|
|
Post by Admin on Mar 12, 2019 18:48:18 GMT
 Figure S3 Admixture Clustering Analysis of K = 2–K = 7 Clusters, Related to Figure 1 We constructed a model where ancient and present-day African populations trace their ancestry to a putative set of nine ancestral populations. As proxies for these populations, three different ancient Near Eastern populations and six African populations were used that, according to our analyses, harbor substantial ancestry related to major lineages present in Africa today. The Mende from Sierra Leone are used in this model to represent a component of ancestry that exists in high proportions in western African populations, the ancient southern African genomes (South_Africa_2000BP) are used to represent the ancestry of southern Africa before agriculture, the Ethiopian individual (Ethiopia_4500BP) is used to represent northeastern African ancestry before agriculture, the Mbuti are used to represent central African rainforest hunter-gatherer ancestry, the individual from an eastern African pastoralist context (Tanzania_Luxmanda_3100BP) is used to represent an early pastoralist lineage from eastern Africa (see below; Figure 2), and the Dinka (from Sudan) are used to represent distinctive ancestry found in Nilotic speakers today. The ancient Near Eastern populations were representative of Anatolia, the Levant, and Iran, respectively (Lazaridis et al., 2016, Mathieson et al., 2015). We used qpAdm (Haak et al., 2015), a generalization of f4 symmetry statistics, to successively test 1-source, 2-source, or 3-source models and admixture proportions for all other ancient and present-day African populations, with a set of 10 non-African populations as outgroups (STAR Methods).  Figure 2 Ancestral Components in Eastern and Southern Africa We find that ancestry closely related to the ancient southern Africans was present much farther north and east in the past than is apparent today. This ancient southern African ancestry comprises up to 91% of the ancestry of Khoe-San groups today (Table S5), and also 31% ± 3% of the ancestry of Tanzania_Zanzibar_1400BP, 60% ± 6% of the ancestry of Malawi_Fingira_6100BP, and 65% ± 3% of the ancestry of Malawi_Fingira_2500BP (Figure 2A). Notably, the Khoe-San-related ancestry in ancient individuals from Malawi and Tanzania is symmetrically related to the two previously identified lineages present in the San (Z < 2; Figure S2), estimated to have diverged at least 20,000 years ago (Mallick et al., 2016, Pickrell et al., 2012, Schlebusch et al., 2012), implying that this was an ancient divergent branch of this group that lived in eastern Africa at least until 1,400 BP. However, it was not present in all eastern Africans, as we do not detect it in the ∼400-year-old individual from coastal Kenya nor in the present-day Hadza. Displacement of Forager Populations in Eastern Africa Both unsupervised clustering (Figure 1B) and formal ancestry estimation (Figure 2B) suggest that individuals from the Hadza group in Tanzania can be modeled as deriving all their ancestry from a lineage related deeply to ancient eastern Africans such as the Ethiopia_4500BP individual (Figure 3A; Table S5). However, this lineage appears to have contributed little ancestry to present-day Bantu speakers in eastern Africa, who instead trace their ancestry to a lineage related to present-day western Africans, with additional components related to the Nilotic-speaking Dinka and to the Tanzania_Luxmanda_3100BP pastoralist (see below; Figure 2). The Sandawe, another population that like the Hadza uses click consonants in their spoken language, are modeled as having ancestry similar to the Hadza but also admixture related to that of neighboring populations (Figure 3A; Table S5) consistent with previous findings (Henn et al., 2011, Tishkoff et al., 2009). Population replacement by incoming food producers appears to have been nearly complete in Malawi, where we detect little if any ancestry from the ancient individuals who lived ∼8,100–2,500 BP. Instead, present-day Malawian individuals are consistent with deriving all their ancestry from the Bantu expansion of ultimate western African origin (Figure 3).  Figure 3 Mixture Events in the Deeper Population History of Continental African Lineages Among the ancient individuals analyzed here, only a ∼600 BP individual from the Zanzibar archipelago has a genetic profile similar to present-day Bantu speakers (Figure 1). Notably, this individual has even more western-African-related ancestry than the present-day Bantu speakers we analyzed from Kenya, who also derive some of their ancestry from lineages related to Dinka and Tanzania_Luxmanda_3100BP (Figure 1B). Using linkage disequilibrium, we estimate that this admixture between western- and eastern-African-related lineages occurred an average of 800–400 years ago (STAR Methods). This suggests a scenario of genetic isolation between early farmers and previously established foragers during the initial phase of the Bantu expansion into eastern Africa (Crowther et al., 2017, Ribot et al., 2010), a barrier that broke down over time as mixture occurred. This parallels the patterns previously observed in genomic analyses of the Neolithic expansion into Europe (Haak et al., 2015, Skoglund et al., 2012) and the East Asian farming expansion into Remote Oceania (Skoglund et al., 2016). However, this process of delayed admixture did not always apply in Africa, as is evident in the absence of admixture from previously established hunter-gatherers in present-day Malawians. Early Levantine Farmer-Related Admixture in an ∼3,100-Year-Old Pastoralist from Tanzania Western-Eurasian-related ancestry is pervasive in eastern Africa today (Pagani et al., 2012, Tishkoff et al., 2009), and the timing of this admixture has been estimated to be ∼3,000 BP on average (Pickrell et al., 2014). We found that the ∼3,100 BP individual (Tanzania_Luxmanda_3100BP), associated with a Savanna Pastoral Neolithic archeological tradition, could be modeled as having 38% ± 1% of her ancestry related to the nearly 10,000-year-old pre-pottery farmers of the Levant (Lazaridis et al., 2016), and we can exclude source populations related to early farmer populations in Iran and Anatolia. These results could be explained by migration into Africa from descendants of pre-pottery Levantine farmers or alternatively by a scenario in which both pre-pottery Levantine farmers and Tanzania_Luxmanda_3100BP descend from a common ancestral population that lived thousands of years earlier in Africa or the Near East. We fit the remaining approximately two-thirds of Tanzania_Luxmanda_3100BP as most closely related to the Ethiopia_4500BP (p = 0.029) or, allowing for three-way mixture, also from a source closely related to the Dinka (p = 0.18; the Levantine-related ancestry in this case was 39% ± 1%) (Table S4). While these findings show that a Levant-Neolithic-related population made a critical contribution to the ancestry of present-day eastern Africans (Lazaridis et al., 2016), present-day Cushitic speakers such as the Somali cannot be fit simply as having Tanzania_Luxmanda_3100BP ancestry. The best fitting model for the Somali includes Tanzania_Luxmanda_3100BP ancestry, Dinka-related ancestry, and 16% ± 3% Iranian-Neolithic-related ancestry (p = 0.015). This suggests that ancestry related to the Iranian Neolithic appeared in eastern Africa after earlier gene flow related to Levant Neolithic populations, a scenario that is made more plausible by the genetic evidence of admixture of Iranian-Neolithic-related ancestry throughout the Levant by the time of the Bronze Age (Lazaridis et al., 2016) and in ancient Egypt by the Iron Age (Schuenemann et al., 2017). |
|
|
|
Post by Admin on Mar 13, 2019 18:02:57 GMT
Displacement of Forager Populations in Eastern Africa Both unsupervised clustering (Figure 1B) and formal ancestry estimation (Figure 2B) suggest that individuals from the Hadza group in Tanzania can be modeled as deriving all their ancestry from a lineage related deeply to ancient eastern Africans such as the Ethiopia_4500BP individual (Figure 3A; Table S5). However, this lineage appears to have contributed little ancestry to present-day Bantu speakers in eastern Africa, who instead trace their ancestry to a lineage related to present-day western Africans, with additional components related to the Nilotic-speaking Dinka and to the Tanzania_Luxmanda_3100BP pastoralist (see below; Figure 2). The Sandawe, another population that like the Hadza uses click consonants in their spoken language, are modeled as having ancestry similar to the Hadza but also admixture related to that of neighboring populations (Figure 3A; Table S5) consistent with previous findings (Henn et al., 2011, Tishkoff et al., 2009). Population replacement by incoming food producers appears to have been nearly complete in Malawi, where we detect little if any ancestry from the ancient individuals who lived ∼8,100–2,500 BP. Instead, present-day Malawian individuals are consistent with deriving all their ancestry from the Bantu expansion of ultimate western African origin (Figure 3).  Figure 3 Mixture Events in the Deeper Population History of Continental African Lineages Among the ancient individuals analyzed here, only a ∼600 BP individual from the Zanzibar archipelago has a genetic profile similar to present-day Bantu speakers (Figure 1). Notably, this individual has even more western-African-related ancestry than the present-day Bantu speakers we analyzed from Kenya, who also derive some of their ancestry from lineages related to Dinka and Tanzania_Luxmanda_3100BP (Figure 1B). Using linkage disequilibrium, we estimate that this admixture between western- and eastern-African-related lineages occurred an average of 800–400 years ago (STAR Methods). This suggests a scenario of genetic isolation between early farmers and previously established foragers during the initial phase of the Bantu expansion into eastern Africa (Crowther et al., 2017, Ribot et al., 2010), a barrier that broke down over time as mixture occurred. This parallels the patterns previously observed in genomic analyses of the Neolithic expansion into Europe (Haak et al., 2015, Skoglund et al., 2012) and the East Asian farming expansion into Remote Oceania (Skoglund et al., 2016). However, this process of delayed admixture did not always apply in Africa, as is evident in the absence of admixture from previously established hunter-gatherers in present-day Malawians. 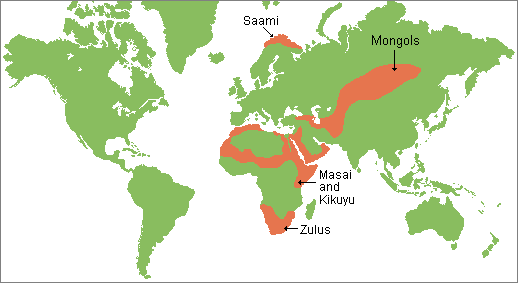 Early Levantine Farmer-Related Admixture in an ∼3,100-Year-Old Pastoralist from Tanzania Western-Eurasian-related ancestry is pervasive in eastern Africa today (Pagani et al., 2012, Tishkoff et al., 2009), and the timing of this admixture has been estimated to be ∼3,000 BP on average (Pickrell et al., 2014). We found that the ∼3,100 BP individual (Tanzania_Luxmanda_3100BP), associated with a Savanna Pastoral Neolithic archeological tradition, could be modeled as having 38% ± 1% of her ancestry related to the nearly 10,000-year-old pre-pottery farmers of the Levant (Lazaridis et al., 2016), and we can exclude source populations related to early farmer populations in Iran and Anatolia. These results could be explained by migration into Africa from descendants of pre-pottery Levantine farmers or alternatively by a scenario in which both pre-pottery Levantine farmers and Tanzania_Luxmanda_3100BP descend from a common ancestral population that lived thousands of years earlier in Africa or the Near East. We fit the remaining approximately two-thirds of Tanzania_Luxmanda_3100BP as most closely related to the Ethiopia_4500BP (p = 0.029) or, allowing for three-way mixture, also from a source closely related to the Dinka (p = 0.18; the Levantine-related ancestry in this case was 39% ± 1%) (Table S4). While these findings show that a Levant-Neolithic-related population made a critical contribution to the ancestry of present-day eastern Africans (Lazaridis et al., 2016), present-day Cushitic speakers such as the Somali cannot be fit simply as having Tanzania_Luxmanda_3100BP ancestry. The best fitting model for the Somali includes Tanzania_Luxmanda_3100BP ancestry, Dinka-related ancestry, and 16% ± 3% Iranian-Neolithic-related ancestry (p = 0.015). This suggests that ancestry related to the Iranian Neolithic appeared in eastern Africa after earlier gene flow related to Levant Neolithic populations, a scenario that is made more plausible by the genetic evidence of admixture of Iranian-Neolithic-related ancestry throughout the Levant by the time of the Bronze Age (Lazaridis et al., 2016) and in ancient Egypt by the Iron Age (Schuenemann et al., 2017).  Direct Evidence of Migration Bringing Pastoralism to Eastern and Southern Africa In contrast to the Malawi and Zanzibar individuals, all three ancient southern Africans show affinities to the ancestry predominant in present-day Tuu speakers in the southern Kalahari more than to present-day Ju|'hoan speakers in the northern Kalahari (Figures S2B and S2C). However, the ∼1,200 BP sample from the western Cape that is found in a pastoralist context has a specific similarity in clustering analyses to present-day Khoe-Khoe-speaking pastoralist populations such as the Nama (Figure 1B), and like them it has affinity to three groups: Khoe-San, western Eurasians, and eastern Africans. This supports the hypothesis that a non-Bantu-related population carried eastern African and Levantine ancestry to southern Africa by at least around 1,200 BP, providing direct evidence for claims previously made based on analysis of present-day populations (Pickrell et al., 2014). We used our modeling framework to show that the South_Africa_1200BP pastoralist individual from the western Cape is consistent with being a mixture of just two streams of ancestry relative to non-southern African populations, with 40.3% ± 2.3% ancestry related to the Tanzania_Luxmanda_3100BP individual (54% ± 7% when restricting analysis to sequences with postmortem damage) and the remainder being related to the South_Africa_2000BP hunter-gatherers (Table S5). This supports the hypothesis that the Savanna Pastoral Neolithic archaeological tradition in eastern Africa is a plausible source for the spread of herding to southern Africa. Even the Ju|'hoan San group with the least genetic affinity to eastern Africans (Ju_hoan_North), have 9% ± 1% of their ancestry most closely related to Tanzania_Luxmanda_3100BP, consistent with previous findings that the ancestries of all present-day San and Khoe were affected by agro-pastoralist migrations in the last two millennia (Pickrell et al., 2014). |
|
|
|
Post by Admin on Mar 13, 2019 20:31:54 GMT
The Earliest Divergences among Modern Human Populations Previous studies have suggested that the primary ancestry in the San is from a lineage that separated from all other lineages represented in modern humans today, before the latter separated from each other (Gronau et al., 2011, Veeramah et al., 2012). Such a model emerges when we automatically fit a tree without admixture to the data (Figure 3A), but we also find that a tree-like representation is a poor fit (Figure S4A), in the sense that ancient southern Africans who lived ∼2,000 BP were not strictly an outgroup to extant lineages in other parts of sub-Saharan Africa. In particular, we find that ancient southern Africans, who have none of the eastern African admixture that is ubiquitous today, share significantly more alleles with present-day and ancient eastern Africans (including Dinka, Hadza, and Ethiopia_4500BP) than they do with present-day western Africans (Figure 3B; Table S6). Even within present-day western Africans, the genetic differences between Yoruba from Nigeria and the Mende from Sierra Leone are inconsistent with descent from a homogeneous ancestral population isolated from ancient southern Africans. The asymmetry between Yoruba and Mende is also observed with non-Africans but is no stronger than in eastern Africans (the most closely related Africans to the ancestral out-of-Africa population), and thus these signals are not driven by admixture from outside Africa and instead likely reflect demographic events entirely within Africa (Figure 3C; Table S6).  Figure S4 Admixture Models of African Population History, Related to Figure 3 We carried out admixture graph modeling of the allele frequency correlations and found two parsimonious models that fit the data. The first posits that present-day western Africans harbor ancestry from a basal African lineage that contributed more to the Mende than it did to the Yoruba, with the other source of western African ancestry being related to eastern Africans and non-Africans (Figures 3D, S4, and S5; Table S7). The second model posits that long-range and long-standing gene flow has connected southern and eastern Africa to some groups in western Africa (e.g., the ancestors of the Yoruba) to a greater extent than to other groups in western Africa (e.g., the ancestors of the Mende) (Figure 3E) (Pleurdeau et al., 2012). The possible basal western African population lineage would represent the earliest known divergence of a modern human lineage that contributed a major proportion of ancestry to present-day humans. Such a lineage must have separated before the divergence of San ancestors, which is estimated to have begun on the order of 200–300 thousand years ago (Scally and Durbin, 2012). Such a model of basal western African ancestry might support the hypothesis that there has been ancient structure in the ancestry of present-day Africans, using a line of evidence independent from previous findings based on long haplotypes with deep divergences from other human haplotypes (Hammer et al., 2011, Lachance et al., 2012, Plagnol and Wall, 2006). One scenario consistent with this result could involve ancestry related to eastern Africans (and the out-of-Africa population) expanding into western Africa and mixing there with more basal lineages. Our genetic data do not support the theory that this putative basal lineage diverged prior to the ancestors of Neanderthals, since the African populations we analyze here are approximately symmetrically related to Neanderthals (Mallick et al., 2016, Prüfer et al., 2014).  Figure 4 Ancient Genomes Provide Evidence of Natural Selection in Present-Day Southern African San Populations A Selective Sweep Targeting a Taste Receptor Locus in Southern Africa The availability of ancient African genomes provides an opportunity to search for genomic footprints of natural selection, manifested as regions of greater allele frequency differentiation between ancient and present-day populations than predicted by the genome-wide background. We compared the two ancient southern African ∼2,000 BP shotgun sequence genomes to six present-day high-coverage San genomes with minimal recent mixture. The small number of ancient individuals does not permit inference of changing allele frequencies at single loci, so we performed a scan for high allele frequency differentiation in 500 kb windows with a step size of 10 kb. Using ∼500 windows spaced at least 5 million base pairs apart as a null distribution, we found that the most differentiated locus was 15 standard deviations from the observed genome-wide mean and overlapped a cluster of eight taste-receptor genes on chromosome 12 (Figure 4A; Table 2). Taste-receptor genes have previously been identified as targets of natural selection in humans, as they modulate the ability to detect poisonous compounds in plants (Campbell et al., 2012). Natural selection on phenotypic traits in humans is expected to only occasionally take the form of sweeps on a single locus, instead acting on multiple genes simultaneously to drive phenotypic adaptation (Coop et al., 2009). While a lack of genome-wide association studies in eastern and southern Africans has left the genetic basis of phenotypic traits far less well documented than it is for other populations, a variety of studies have linked broad functional classes of genes to phenotypic traits. To test for evidence of selection on specific functional categories of genes in present-day San since the divergence of the two ancient genomes from southern Africa (Figure 4B), we estimated allele frequency differentiation for 208 gene ontology categories with 50 or more genes in each, and we computed weighted block jackknife standard errors. The functional category that displays the most extreme allele frequency differentiation between present-day San and ancient southern Africans is “response to radiation” (Z = 3.3 compared to the genome-wide average). To control for the possibility that genes in this category show an inflated allele frequency differentiation in general, we computed the same statistic for the Mbuti central African rainforest hunter-gatherer group but found no evidence for selection affecting the response to radiation category (Figure 4C). Instead, the top category for the Mbuti is “response to growth,” suggesting the possibility that the small stature of rainforest hunter-gatherer populations such as the Mbuti may be an acquired adaptation (although we have no ancient central African genome and thus no information about the time frame of selection). We speculate that the signal for selection in the response to radiation category in the San could be due to exposure to sunlight associated with the life of the ‡Khomani and Ju|'hoan North people in the Kalahari Basin, which has become a refuge for hunter-gatherer populations in the last millenia due to encroachment by pastoralist and agriculturalist groups (Morris, 2002). Discussion This study, which multiplies by 16-fold the number of individuals with genome-wide ancient DNA data from sub-Saharan Africa, highlights the power of ancient African genomes to provide insights into prehistoric events that are difficult to discern based solely on analysis of present-day genomes. We reveal the presence of a hitherto unknown cline of geographically structured hunter-gatherer populations stretching from Ethiopia to South Africa, which we show existed prior to the great population transformations that occurred in the last few thousand years in association with the spread of herders and farmers. We also document deeper structure in western Africa, possibly predating the divergence of the ancestors of southern African hunter-gatherers from other population lineages. We finally provide case examples of how populations in eastern and southern Africa were transformed by the spread of food producers and show how the process gave rise to interactions with the previously established hunter-gatherers, with the outcomes ranging from no detectable mixture in present-day populations to substantial mixture. Our documentation of a radically different landscape of human populations before and after the spread of food producers highlights the difficulty of reconstructing the African past based solely on analysis of present-day populations and the importance of using ancient DNA to study deep African population history in an era in which technological improvements have now made this feasible. It is clear that ancient DNA studies with larger sample sizes and covering a broader chronological and geographic range have the potential to make major progress in improving our understanding of African prehistory. Cell, VOLUME 171, ISSUE 1, P59-71.E21, SEPTEMBER 21, 2017 |
|
|
|
Post by Admin on Mar 27, 2019 18:22:15 GMT
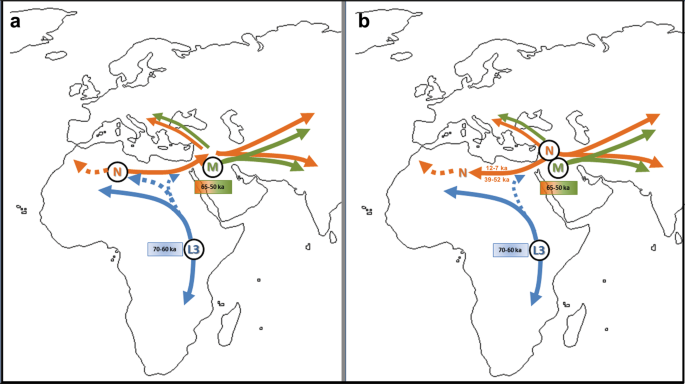 Figure 4 Fossil remains attest to the presence of Anatomically Modern Humans (AMH) in the Near East around 130–100 kilo annum (ka), but their decisive spread from Africa all over the world seems to be the result of a migration that occurred after 70 ka1,2,3. African genetic variability is well known only for current times, but local population genetics dynamics and back-migrations from Eurasia could have modified haplotype frequencies, thereby masking ancient genetic scenarios4,5,6,7. Several variables, such as temperature, pH and salinity, influence molecular degradation8,9 and environmental conditions in hot and arid regions of Africa do not favour DNA preservation. For these reasons, only in the last years, thanks to methodological improvements in the ancient DNA field, genetic data from ancient human samples have been obtained for the African continent10,11,12,13,14,15,16. Here we present the earliest and first genetic data from the Saharan region and contextualize them into the mitochondrial phylogeny, in order to enhance the understanding of past African genetic variability and human lineage evolution and dispersal. The analysed material comes from Takarkori rockshelter in the southern Tadrart Acacus massif of Libya’s central Sahara (Fig. S1). The rockshelter’s thick stratigraphy spans several millennia of the Holocene and documents the transition from hunting and gathering to pastoralism17. Fifteen burials mostly referred to the Early and Middle Pastoral (Neolithic) date to between ~8.9 and ~4.8 ka. Located in a recessed area of the shelter, these burials were exclusively of women of reproductive age, children and juveniles. Strontium isotope analysis revealed all have the same local geographic origin. These patterns suggest a kinship system based on matrilineal descent with repeated use of the rockshelter as deathplace by early pastoral communities18. The DNA analysis focuses on two individuals of Middle Pastoral age, who present signs of natural mummification: TK RS H1 and TK RS H9 (Fig. 1). The samples are directly radiocarbon dated to 6090 ± 60 BP (7.1–6.7 ka; 7159–6797 calBP, 95.4% probability) and 5600 ± 70 BP (6.5–6.2 ka; 6547–6280 calBP, 95.4% probability) respectively (Table S1). Both belong to adult females18. We extracted DNA from several bone and skin samples (Table S2) and performed a capture enrichment for the mitochondrial genome (mtDNA). 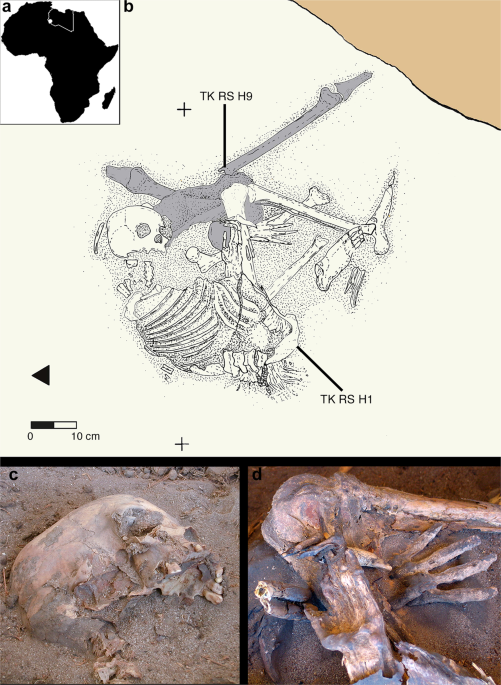 Figure 1 Ancient DNA analysis A tooth sample for TK RS H1 gave a mean coverage of the mtDNA of 72x with 99.99% of positions covered at least threefold (Table S3). Two bone fragments from the fibula of TK RS H9 gave a mean coverage of the mtDNA of 8x and 8.4x with 94.13% and 89.75% of positions covered at least threefold respectively (Table S3). Nucleotide misincorporation and fragmentation patterns of the molecules were consistent with the samples’ age and environmental conditions8 (Table S3, Figs S2, S3). Modern human DNA contamination is absent (Table S4). Phylogenetic analysis Comparing the consensus sequences of individuals TK RS H1 and TK RS H9, the same haplotype was found except for possible private mutations in TK RS H9 masked by missing data (Table S5). According to radiocarbon datings (Table S1) the two individuals were separated by a maximum of 879 and a minimum of 250 years, corresponding to 35–10 25-year-long generations, hence the perfect match of the sequences obtained sustains the hypothesis of close maternal kinship. This supports the cultural interpretation of the repeated use of the rockshelter to dispose of members of the same lineage18. The sequences were analysed regarding their position in the human mitochondrial phylogeny and were attributed to the N haplogroup with some private mutations. A Median Joining Network was drawn by combining the sequences obtained for TK RS H1 and TK RS H9 with an extended reference dataset of 536 modern individuals belonging to the different branches of macro-haplogroups N and L3, from which N originated. Further, we included 8 ancient European and African samples, selected among the published mitogenomes, with a special emphasis in basal informative haplogroups and added the rCRS and RSRS reference sequences for comparison (Fig. 2; Table S6). A Bayesian phylogenetic tree was inferred from a multi-sequence alignment of the mtDNA sequence of TK RS H1, as representative of the haplotype found in Takarkori, 167 modern samples with lineage relevance for the main haplogroups and sub-haplogroups and 42 published complete genomes of dated ancient samples as tip calibration points (Fig. 3; Fig. S4; Tables S6, S7). Both the Median Joining Network and the phylogenetic tree confirmed the attribution of the sample to the N haplogroup, highlighting it as a basal lineage, which branches off immediately after the Palaeolithic sample Oase 119 and before all present-day N-derived mtDNAs. A mutation rate of 2.26 × 10−8 mutation/site/year (95% HPD: 1.85–2.68 × 10−8) was estimated by the Bayesian analysis, consistent with the results of previous studies3,20. The time to the most recent common ancestor (TMRCA) for N clade and for the branch leading to Takarkori sequence was estimated at 64,103 years BP (95% HPD: 54,535–74,578) and 61,618 years BP (95% HPD: 52,517–71,405) respectively. The tip date for the TK RS H1 terminal node was estimated to be 12,325 years BP (95% HPD: 1–25,119). The large confidence interval, presumptively affected by the small number of sites of the mtDNA genome and short evolutionary time scale, is consistent with the radiocarbon date of the sample taking into account the confidence intervals and provides further evidence for the authenticity of the sequence. 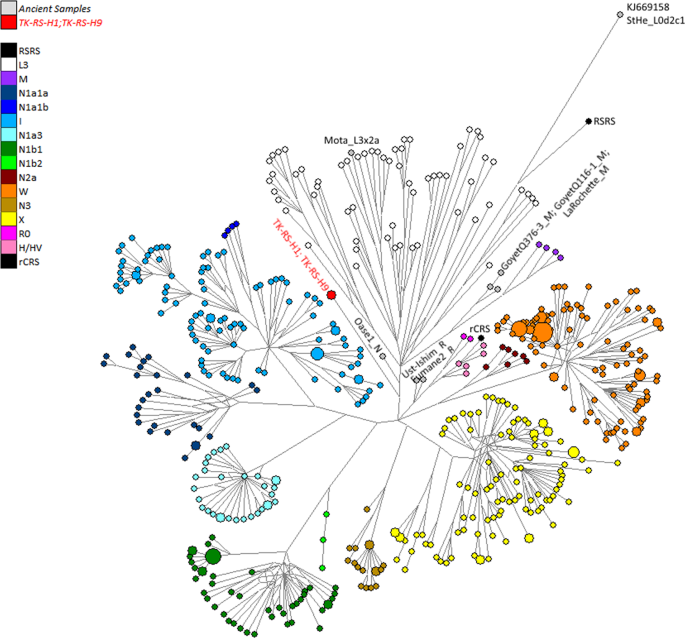 Figure 2 Until now, the origin of African haplogroup L3, from which N originated, has been dated around 60–70 ka and its expansion in Eastern Africa linked with the exit of AMH from the continent. The M and N haplogroups, which lie at the base of Eurasian mtDNA diversity, are today globally distributed outside Africa and are dated to around 50–65 ka, very close to the ancestral L3 clade. Their divergence from it is commonly considered to have occurred outside Africa or during the expansion1,2,21,22. The Arabian Peninsula represents a possible area where this occurred and a cradle from which the new branches spread toward Eurasia and back to Africa, including N1a and R0a, both of which are found in East Africa1,7,23,24. The uneven geographical distribution of existing data could bias the representation of real past genetic variability: sampling gaps characterize the African continent, and several studies focus only on particular haplogroups. Moreover, the past genetic scenario is still poorly known for this large and crucial area, and the ancient mitochondrial sequences available refers only to few sites distributed in Ethiopia, Egypt, Morocco, Kenya, Malawi, Tanzania and South Africa and do not extend back in time more than ~15000 years. 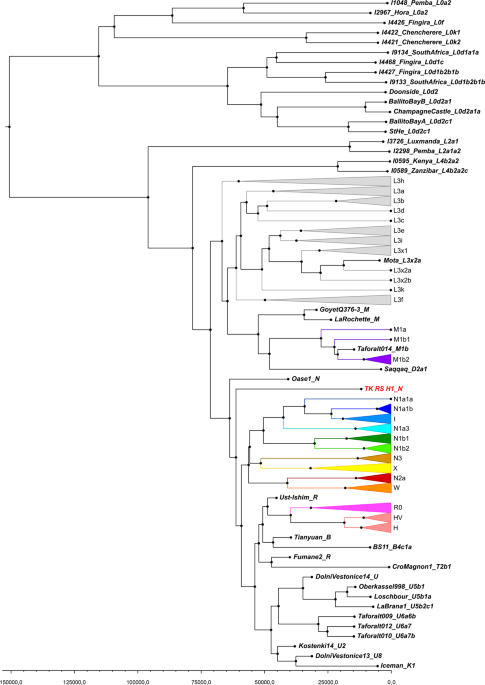 Figure 3 Our research reveals that the Neolithic Saharan individuals from Takarkori present a haplotype not previously identified in Africa, that belongs to a basal branch of haplogroup N. This discovery needs to be addressed cautiously, given its potential geographical, chronological and archaeological implications (Fig. 4). As recently suggested23, the presence of an unexpected branch where other clades prevail in the population may provide an indication of ancestry, but more data are necessary. The Saharan region was interested by strong climatic oscillations. Repeated peaks of humidity and the presence of several intermittent pulses of lake activity occurred between 125 and 11 ka25. Warmer and wetter environmental conditions characterized the Late Glacial Bølling/Allerød Interstadial26 allowing population growth and spread. The return of cooler, drier conditions during the Younger Dryas may have prompted human groups to exploit glacial refugia across this region. Interestingly the molecular date of the Takarkori sequence (12,325 BP) falls into the context of the Interstadial expansion. The analysed samples, dated to ~7000 BP, could represent a signal of a mitochondrial lineage that later disappeared because of genetic drift due to population contraction and isolation with the beginning of desertification27,28. A possible scenario envisages an introgression from Eurasia in ancient times that carried haplotypes that have since disappeared from Africa. The timing of this migration remains difficult to define. Late Pleistocene dispersal from Western Asia into Africa around 39–52 ka is suggested by the expansion of the U6 haplogroup29,30, with a potentially corresponding archaeological signature in the MSA Dabban industry of Cyrenaica, Libya, ca. 45–40 ka31. Individuals carrying a N haplogroup basal lineage could have followed the same dispersion pattern as U6: their legacy could have been survived up to ∼7000 years ago in the central Sahara thanks to the climatic conditions previously described, but replaced and disappeared in other parts of North Africa. Genomic data for seven 15,000-year-old individuals attributed to the Iberomaurusian culture in Taforalt (Morocco) suggest a connection with Epipaleolithic Natufians from Near East, while seem to exclude a possible gene flow from Upper Paleolithic Europe32. Our samples postdate the Taforalt individuals by up to 8,000 years and belong to Neolithic pastoral cultures of the Middle Holocene. It is known that livestock was introduced from Southwest Asia33 and early pastoralist connections between Northeast Africa and Arabia are indicated by a few sites along the Red Sea with sheep/goat dated to ~8.1–7.5 ka34,35,36. Thus, the spread of pastoralism from the Levant to Northeast Africa could probably represent the context for the introgression of the N haplogroup into the central Sahara, even if it is commonly associated with derivative lineages (N1)1,37. It is worth noting, however, that when geometric morphometric analysis of the skull of TK RS H1 is compared with a large published dataset it shows closer affinities with sub-Saharan contests38, such as Gobero in Niger whose occupation is dated from ~9.6–4.8 ka39. Unfortunately, no genetic data are available for this region that could help understanding the possible origin of the haplotype found at Takarkori. Scientific Reportsvolume 9, Article number: 3530 (2019) |
|
|
|
Post by Admin on Jun 6, 2019 21:13:31 GMT
Within the intensively studied field of early hominin evolution, a crucial question is the split of our own clade from the Panini. Over the last decades the fossil record of potential early hominins increased with taxa such as Ardipithecus, Orrorin and Sahelanthropus [1–3]. Recent molecular data propose a divergence time of Pan and Homo between 5 and 10 Ma [4] and Langergraber et al. [5] propose an age of at least 7–8 Ma. These estimations largely coincide with the evidence obtained from the fossil record across Africa and Eurasia [6, 7].  Ouranopithecus In the present study, we define ‘hominoid’ as ‘apes’; ‘hominid’ as ‘great apes and humans’; ‘hominine’ as ‘African apes and humans’; and ‘hominin’ as ‘humans and their non-ape ancestors’. Currently, the fossil record reveals three Miocene candidates with potential hominin affinity. Ardipithecus kadabba is dated to between 5.2 and 5.8 Ma. It is more primitive than Ardipithecus ramidus and may not belong to the same genus [8], but it does show hominin affinities such as evidence of bipedalism and canine reduction [9, 10]. Orrorin tugenensis is dated to ~5.8–6.0 Ma and shows an upright posture [2, 11]. Sahelanthropus tchadensis is dated to ~6–7 Ma [3, 12] and provides several derived cranial and dental features that suggest hominin affinity. Lebatard et al. [13] propose an age of 7.2–6.8 Ma for Sahelanthropus. We do not consider this age determination to be reliable given the circumstances of the provenance of the skull [14] and the relatively low accuracy of the method [15]. The overwhelming effort to reconstruct hominin origins have been focused on the African continent. However, ancestral lineages remain largely unknown [16]. A crucial problem in identifying ancestral lineages is the prevalence of homoplasy and the relative lack of derived morphological features that reduces the phylogenetic resolution around lineage divergence [17, 18]. Root morphology might be a potential feature, which is less affected by homoplasy. Studies on fossil hominids, extant great apes and humans indicate that the premolar root number is not primarily linked to a functional adaptation, and is interpreted to represent a genetic polymorphism [19, 20]. Hence, homoplasy is only a minor consideration for the traits of premolar root numbers, which therefore may provide a useful phylogenetic signal. Nevertheless, some relations of root and crown morphology indicate overlaying masticatory adaptations that may attenuate the phylogenetic signal [21, 22].  Fig 1. Studied specimens and virtual reconstructions of the holotype of Graecopithecus. a, Type mandible of G. freybergi from Pyrgos, Greece. b, RIM 438/387 –Left P4 of cf. Graecopithecus sp. from Azmaka, Bulgaria. From left to right: distal, mesial, lingual, buccal, occlusal and apical. c-i, μCT based 3D reconstructions of the type mandible showing the partially preserved roots and pulp canals from c-m3 and the crowns of right p4-m2. Further images with a magnification of the virtually isolated teeth and pulp canals are provided in S1 Fig. c, Occlusal view. d-e, Apical view. f, Buccal view of the left hemimandible. g, Buccal view of the right hemimandible. h, Lingual view of the left hemimandible. i, Lingual view of the right hemimandible. Scale bars, 10 mm Of special importance for hominin evolution is the lower fourth premolar (p4), as its morphology seems to be diagnostic for the hominin lineage. Taxonomic attempts have been made concerning its crown morphometry [23–25] and especially its root configuration [26, 27], which turns out to be a powerful tool for early hominin phylogeny [28]. Several morphological traits of putative early hominin p4s (Sahelanthropus, Ar. kadabba, Ar. ramidus) point to a reduced configuration. A two-rooted, but narrow state is documented in Sahelanthropus [28, 29]. A Tomes’ root is present in Ardipithecus kaddaba and a single-rooted p4 is characteristic for Ardipithecus ramidus [1, 30, 31] and Homo. The plesiomorphic p4 root configuration shown by extant great apes, basal hominids like Proconsul and Miocene hominines (Ouranopithecus) differs significantly, showing two or three clearly diverging roots and four pulp canals [28, 32]. The p4 root number in australopithecines (Au. anamensis, Au. afarensis, Au. africanus; [33–37]) is highly variable, from a Tomes’ root up to a three-rooted condition [26]. Another p4 root morphology, which has two roots that are fused on their basal buccal part, is recently described for some specimens of P. robustus, Au. africanus and australopithecines from Woranso-Mille [25, 36]. In this study, we propose based on root morphology a new possible candidate for the hominin clade, Graecopithecus freybergi from Europe. Graecopithecus is known from a single mandible from Pyrgos Vassilissis Amalia (Athens, Greece) [38] and possibly from an isolated upper fourth premolar (P4) from Azmaka in Bulgaria [39] (Fig 1A and 1B). A new age model for the localities Pyrgos Vassilissis and Azmaka, as well as the investigations on the fauna of these localities [40] confirms that European hominids thrived in the early Messinian (Late Miocene, 7.25–6 Ma) and therefore existed in Europe ~ 1.5 Ma later than previously thought [39]. This, and recent discoveries from Çorakyerler (Turkey), and Maragheh (Iran) demonstrate the persistence of Miocene hominids into the Turolian (~8 Ma) in Europe, the eastern Mediterranean, and Western Asia [41, 42]. |
|












