|
|
Post by Admin on May 7, 2022 17:24:58 GMT
Whole genome composition and genetic ancestry Balatonkeresztúr site samples According to the principal component analysis (PCA) based on 590k nuclear SNPs (Fig. 3 a.) Bk-I is clearly separated from Bk-II and Bk-III, where Bk-II has a strong shift towards HG samples23 overlapping with only a fraction of known archaic samples23 and Bk-III. Admixture and qpAdm analyses for assessing genetic components (Supplementary Tables S9, S12-16; Supplementary Information sections 5.2 and 5.5.2) revealed ~17% HG, ~40% European farmer, and ~43% steppe ancestry for Bk-I, similar to average Bronze Age Europeans. Bk-I is most likely derived from a single source that is genetically related to a Poland Southeast Bell Beaker culture (BBC) associated population (p = 0.784) in line with archaeological observations24. Bk-II comprises a unique makeup of ~42% HG, ~41% European farmer, and ~17% steppe ancestries. qpAdm analysis revealed most plausible sources of Neolithic Sweden Funnel Beaker culture (~32±8%), Poland Southeast BBC (~41±6%) and an extra HG (~29±3%) ancestry of yet unknown origin. Despite the lower supported models Bk-I can not be excluded as an ancestry component for Bk-II, while the affinity of Sweden Funnel Beaker culture associated population likely reflects a more closely related group, such as population related to the Poland Globular Amphora culture, see Supplementary Information section 5.5.2.2 and Supplementary Tables S12 and S15. Bk-III shows a shift in ancestry composition (~29% HG, ~46% European farmer, ~25% steppe). qpAdm analyses revealed that the main ancestry component for Bk-III is Bk-II (~53±5%), while “dilution” of Bk-II to Bk-III is mostly driven by population events that are yet to be uncovered.  Fig. 3 a) Principal Component Analysis based on 590k SNPs calculated by the smartpca software. Bk-II clearly separated from any known archaic Central-Eastern European populations. b) Highlighted genetic outliers from previous studies (Supplementary Information section 5) that show a similar hunter-gatherer (HG) ancestry component to Bk-II among ancient datasets. The origin of the HG component likely lies between the Carpathian mountains and the Dnieper River or the Black Sea, from where it expanded further North and from where the Bk-II population most likely originated from. Outliers signalise a westward migration route of this component detected in Bk-II. Genetic outliers from previous studies Many samples were defined as genetic outliers in their genetic context by previous studies from Bronze Age Europe. We selected such outlier individuals with high HG ancestry components to assess whether they are related to Bk-II. Selection was based on previous observations and also by using Dixon’s Q-test25 at 90% confidence interval on HG component’s upper deviation using results of the Admixture analysis. First, we ran f4-statistics in the form of f4(W=test outlier, X=corresponding population, Y=Bk-II, Z=Yoruba)26. This test resulted in positive values for some outliers (W) meaning that these are genetically closer to Bk-II (Y) than its presumed population (X). However, Z-scores are low in many cases, and false positives may appear solely by high HG component, not by true relationship (see Supplementary Table S10 and Supplementary Information section 5.3). To check true relationship between Bk-II and groups/samples with high HG ancestry, we performed an outgroup f3-statistics in the form of f3(X=Bk-II, Y=test HG-s, Z=Yoruba) for all relevant archaic populations and outliers23 (Supplementary Table S11) that resulted a table of allele frequency based distances between test pairs Xs and Ys. Euclidean clustering based on the results of f3-statistics revealed that a number of samples and even three populations from the Baltic (Fig. 3 b, Supplementary Information section 5.4) from AADR23 form a cluster with Bk-II and Bk-III, suggesting actual genetic connection via a common HG ancestry source. |
|
|
|
Post by Admin on May 7, 2022 20:25:04 GMT
Discussion
The Carpathian Basin was inhabited by the Baden cultures’ population at the end of the Copper Age, and their genetic composition was represented by an early farmer and a slightly increased HG genetic component, compared to the previous Neolithic populations of the region6. Here we demonstrated that in the early phase of this culture, a group of Western European origin appeared in Transdanubia, diversifying what we knew about the region’s Late Copper Age substrate up to now.
The Carpathian Basin experienced the influx of steppe-related genetic ancestry at the dawn of the Bronze Age5,8, and this transformation was already detectable at Balatonkeresztúr-Réti-dűlő site as well, where we could examine multiple populations. The earliest Bronze Age horizon Bk-I (representative of the Somogyvár-Vinkovci culture) shows similarities to Poland Southeast BBC associated population with high steppe ancestry that was replaced by the Kisapostag culture associated Bk-II around the 23-22th century BCE, while at least some sort of genetic ancestry of Bk-I in this population can not be excluded. According to our results, the Bk-II population had outstandingly high HG genetic ancestry levels, compared to other Bronze Age groups of the region, which can be traced back to today’s Ukraine, Belarus, Moldavia or Romania, pointing to a long standing previously unsampled population with dominant HG ancestry. Calculated admixture dates10 suggest the presence of a genetically pure or at least highly HG specific population in Eastern Europe as late as the end of the Copper Age. Part of this group subsequently admixed with populations of mainly steppe (likely Poland Southeast BBC) and early farmer (most likely a Globular Amphora culture related) ancestry during their westward migration on a Northern route, leaving genetic traces in Corded Ware culture, BBC, and other Bronze Age populations. The paternal lineage of BK-II was likely linked to the farmer component, as I2a-M223 (upstream to I2a-L1229) was a frequent paternal lineage among Globular Amphora culture and related populations. Looking for the possible source areas of the Kisapostag culture, a number of archaeological theories need to be considered. The pottery decoration technique originated either from Corded Ware in the Middle Dnieper region (Ukraine), epi-Corded Ware groups (northern Carpathians), e.g. Chłopice-Veselé or Nitra groups (Slovakia), the latter two is also supported by inhumation practises31–36. However, connections with the Litzenkeramik or Guntramsdorf-Drassburg group (eastern Austria, Slovenia, western Croatia) were also raised37,38. Pottery forms were connected to local development of communities with eastern (Makó–Kosihy–Čaka) or southern (Somogyvár–Vinkovci) origins, too39. BBC influence was also mentioned based on connections of pottery and craniometry data (so called Glockenbecher or brachycran skull type40–42). The results of this study fit best with the Middle Dnieper area origin of BK-II, especially when we consider individual I4110 from Dereivka I (Ukraine Eneolithic) as one of the earliest representatives of their genomic makeup.
Strontium isotope (87Sr/86Sr ratio) data, representing through nutrition the bioavailable Sr in the area where people lived in a certain age interval, shows local values for both sexes in both Bk-II and Bk-III. These results push back the timing of their arrival a few generations, meaning that local or southern impact of cultural traits and maybe even genetic admixtures likely occurred during this short period as well, which also could explain the culture’s archaeological heterogeneity.
The population of Bk-III was the direct descendant of Bk-II, forming not just cultural (Encrusted pottery) but also genetic continuity for at least ~500 years, even if the radiocarbon sequences allow a few decades of hiatus at the studied site. Continuous female-biased admixture with various groups occurred during this period according to our and previous genetic10 and archaeological31,43 evidence, diluting the BK-II genetic ancestry.
In both periods, the homogeneity of paternal lineages suggest a similar social organisation described in9,10 of a patrilocal residence system. However, strontium isotope data shows local values for both sexes, which along with similar genomic makeup of females and males suggest exogamy most probably between villages of the same population. Two pairs of half-sibling graves in the two periods may indicate polygamy, although remarriage for high female mortality is more plausible. Notably, almost none of the uniparental markers are identical even at the haplogroup level with individuals from the Croatian Encrusted Pottery culture Jagodnjak site, despite high similarities in cultural traits, social structure and genomic composition of the communities. This points to clan-like or patriarchal superfamily structure of Kisapostag and Encrusted Pottery groups. The relatively limited presence of female and children burials in both Bk-II and Bk-III periods may suggest distinctive treatment or another (here undiscovered) burial group for women and children at the same site. Although, in other Bronze Age cemeteries, e.g. Ordacsehi and Bonyhád in Hungary, males, females and children were buried close to each other, suggesting high variance of burial practises34,35,44.
While low genomic coverage did not allow fine SNP recovery, we did find evidence for malignant variants within all of our tested groups, and undoubtedly showed the presence of LHON and Jacob’s syndrome within Bk-II. Additionally, the disease panel we created can be extended and used in future studies, providing insight into past population health qualities.
Considering the unstructured age and kinship distribution in the mass grave Bk-III compared to Bk-II, the coetaneous death of eight people at least, the absence of traumatic or ritual events on bones, and non-cremated nature of the burial all signals a sudden tragic event in the Encrusted pottery period, most likely an epidemic, as first suggested based on the anthropological analyses45. Interestingly, comparative strontium isotope analyses on the first and third molar of the individuals in the BK-III mass grave indicate that subadult males - including a severely disabled individual (S15) with hip dysplasia - left their community for a while and then returned to their birthplace prior to their death, raising further questions for future studies on prehistoric lifeways and social organisations.
|
|
|
|
Post by Admin on May 8, 2022 1:28:31 GMT
Materials and Methods
Isotope analyses
Radiocarbon dating was performed at the HEKAL AMS C-14 facility of the Institute for Nuclear Research in Debrecen, Hungary (see Supplementary Information section 1.8). 87Sr/86Sr isotope measurements were performed in the ICER Centre, Institute for Nuclear Research Debrecen, Hungary and at Quinnipiac and Yale University, Connecticut, USA (see Supplementary Information section 1.9).
Ancient DNA laboratory work
Petrous bones and teeth were taken from skulls for genetic investigation (Supplementary Table S1). Laboratory work was performed in a dedicated ancient DNA laboratory facility (Institute of Archaeogenomics, Research Centre for the Humanities, Eötvös Loránd Research Network, Budapest, Hungary). Each step was carried out in separate rooms under sterile conditions, during work protective clothing was used. Irradiated UV-C light, DNA-ExitusPlus™ (AppliChem) and/or bleach were applied for cleaning after and between work stages, and also, blank controls were utilised at all times.
Sample surfaces were cleaned by sandblasting and mechanically milled to powder. DNA extraction was performed according to Dabney et al. 201346 with minor changes according to Lipson et al. 20176. DNA extraction success was verified by PCR using mtDNA primer pairs (F16209-R06348; F16045-R06240). Half-UDG treated libraries were used according to Rohland et al. 201547 with minor changes. Unique double internal barcode combinations were used for each library (Supplementary Table S1). Libraries were amplified with TwistAmp Basic (Twist DX Ltd) and purified with AMPure XP beads (Agilent). Then, concentration measurements were taken on Qubit 2.0 fluorometer, fragment sizes were checked on Agilent 4200 TapeStation System (Agilent High Sensitivity D1000 ScreenTape Assay).
Hybridisation capture method for mtDNA and 3k nuclear SNP was used besides whole genome shotgun, as described by Haak et al. 2015, Lipson et al. 2017 and Csáky et al. 20204,6,48. Bait production was based on Fu et al. 20161 and N. Rohland’s personal communication, the oligos as a pool was ordered from CustomArray Inc. Both for shotgun and capture libraries, universal iP5 and unique iP7 indexes were used.
Sequencing was done on Illumina MiSeq and NovaSeq platforms with custom setup and 150, 200 and 300 cycles, respectively.
Additionally, we investigated Y chromosome STR profiles (17 markers) with AmpFISTR® Yfiler® PCR Amplification Kit (Applied Biosystems), having one blank and one positive control at each reaction preparation. The workflow followed the recommended protocol except the PCR cycles were increased from 30 to 34 and reactions were halved in volume. Two repeats were done where at least 4 markers yielded results. Data analyses were carried out in GeneMapper® ID Software v3.2.1 (Applied Biosystems), results are summarised in Supplementary Table S3.
Bioinformatic analyses
Illumina sequencing reads were processed by the PAPline, for details, see Supplementary Information section 6. We used the GRCH37.p13 reference sequence for calling pseudohaploid genomes. For kinship inferences we applied the READ software49 and a custom script (named MPMR, see Supplementary Information section 2.3 and Supplementary Table S2). MtDNA analyses included phylogenetic analyses using the MrBayes v3.2.650 and the BEAST v1.10.451 softwares and diversity tests using the Popgenome52 R package, see Supplementary Information section 2.1. For Y chromosome haplogroup determination the Yleaf v153 software was applied. For network analysis of STR data we used Network v10.1.0.0 and Network publisher v2.1.2.554,55, see Supplementary Information section 2.2. Due to low genomic coverages (<10,000 SNPs) we discarded individuals S2, S5 and S17 from the population genetic analyses. PCA was made by the Eigensoft smartpca software56 using the Human Origins Panel SNP set26, for other analyses the 1240k array12 was used for SNP call, for results, see Supplementary Table S7. Individuals S4, S5, S6 and S20 were discarded from further tests for them being first degree relatives of other samples. For investigating ancestry estimates we used supervised admixture analysis calculated by the ADMIXTURE v1.3.0 software57. The results were visualised by custom R scripts. f-statistics and qpAdm were performed using the admixr v0.9.158 and the admixtools v2.0.026 R packages.
|
|
|
|
Post by Admin on Jun 6, 2022 19:03:24 GMT
Y-chromosomal connection between Hungarians and geographically distant populations of the Ural Mountain region and West Siberia Helen Post, Endre Németh, László Klima, Rodrigo Flores, Tibor Fehér, Attila Türk, Gábor Székely, Hovhannes Sahakyan, Mayukh Mondal, Francesco Montinaro, Monika Karmin, Lauri Saag, Bayazit Yunusbayev, Elza K. Khusnutdinova, Ene Metspalu, Richard Villems, Kristiina Tambets & Siiri Rootsi Scientific Reports volume 9, Article number: 7786 (2019) Abstract Hungarians who live in Central Europe today are one of the westernmost Uralic speakers. Despite of the proposed Volga-Ural/West Siberian roots of the Hungarian language, the present-day Hungarian gene pool is highly similar to that of the surrounding Indo-European speaking populations. However, a limited portion of specific Y-chromosomal lineages from haplogroup N, sometimes associated with the spread of Uralic languages, link modern Hungarians with populations living close to the Ural Mountain range on the border of Europe and Asia. Here we investigate the paternal genetic connection between these spatially separated populations. We reconstruct the phylogeny of N3a4-Z1936 clade by using 33 high-coverage Y-chromosomal sequences and estimate the coalescent times of its sub-clades. We genotype close to 5000 samples from 46 Eurasian populations to show the presence of N3a4-B539 lineages among Hungarians and in the populations from Ural Mountain region, including Ob-Ugric-speakers from West Siberia who are geographically distant but linguistically closest to Hungarians. This sub-clade splits from its sister-branch N3a4-B535, frequent today among Northeast European Uralic speakers, 4000–5000 ya, which is in the time-frame of the proposed divergence of Ugric languages. Introduction The Uralic languages cover today a wide territory of North Eurasia from West Siberia in the east to Northeast Europe in the west. Hungarians with about 13 million speakers1 are the largest Uralic speaking group in the world2, who today reside in Central Europe (Fig. 1a) far apart from the rest of the members of their language family. Linguistically closest to Hungarians are geographically very distant West Siberian Mansi and Khanty (Fig. 1a), with whom they belong to the Ugric branch of the Uralic linguistic family2,3,4. In addition, the Hungarian language has been intensively influenced by several Turkic languages during the second half of the first millennium AD5. Agreeing with the linguistic results most of the archaeologists propose that the putative homeland of the ancestors of the Hungarian speaking population must have been in West Siberia6,7,8,9 (Fig. 1a). Figure 1 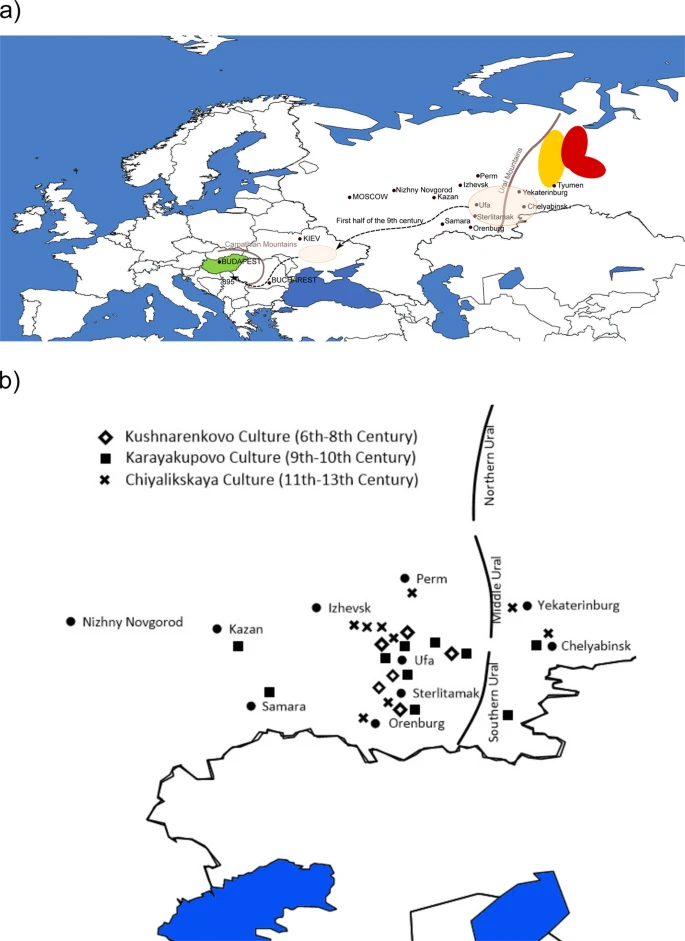 (a) Map of Western Eurasia and the putative migration route of early Hungarians based on archaeological data. Hungary is shown in green. Geographic locations of Mansis and Khantys are indicated with yellow and red, respectively. The pink transparent colour marks the geographic regions where archaeological evidence of Hungarian ancestors has been found. (b) Volga-Ural archaeological cultures from 6th to 13th century. Background maps from Surfer® (v.8, Golden Software, Inc, Golden, CO, USA). Despite of the eastern roots of the Hungarian language the present-day gene pool of Hungarians is very similar to the neighbouring non-Uralic speaking Central Europeans according to autosomal10,11, Y-chromosomal (chrY)12,13,14,15,16 and mitochondrial DNA (mtDNA) data13,17. One of the main results of the study by Tambets et al.10 was that the Hungarians differ from the majority of Uralic-speakers – they do not show any specific link with their linguistic relatives compared to their non-Uralic neighbours in Central Europe. These results are in line with the earlier observations of anthropologists who have suggested that since the forefathers of Hungarians arrived in the Carpathian Basin at the end of the 9th century6,18, the population has changed significantly during the demographic processes that homogenized different ethnic groups in the area within the last 1100 years19. Anthropological differences detected among ancient Hungarians from different macro-regions in the Carpathian Basin were interpreted as a possible reflection of their heterogeneous geographical origin and relatively recent admixture of ancient Hungarian groups20. Archaeogenetic studies also confirm the admixed genetic background of the early Hungarians. Comparing the Hungarian Conqueror mtDNA dataset to a large modern-day population dataset and archaeogenetic database, researchers found strong genetic affinities towards modern populations of Inner Asia, North and East Europe, Central Russia, and Late Bronze Age populations of the Baraba region, situated between the rivers Ob and Irtis21. Also, most researchers agree that the size of the Avar population, who resided in the Carpathian Basin in the 8–9th century AD, was unequivocally greater than the number of putative Hungarian ancestors22. It is notable, that the mtDNA gene pool of Avar “commoners” and mixed Avar-Slavic cemeteries showed significantly lower genetic distances toward medieval European populations than Asian populations, indicating that the genetic imprint of the Inner Asian Avar elite through their mtDNA was rather weak in those populations23,24. In addition, an ancient DNA study dealing with 6th century barbarian migrations shows that Y-chromosomes of 21 ancient individuals from a Szólád cemetery (Hungary) belong to predominantly Central and Southern European haplogroups (E, I1, I2a2, T, R1a and R1b)25. Considering the possibility of sometimes intensive migration to Carpathian Basin from the neighbouring regions26, it is reasonable that present-day gene pool of Hungarians has become very similar to neighbouring populations. However, it is an open issue if there is any trace in the recent Hungarian gene pool reflecting their possible homeland in the East. |
|
|
|
Post by Admin on Jun 7, 2022 19:17:04 GMT
Certain chr-Y lineages from haplogroup (hg) N have been proposed to be associated with the spread of Uralic languages27. So far, hg N3 has not been reported for Indo-European speaking populations in Central Europe14,28,29,30, but it is present among Hungarians, although the proportion of hg N in the paternal gene pool of present-day Hungarians is only marginal (up to 4%) compared to other Uralic speaking populations27. It has been shown earlier that one of the sub-clades of hg N – N3a4-Z1936 – could be a potential link between two Ugric speaking populations: the Hungarians and the Mansi31. It is also notable that some ancient Hungarian samples from the 9th and 10th century Carpathian Basin belonged to this hg N sub-clade32: Three Z1936 samples were found in the Upper-Tisza area (Karos II, Bodrogszerdahely/Streda nad Bodrogom) and two in the Middle-Tisza basin cemeteries (Nagykörű and Tiszakécske). The haplotype of the Nagykörű sample is identical with one contemporary Hungarian sample from Transylvania that tested positive for B545 marker downstream of N3a4-Z193632. Similar findings come from the maternal gene pool of historical Hungarians: the analyses of early medieval aDNA samples from Karos-Eperjesszög cemeteries revealed the presence of mtDNA hgs of East Asian provenance21.
Archaeogenetic studies based on mtDNA haplotypes have shown that ancient Hungarians were relatively close to contemporary Bashkirs33 who are a Turkic speaking population residing in the Volga-Ural region. Another study reported excessive identical-by-descent (IBD) genomic segments shared between the Ob-Ugric speaking Khantys and Bashkirs but a moderate IBD sharing between Turkic speaking Tatars and their neighbours including Bashkirs34. According to this study the gene pool of Bashkirs is a mixture of Turkic, Ugric and Indo-European contributions. The complexity of genetic composition of Bashkirs was shown by Yunusbaev et al.35, both in autosomal and in chrY data.
In this study we use, for the first time, the chrY high coverage sequencing data of Hungarian samples together with samples from the populations from regions of the Ural Mountains and West Siberia. We refine the phylogeny of hg N, estimate the divergence times of sub-clades of hg N3a4 and, together with a large genotyping dataset, reveal the spatio-temporal distribution pattern of hg N3a4. This lineage is the genetic link between present-day Hungarians and populations from the Ural region and West Siberia, the proposed region of origin for the Hungarian language.
|
|



