14K [61]. Populations associated with this ancestry likely contributed to the ancestors of Native Americans and populations associated with Paleosiberian ancestry.
Fujian ancestry—ancestry associated with populations in the Fujian region of coastal, southern China from 12,000–4,000 years ago. The oldest individual sampled to date is an individual from Fujian, i.e. Qihe3 [62]. Populations associated with this ancestry contributed to Austronesian speakers today.
Guangxi ancestry—ancestry associated with a 10,500-year-old individual from Guangxi, i.e. Longlin [62]. Populations associated with this ancestry are partially observed in 8,000–6,000-year-old hunter-gatherers from Guangxi and are not observed in historical Guangxi individuals and present-day East and Southeast Asians.
Hòabìnhian ancestry—ancestry on the ESEA lineage associated with 8,000–4,000-year-old hunter-gatherers [63] associated with the Hòabìnhian culture in Laos and Malaysia. This ancestry is deeply diverged from the common ancestor of present-day East and Southeast Asians and Tianyuan ancestry.
Jōmon ancestry—ancestry associated with 8,000–3,000-year-old individuals in the Japanese archipelago. The oldest individual sampled to date is Higashimyo, from Kyushu, Japan [64]. Populations associated with this ancestry contributed partially to present-day Japanese populations.
Tianyuan ancestry—ancestry on the ESEA lineage associated with Upper Paleolithic individuals dating to 40,000–33,000 years ago in northern China and Mongolia, i.e. Tianyuan, Salkhit, and AR33K [61,65,66]. This ancestry is deeply diverged from the common ancestor of present-day East and Southeast Asians and Tianyuan ancestry.
Tibetan ancestry—ancestry associated with 3,000–1,000-year-old individuals in the Himalayan region of the Tibetan Plateau [67]. Populations associated with this ancestry contributed to present-day Tibetan and Sherpa populations.
Yellow River ancestry—ancestry associated with populations in the Yellow River region, with the oldest individual sampled to date represented by a 9,500-year-old individual from the lower reaches of the Yellow River in Shandong, i.e. Bianbian [68]. Populations associated with this ancestry greatly impacted most present-day East and Southeast Asians.
Ancestries not deriving completely from the AA, AASI, or ESEA lineages
Ancient Northern Siberian (ANS) ancestry—ancestry associated with 33,000-year-old individuals from near the Yana River in northern Siberia [69]. 24,000 and 17,000-year-old individuals from the Lake Baikal region of Siberia [51], associated with Ancient North Eurasian ancestry, derive from a lineage associated with ANS ancestry. This ancestry is more closely related to ancestry found in present-day Europeans rather than ancestry found in present-day East and Southeast Asians.
Basal ancestries—ancestries associated with populations that diverged very early in the dispersal out of Africa. At least three distinct ancestries have been reported, including one related to a 45,000-year-old individual from Ust’-Ishim Cave in western Siberia [70], and two related 12,000–7,000-year-old individuals from the Levant and Iran, who are some of the earliest agriculturalists sampled to date [71]. Populations associated with Basal Iranian ancestry contributed to populations in Central and South Asia.
Indus Periphery (IP) ancestry—ancestry associated with 5,000–4,000-year-old individuals from near the Indus Valley [59,60]. This ancestry is a mixture of ancestry related to the AASI lineage and Basal Iranian ancestry, and populations associated with this ancestry contributed to populations in South Asia.
Native American ancestry—ancestry associated with present-day Native Americans. This ancestry is a mixture of ancestries associated with Amur ancestry and ANS ancestry, and it is closely related to Paleosiberian ancestry. The oldest sequenced individual associated with this ancestry is an 11,500-year-old individual from Upward Sun River, Alaska [72].
Paleosiberian ancestry—ancestry associated with 14,000–10,000-year-old individuals from the Lake Baikal region of Siberia (Ust-Kyakhta-3) and Far East Siberia (Kolyma1) [69,73]. This ancestry is a mixture of ancestry related to ANS and Amur ancestries, and it is closely related to ancestry associated with Native Americans.
Steppe ancestry—ancestry associated with 5,000–3,500-year-old individuals from the central and western steppe, related to the Yamnaya and Afanasievo cultures [42,74]. This ancestry is a mixture of ancestries related to populations associated with ANS ancestry, basal ancestry from the Levant and Iran, and ancestry related to hunter-gatherers from the Caucasus regions of eastern Europe.
|
|

