|
|
Post by Admin on Feb 21, 2017 20:17:13 GMT
 Figure 3. Map of genetic distances between modern-day populations of Eurasia and from aUzPo and aBOO. The hg distribution in the Mesolithic aUzPo population: U4 (37%), C (27%), U2e (18%), U5a (9%), and H (9%), indicated an ‘admixed’ composition of ‘European’ (U4, U2e, U5a and H, 73%) and ‘Central/East Siberian’ (C, 27%) hgs, based on the PCA plot (Figure 2). Interestingly, the population of aUzPo did not group with modern NEE populations, including Saami, but fell instead between the present-day ‘European’ and ‘Central/East Siberian’ clusters on the PCA graph, and more precisely between populations of the VUB (in light green) and West Siberia (in dark green). The high frequency of hg U4 is a feature shared between Mesolithic aUzPo, present-day VUB (Komi, Chuvashes, Mari), and West Siberian populations (Kets, Selkups, Mansi, Khants, Nenets), with the latter group also being characterized, like aUzPo, by the presence of hg C. The genetic affinity between Mesolithic aUzPo and present-day West Siberian populations could be visualized on the genetic distance map of North Eurasia (Figure 3A), on which locally lighter colorings indicated low values of genetic distances, and therefore an affinity between aUzPo and extant West Siberians. In order to test the potential population affinities formulated on the basis of the hg-frequency PCA and the distance map, we examined the present-day geographical distribution of the haplotypes found in aUzPo via haplotype sharing analyses (Figure 4). These analyses are less impacted by biases due to small population sizes or unidentified maternal relationships in ancient populations, and thus are less prone to artefacts. Although the highest percentages of shared haplotypes for aUzPo were observed in pools of West Siberian Khants/Mansi/Nenets/Selkups (2.8%), South Siberian Altaians/Khakhassians/Shors/Tofalars (2.2%) and Urals populations (Chuvash/Bashkirs, 2.0%), matches were widely distributed across Eurasia. This was consistent with the observation that most haplotypes sequenced in aUzPo were basal and hence, not informative in terms of geographical population affinity. Haplogroup-based analyses suggested that the genetic affinity between aUzPo and present-day West Siberians was partly due to the presence of hg C, implying that the non-basal haplotype C1 found in aUzPo (16189C-16223T-16298C-16325C-16327T, detected in three individuals) could be a clear genetic link with extant Siberian populations. However, the C1 haplotype found in aUz did not belong to hg C1a, the only C1 clade restricted to Asia (characterized by a transition at np 16356 [48]). Indeed, no exact match was found for the C1 haplotype in the comparative database of Eurasian populations (comprising 168,000 haplotypes), although 47 derivatives (showing one to three np differences) were found in extant populations broadly distributed throughout Eurasia (Table S1). Therefore, the C1 haplotype sequenced in aUzPo is currently uninformative about population affinity. In addition, all three aUzPo individuals showed identical C1 haplotypes, which meant that a close maternal kinship between these individuals could not be rejected. Biases due to the overestimation of the hg C1 frequency and small sample size of aUzPo may have led to an overestimation of the genetic affinity with modern-day West Siberians in the hg-based analyses. To account for this, we assumed a scenario of extreme maternal kinship, in which identical haplotypes found in several individuals at the same site (redundant haplotypes) were only counted once (Figure S2A). Under this scenario, the genetic affinity between aUzPo and present-day Western Siberians was less distinctly pronounced (Figure S2B).  Figure 4. Percentages of haplotypes from aUzPo and aBOO matched in modern-day Eurasian population pools. To further evaluate the apparent significant genetic discontinuity between aUzPo and modern extant populations of NEE and Saami, we analyzed Bayesian Serial SimCoal (BayeSSC) coalescent simulations [49] using Approximate Bayesian Computation (ABC, [50]) and tested whether discontinuity could be better explained by genetic drift or by migration. Models of genetic continuity between aUzPo and the present-day population of NEE or Saami (H0a) were compared to models in which genetic discontinuity between aUzPo and the extant population of NEE was introduced by migration (H1a, Figure 5). Ancestors of individuals from CE were selected as a source population for the migration on the basis of the PCA plot (Figure 2) showing that present-day populations of NEE shared the most genetic similarities with those of CE. The model of genetic discontinuity between aUzPo and present-day Saami was not tested since no source population for a potential migration could be identified from the PCA plot. The model of genetic continuity between aUzPo and present-day Saami was found to fit the observed data better than the model of genetic continuity between aUzPo and present-day NEE. This can be attributed to the low haplotypic diversities (0.74 and 0.81, respectively, in contrast to 0.98 for NEE; Table 2) of both aUzPo and Saami populations. The migration model provided a better fit for the genetic data than the model of genetic continuity (H0a), as indicated by a low Akaike Information Criterion (AIC, [51]) and a high Akaike weight ω [52]–[53]. The lowest AIC (Figure 5) and highest Akaike's ω (Table 3) were obtained for migration models, the best fit being obtained for the model involving 10% of migrants over the last 7,500 years (H1b; ω = 1.00E+0 as opposed to ω = 2.57E-7 for the continuity model H0a). Our analyses of coalescent simulations therefore supported a genetic discontinuity between aUzPo and the present-day population of NEE, which was better explained by a migration from CE than by genetic drift.  Figure 5. Graphical representation and Akaike Information Criterions of the demographic models compared by coalescent simulation analyses. The timeline indicates the age of populations in generations (G). For models H0a to H0e, genetic continuity is tested between combinations of ancient populations and present-day populations of North East Europe (NEE) or Saami (saa), as indicated in the column ‘P0’. For models H1a and H1b, genetic discontinuity between aUzPo or aBOO, and NEE is tested assuming a migration from Central Europe (CE). The percentage of migrants from the source population into the sink population (10%, 50% and 75%) is indicated in the column ‘%’. The cells containing Akaike Information Criterion (AIC) values were colored according to the gradient of AIC represented below the figure: from white for the highest value of AIC (worst model fit, 199.1 for H0b) to red for the lowest value of AIC (best model fit, 81.9 for H0a). |
|
|
|
Post by Admin on Feb 22, 2017 20:20:46 GMT
 Evidence for Siberian gene flow into far north-eastern Europe. At the 3,500 uncal. yBP site of aBOO, we observed 39% ‘European’ hgs: U5a (26%), U4 (9%), T (4%), and 61% ‘Central/East Siberian’ hgs: C (35%), Z (13%), D (13%). Concordant with this admixed hg make-up, PCA indicated a position close to present-day Siberians (Figure 2). This position did not change when potential maternal relationships among individuals were accounted for by excluding redundant haplotypes (Figure S2B). The genetic relationship between aBOO and Siberians was also evident on the genetic distance map, where the area representing the lowest genetic distance covered a broader area of Siberia than for aUzPo (Figure 3B). The extant populations that showed most genetic similarity to aBOO were found in Central and East Siberia. In contrast, the area of maximum similarity for aUzPo lay in West Siberia (Figure 3A); this observation however could be influenced by low sample size in aUzPo.  A Tuvinian girl Haplotype sharing analyses for aBOO confirmed the genetic affinity with modern-day West and Central/East Siberians inferred from the PCA (Figure 4), but also identified a close relationship with the VUB population pool. The distribution of haplotype matches observed in pools of the VUB, West Siberia and Central/East Siberia was partly due to the presence of basal C* (16223T-16298C-16327T) and D* (16223T-16362C) haplotypes in these pools, whereas these types were absent in Middle Eastern and European pools. Central Siberian Tuvinians displayed the highest percentage of shared haplotypes with aBOO (12.2%) although all shared haplotypes belong to hgs C* and D*. A more explicit genetic link between aBOO and extant East Siberians was seen in the presence of the derived C5 haplotype (16148T-16223T-16288C-16298C-16311C-16327T) in aBOO and in one single Buryat individual of Central Siberia [54]. The Z1a haplotype (16129A-16185T-16223T-16224C-16260T-16298C) detected in aBOO had a broad but interesting distribution in Eurasia. It was found in all Central/East Siberian pools except in Tuvinians, but also in the Bashkirs of the Urals, in the VUB pool, as well as in Scandinavian and Baltic populations (Norwegians, Swedes, Finns, Ingrians, Karelians, and the Saami). Although haplotype sharing analyses revealed genetic links between aBOO and extant populations of NEE, a strong genetic differentiation was obvious between aBOO, modern-day NEE and Saami. This genetic discontinuity was further supported by BayeSSC analyses (Figure 5; Table 3). Similarly to aUzPo, a better fit was obtained for the model involving a 10% migration from CE over the last 3,500 years (H1b; ω = 1.00E+0) than for the model of genetic continuity between aBOO and NEE (H0b; ω = 3.86E-10).  Figure 6. Percentages of haplotypes from aUzPo and aBOO matched in selected ancient Eurasian populations. Previously described populations of hunter-gatherers of Central/East Europe (aHG [12], [14]) and Scandinavia (aPWC, [13]) were characterized by high frequencies and diversity of hg U4, U5a and U5b, which caused the two ancient datasets to group outside the cluster of extant European populations on the PCA plot (Figure 2). This matches previous studies that have shown that genetic continuity between hunter-gatherers and present-day Europeans can be rejected [12]–[13]. Like other European hunter-gatherers, aUzPo is characterized by high frequencies and diversity of hgs U4 and U5, but was genetically differentiated from aHG and aPWC due to the occurrence of hg C. Despite the fact that high frequencies of hgs U5b and V cluster the aHG and aPWC hunter-gatherer groups on the PCA plot (Figure 2), and that these hgs are also common in modern-day Saami, the ‘Saami motif’ is absent from aPWC and genetic continuity between aPWC and modern-day Saami was rejected [13]. Although the aBOO individuals were also characterized by high frequencies of hg U, the group appeared less close to the Palaeolithic/Mesolithic hunter-gatherers aHG and aPWC on the PCA plot than aUzPo. Haplotype sharing analyses (Figure 6) also showed that aBOO shared less haplotypes with aHG and aPWC than aUzPo (4.76% and 0.00%, respectively, versus 9.52% and 36.84%). This observation was confirmed by the analyses of our coalescent simulations, in which a model of genetic continuity between aHG, aPWC and aUzPo (ω = 9.91 E-1; H0d) was better supported than a model of genetic continuity between aHG, aPWC and aBOO (ω = 1.10 E-4; H0e). As demonstrated above, aBOO exhibited greater genetic affinities with extant populations of Siberia than aUzPo. Accordingly, aBOO shared more haplotypes with ancient samples from Siberia aEG (10.87% [55]) and aKUR (7.69% [56]) than aUzPo (0.00% and 7.69%, respectively; Figure 6). 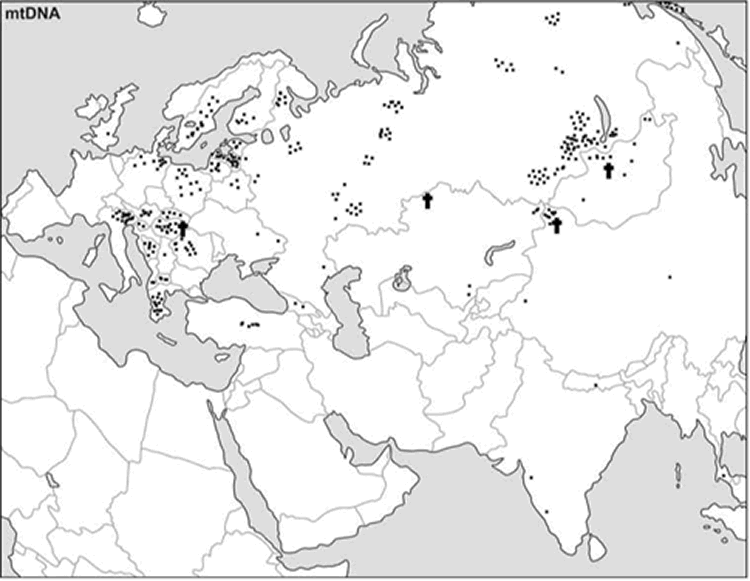 Current distribution pattern of the Y-STR haplotypes found in the ancient Siberians under study. Each square represents a present-day individual sharing the same Y-haplotype of an ancient specimen. To date, all studies on ancient Mesolithic/Palaeolithic hunter-gatherers from Europe have reported large proportions of hg U: 64% in aUzPo, 73% in aHG, 74% in aPWC; and hg U was also found in three out of five Mesolithic individuals of Spain [20], [57]. On the basis of the distribution of hg U5b, it was proposed that the Mesolithic population has remained genetically homogeneous over a wide geographical area and for a long period of time [57]. The new data from aUzPo suggests that hg U5a may be a representative of Central and North East Europe's Mesolithic mtDNA diversity, whereas elevated frequencies of hg U4 appear more characteristic of populations of the peri-Baltic area (aUzPo and aPWC). Haplogroup U also represents a significant genetic component of aBOO (35%), as well as Bronze Age Central Asians (14% in aKAZ; 2,700–3,400 yBP), and pre-Iron Age Siberians (54% in aKUR; Andronovo and Karasuk cultures; 2,800–3,800 yBP). Today, hg U is found in 7% of Europeans and displays a wide distribution in Europe, West Siberia, south west Asia, the Near East and North Africa [5]. Both the widespread distribution and high variability of hg U in extant and prehistoric populations are consistent with the description of hg U as one of the oldest hgs in Europe. On the basis of modern genetic data, hg U was proposed to have originated in the Near East and spread throughout Eurasia during the initial peopling by anatomically modern humans in the early Upper Palaeolithic (around 45,000 yBP, [5]). It is then plausible that hg U constituted the major part of the Palaeolithic/Mesolithic mtDNA substratum from Southern, Central and North East Europe to Central Siberia. It can also be suggested that the Palaeolithic/Mesolithic mtDNA substratum has been preserved longer in NEE than in Central and southern parts of Europe, where new lineages arrived with incoming farmers during the Neolithisation from the Near East [16]. This is supported by ancient genomic data obtained from hunter-gatherers of Scandinavia [58] and Spain [57], that shows a genetic affinity between Mesolithic individuals and present-day northern Europeans and supports genetic discontinuity between Mesolithic and Neolithic populations of Europe. The genetic links between the sample populations of aUzPo/aBOO and the extant populations of Siberia follow a general pattern discussed for the early and mid-Holocene (6,000–10,000 yBP). Facilitated by the East-West extension of vegetation zones between the Russian Far East and Eastern Europe [65], long-distance contacts and connections across Eurasia have been proposed for a number of cases. For example, the North East and East European hunter-gatherer pottery is thought to have originated in the early ceramic traditions of the Russian Far East and Siberia [66]–[68]. An eastern Asian origin followed by a westward expansion was also discussed for domesticated broomcorn millet (Panicum miliaceum L.) [69]. While the exact scenario behind these two examples of long-distance connections is unclear, migrations are a common interpretative model for evidence from later periods [70]. In any case, long-distance connections across Eurasia are not unusual. A later migration from the East was associated with the spread of the Imiyakhtakhskaya culture from Yakutia (East Siberia) through northwestern Siberia to the Kola Peninsula during the Early Metal Age (3,000–4,000 yBP, [24]). Interestingly, one individual of the aBOO site (grave 10, not sampled for aDNA here) was archaeologically associated with this culture, but its cultural relationships to other individuals of the same site remain unclear. Der Sarkissian C, Balanovsky O, Brandt G, Khartanovich V, Buzhilova A, Koshel S, et al. (2013) Ancient DNA Reveals Prehistoric Gene-Flow from Siberia in the Complex Human Population History of North East Europe. PLoS Genet 9(2): e1003296. doi:10.1371/journal.pgen.1003296 |
|
|
|
Post by Admin on Jun 8, 2018 18:24:26 GMT
 Figure 1: Schematic Representations of the U7, U7a and U7b Phylogenies. The maximum-parsimony reconstruction of 367 sequences of hg U7 yielded a tree with a basal hard polytomy that cannot be resolved (to a dichotomous one) at the level of whole-mtDNA sequence data: we identified eight independent branches that coalesce at the root of U7 (Fig. 1A and Supplementary Figure S1). Consistent with previous studies, we found that three major branches, U7a–c, capture most (96%) of the U7 mitogenomes. Besides these three previously known branches, we identified three additional clades, hereby designated as U7d, U7e, and U7f (Table 1). These were exclusively seen in Iran and the Caucasus. Finally, two mitogenomes – also from Iran and the Caucasus – did not cluster with any of hgs U7a–f and remained as unlabelled single lineages. In agreement with previous observations16, U7c appears to be restricted to South Asia (Fig. 1A and Supplementary Figure S1). In contrast, U7a is the dominant branch of U7 throughout the Near East and South Asia with subclades specific to Central Asia (U7a12–15), Mediterranean and Southeast Europe (U7a17 and U7a19; Figs 1B and 2B, Supplementary Figure S1). U7b exhibits a higher frequency than U7a in Europe with elevated levels of diversity in the Mediterranean and southeastern regions (Figs 1C and 2C and Supplementary Figure S1). It is distributed also in the Near East, South and Central Asia.  Figure 2: Spatial Frequency Distribution Maps of Haplogroups U7, U7a and U7b. We estimated a coalescence time for hg U7 at ~15.6–18.6 kya (Table 1), in agreement with previous maximum-likelihood estimates31. This confirms that U7 is the youngest major clade within the macro-hg U and the only one with the most recent common ancestor after the Last Glacial Maximum (LGM), presumably resulting from a severe glacial bottleneck. All other hg U subclades (U1, U2, U3, U4′9, U5, U6, and U8) display considerably older ages (~30–43 kya)31. This loss of genetic diversity of the ancestral U lineage, which eventually led to the formation of hg U7 is consistent with the survival of a small number of founders during the LGM; a pattern similar to that observed for mtDNA hgs N1a3 (previously N1c), N3, W, R2, HV, and within hgs M1 and U611,16,20,21,43,44,45. The hg U7 Bayesian skyline analysis (Fig. 3) shows a clear signal for an overall demographic expansion after the LGM. U7a drives the early stages of this demographic expansion, whereas the signal for U7b (the predominantly European subclade of hg U7) occurs much later, ~8–5 kya (Table 1 and Fig. 3). The subclades of U7a that are common in the Near East and South Asia (U7a1, U7a2, U7a3, and U7a10) are characterized by coalescence dates and a growth phase prior to the Holocene (Supplementary Figure S1 and Supplementary Table S4). Among those, U7a3 is both the oldest (~19 kya) and most frequent throughout these two areas, whilst U7a1, U7a2 and U7a10 are older than 12 kya. Clades U7a2, U7a3 and U7a10 have individual components, specific to the Near East and South Asia, suggesting that U7a was already differentiated in both regions by the end of the Pleistocene. |
|
|
|
Post by Admin on Jun 9, 2018 18:15:52 GMT
 Figure 3: Bayesian Skyline Plots of Haplogroups U7, U7a and U7b. Central Asia has four regionally specific clades (U7a12–15), whilst U7a11 is shared with South Asia (Fig. 1B and Supplementary Figure S1). U7 is distributed unevenly in Central Asia; it is most frequent in its southern areas adjacent to the Near East and South Asia (Fig. 2), with Afghanistan forming a “buffer zone”. Despite this geographical proximity, there are very few instances of shared lineages among the three regions, which, combined with the early coalescence date for U7a12 (~12 kya) (Supplementary Table S4), is consistent with a regional differentiation prior to the Holocene. In contrast to U7a, U7b shows signs of a significantly (t-test p < 0.001) later expansion and is characterized by low frequencies in the Near East, South Asia, and Central Asia, while it has a higher frequency in Europe (Table 1, Supplementary Tables S3 and S4). This differentiation is also reflected in the number of U7b subclades that are restricted to Europe – four out of nine subclades identified in the current phylogeny (Fig. 1C and Supplementary Figure S1). In addition, many single lineages (eight out of eighteen) in U7b are from Europe. The major sub-branches of U7b are characterized by star-like radiations and growth ~8–5 kya (Fig. 3). Subclades U7b1c and U7b1d, which are exclusive to Europe, expanded ~5 kya (Supplementary Figure S1). Hence, we consider this time as a minimum age for the presence of U7b in Europe. Taking into account the U7b coalescence age in Europe (Table 1), we think that U7b may have appeared there between 5 and 10 kya. This timeframe overlaps significantly with the time of the Neolithic demographic transition in Europe.  Expansions within this timeframe are also observed for the European-specific clades U7a17 and U7a19, whose distributions are centred on Mediterranean and Southeast Europe, along one of the preferred routes for the initial dispersal of farming46,47,48,49. Elevated frequencies in the Mediterranean area are also witnessed for many subsets of U7b (Fig. 2C), and the age of U7b in Europe (5–10 kya) is incompatible with its presence there prior to the Holocene. To date, aDNA studies have not found any example of U7 in either Neolithic or pre-Neolithic contexts. In contrast, other clades of hg U were common during the postglacial re-expansion period, including U8a, which is extremely rare today24. However, the number of analysed ancient samples from the Mediterranean area, where U7b has elevated frequencies today, is still rather small. The reduction in frequency of U7 in Europe from south to north is mirrored by the main components of hg K (a sister clade of U8b1). These have also been argued to have arrived into Europe during the early Neolithic from the Near East2,59,60,61, and display a clear northward frequency cline23,62. According to aDNA evidence, Neolithic populations in Europe display a distinct mtDNA lineage make-up, argued to be derived from Near Eastern sources. This early colonisation was probably followed by a complex process of assimilation of autochthonous hunter-gatherer diversity, seen most clearly in the autosomes. Notably, the distribution of nuclear genetic variants from Neolithic migrants among modern-day European populations3,55,64,65,66,67,68,69,70,71 resembles the phylogeography of hg U7 in Europe today. |
|
|
|
Post by Admin on Jun 10, 2018 18:24:52 GMT
 Another major episode of gene flow affecting the European gene pool appears to have occurred during the Late Neolithic and Early Bronze Age, from a source in the Pontic-Caspian Steppe region north of the Caucasus3,54,66,72. It has been suggested that this migration resulted in a further substantial shift in the genetic profile of Europeans and was a major vehicle for the movement of Indo-European languages to Europe3,72, and likely also to South Asia54. Interestingly, the autosomal genetic component in Europeans considered to derive from the Steppe is almost fixed in two pre-Neolithic ancient genomes from the South Caucasus. This component is distributed eastwards towards South Asia as well54, where it mimics the distribution of U7 (Pearson’s r = 0.65, p = 0.01). Our time estimates for the expansion and differentiation of hg U7 in the Near East, Central Asia, South Asia, and Europe, however, predate these putative late Neolithic-early Bronze Age migrations and thereby rule them out as a major vehicle for the spread of U7 to Europe and South Asia. In this respect, it is also noteworthy that Yamnaya herders of the Steppe so far analysed (n = 43) show no traces of U73,55,72,73 – and U7 is rarely found in this region today (Fig. 2). The expansion time of hg U7 in the Near East, Central Asia and South Asia is more consistent with autosomal multi-locus estimates for the genetic separation of these regions during the Terminal Pleistocene74, suggesting a common demographic process, whose origin was unclear previously. Here, we show that the frequency and distribution of U7b lineages indicate an origin of this clade in the Near East, whilst for U7a these statistics cannot differentiate between South Asia and the Near East (including the Caucasus) as a possible homeland. Within the Near East hg U7 is most frequent and diverse in Iran, whilst in South Asia its frequency and diversity peaks are in the Indus Valley region (Fig. 2 and Supplementary Figure S1). The demographic histories of the Near East and South Asia show marked differences during the LGM with the latter being less affected75. This is consistent with the long-term high effective population size and deep structure of autochthonous mtDNA haplogroups in South Asia76,77,78, in striking contrast to the severe population reduction affecting hg U7 during the last glacial period. Conversely, hg U2 in South Asia dates to more than 35 kya31, indicating its presence prior to the LGM.  Moreover, other haploid (Y chromosomal hg J2)79 and diploid80,81 genetic markers provide support for a post-glacial dispersal from the Near East to South Asia before the Bronze Age. Interestingly, recent aDNA studies have revealed a significant shared ancestry between Neolithic populations from the Zagros Mountains in Iran and contemporary populations from South Asia, suggesting eastward migration of people from that region to South Asia already at least in the Neolithic timeframe34,82. In conclusion, the Near East is the most likely ancestral homeland of U7. Our analyses reveal two temporally and geographically distinct signals of U7 expansion that disseminated from this region. The first signal dates shortly after the LGM and this dispersal is responsible for the spread of U7 towards South and Central Asia prior to the Holocene, while the more recent expansion explains its spread in Mediterranean Europe most probably during the early Holocene. These dispersals of hg U7 towards South Asia and Europe preclude any major association of U7 with the putative Bronze Age expansion of the Indo-European language family to these regions.  Scientific Reports volume 7, Article number: 46044 (2017) |
|










