|
|
Post by Admin on Oct 13, 2016 21:37:47 GMT
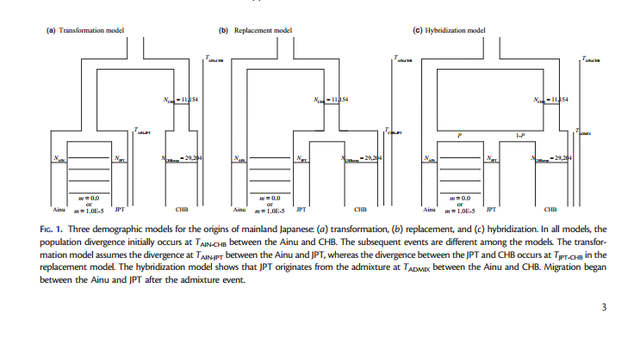 FIG. 1. Three demographic models for the origins of mainland Japanese: (a) transformation, (b) replacement, and (c) hybridization. In all models, the population divergence initially occurs at TAIN-CHB between the Ainu and CHB The dual structure model for the population history of the Japanese people is a hybridization model that Hanihara (1991) proposed concerning the demographic history of prehistoric Japanese based on morphological analyses of crania and teeth (Hanihara 1991). The dual structure model proposes two main events of (I) migration and (II) admixture: (I) the Jomon people originated somewhere in Southeast Asia and came to the Japanese archipelago around 12,000 years ago, whereas the Yayoi people came from northeast Asia around 2,000 years ago and entered the northern part of Kyushu, which is the western island of Honshu (main-island Japan), probably via the Korean peninsula. Then, (II) the Yayoi migrants admixed with the indigenous Jomon people gradually from west to east. Because the degree of Jomon or Yayoi ancestry may vary among local populations, the Japanese populations can be characterized by the dual structure of ancestral populations: people in main-island Japan have a high degree of genetic contribution from the Yayoi migrants, whereas the Ainu on Hokkaido (the northernmost island) and the Ryukyuans in Okinawa (the southernmost islands of the Japanese archipelago) have a higher degree of genetic contribution from the indigenous Jomon. Meanwhile, recent morphological studies have reported that Jomon skeletal series vary among geographic regions and that cranial variation is clearly distinct between the northeast and the southwest portions of the Japanese archipelago (Hanihara and Ishida 2009; Nakashima et al. 2010; Fukase et al. 2012a, 2012b). Therefore, the Jomon people are likely to have already diversified among the local regions before the Yayoi migration happened; thus, the population history in the Japanese archipelago may be complicated with the population structure of the Jomon. 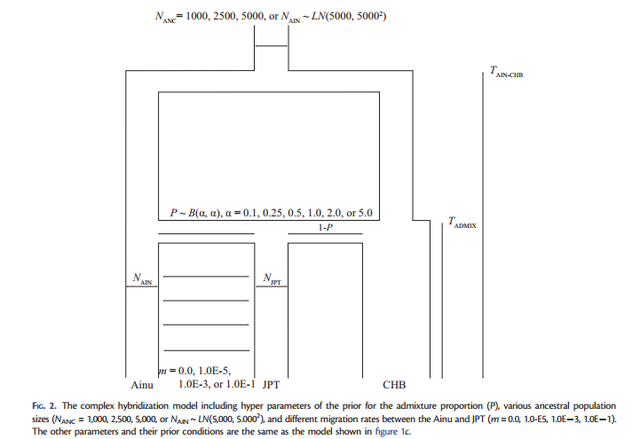 FIG. 2. The complex hybridization model including hyper parameters of the prior for the admixture proportion (P), various ancestral population sizes and different migration rates between the Ainu and JPT. The demographic models (transformation, replacement, and hybridization) are illustrated in figure 1, including three parameters common to the three models (NAIN: population size in Ainu; NJPT: population size in JPT; TAIN-CHB: time of population divergence between Ainu and CHB) and four parameters specific to each model (TAIN-JPT: time of population divergence between Ainu and JPT; TJPT-CHB: time of population divergence between JPT and CHB; TADMIX: time of admixture between Ainu and CHB; P: the admixture proportion from the Jomon ancestry). A migration parameter (m= 1.0E5) after the admixture event was incorporated between Ainu and JPT to evaluate this effect on the model selection. The transformation model was characterized as the modern Japanese as direct descendants of the Jomon people. Next, we considered that the Ainu split from CHB at TAIN-CHB and that JPT diverged from Ainu at TAIN-JPT. In contrast, the replacement model stated that the modern Japanese were of Yayoi ancestry and we postulated that JPT branched off from CHB at TJPT-CHB. The hybridization model was expressed as the admixture between Ainu and CHB that happened at TADMIX subsequent to the divergence between them at TAIN-CHB. Because complete isolation between Ainu and JPT seems to be an extreme assumption, we also incorporated a continuous gene flow between these two geographically proximate populations into the three models (fig. 1). Then, we tested the model selection with or without migration. We used kernel-approximate Bayesian computation (kernelABC), which is a likelihood-free approach for Bayesian inferences, to incorporate high dimensional summary statistics and improve approximation of posterior estimates given the data (Fukumizu et al. 2013; Nakagome, Fukumizu, et al. 2013; Nakagome, Mano, et al. 2013; Osada et al. 2013; Sato et al. 2014). 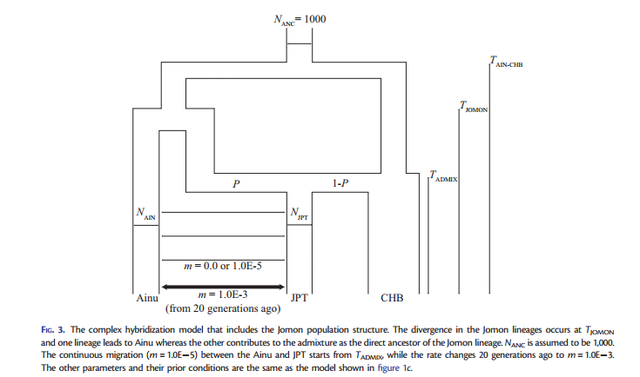 FIG. 3. The complex hybridization model that includes the Jomon population structure. The simplified hybridization model chosen from our model selection can be used as a starting point for further investigation into more realistic scenarios consisting of various prior conditions. Here, we hypothesized several complex hybridization models by including different sets of priors for three parameters (fig. 2): 1) symmetric beta distributions, which is B(, ), for the prior of admixture proportion (P) (supplementary fig. S2, Supplementary Material online); 2) ancestral population sizes before the divergence between Ainu and CHB (NANC = 1,000, 2,500, 5,000, or NAIN which follows lognormal (LN; 5,000, 5,0002 )); and 3) migration rates between Ainu and JPT (m= 0.0, 1.0E5, 1.0E3, or 1.0E1). First, the hybridization model is characterized by the admixture event between Ainu and CHB; hence, the choice of hyper parameters for the prior of P can be crucial in estimating the posterior distribution. We compared MLs of six hyper parameters ( = 0.1, 0.25, 0.5, 1.0, 2.0, and 5.0) under NANC =NAIN and m= 0.0 (supplementary table S2, Supplementary Material online). Although differences in their aMLs were less signifi- cant in terms of aBFs, the U-shaped prior with = 0.5 exhibited the highest likelihood among the hyper parameters. Second, NANC is likely to greatly contribute to shaping the amount of genetic variation among the three populations. We tested four conditions in which NANC was the same as NAIN or fixed at 1,000, 2,500, and 5,000 under m= 0.0 (supplementary table S3, Supplementary Material online). The fixed NANC = 1,000 showed a significantly higher likelihood than the other conditions. Third, continuous migration between Ainu and JPT after the admixture event can be important to defining shared genetic variation (supplementary table S4, Supplementary Material online). The likelihood of m= 1.0E5 was significantly higher than m= 1.0E3 and 1.0E1. Meanwhile, the difference in the likelihoods between m= 0.0 and 1.0E5 was less significant, as indicated in table 1. 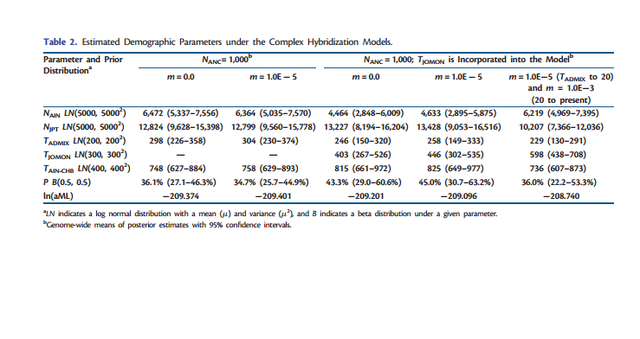 Table 2. Estimated Demographic Parameters under the Complex Hybridization Models. The highest likelihood model indicated an old divergence of Ainu and CHB populations an estimated 736 generations ago (95% confidence interval: 607–873), which was translated into 18,400 years ago based on the assumption of 25 years per generation. This model predicts the divergence of Jomon lineages occurred 598 generations ago (438–708) (14,950 years ago) with admixture occurring 229 generations ago (130– 291) (5,725 years ago) with 36.0% (22.2–53.3%) of the admixture proportion from Jomon ancestry to the mainland Japanese. NAIN was estimated at 6,219 (4,967–7,395) which is approximately a half of NJPT= 10,207 (7,366–12,036). In addition, the other models, either with or without assuming the population structure of the Jomon, indicated that TAIN-CHB was old and NAIN was a half or a third of NJPT. Meanwhile, the posterior estimates of TADMIX are still older than 120 generations ago (3,000 years ago), which archaeological and anthropological studies have traditionally proposed. Although estimating the parameters by fixing TADMIX at 120 revealed that the other parameters of NAIN, NJPT, and P were scaled down to half of those estimates from the models with unfixed TADMIX, the likelihoods were similar between the models with unfixed and fixed TADMIX (table 2 and supplementary table S6, Supplementary Material online). These results suggest that the estimates of TADMIX strongly depend on the other parameters, implying the difficulty in estimating TADMIX under the complex hybridization models. Our combined approaches of parameter estimation with model selection support the new model that the Jomon people were already diversified in the Japanese archipelago before the admixture event with the Yayoi occurred, and one of the Jomon lineages is a direct ancestor of the modern Japanese, yet the other lineage leads to the contemporary Ainu. Our inference shows that the Ainu and the CHB share an ancestral population that dates to the late Pleistocene; archaeological evidence shows the peopling of the Japanese archipelago began during this time frame. From these results we conclude that the ancestral people of the Ainu (or some of their ancestors) established the Jomon culture sometime at the end of Pleistocene, while a part of the ancestral population of CHB migrated to Japan and started admixing with the Jomon people about 5,000– 6,000 years ago. Our results strongly support the dual structure model for the peopling in the Japanese archipelago produced through hybridization between the Jomon and Yayoi people. Furthermore, we propose a more complicated model in which the Jomon people have diversified on the Japanese archipelago because they were isolated from the continent around 18,000 years ago. Admixture did occur between the Jomon and Yayoi people, yet some of the Jomon people still kept their own lineages without being strongly affected by the gene flow from the Yayoi people. The older divergence time of the Jomon and Yayoi lineages is compatible with the Hanihara’s hypothesis (1991), which claims the Jomon people are direct descendants of late-Paleolithic populations somewhere in Southeast Asia, whereas the Yayoi people were a population adapted to a cold climate in Northeast Asia. As the first step for resolving the peopling of Japan statistically and quantitatively, we simplified the models to make it possible to test the three hypotheses established from the previous studies. Then, we tested complex scenarios under the hybridization model to fit them with the gwSNP data. However, there should be other possible scenarios that we have to consider, and further studies taking account of realistic scenarios should be necessary to untangle a true evolutionary history of Japanese. To this end, our results can be used as a starting point to test more complicated hypotheses using a genomic-scaled data. In particular, the origins of the Jomon and Yayoi people and their demographic history, including the source of gene flow to Ainu, still remain unresolved. Future studies, including ancient genome analysis of Jomon and Yayoi skeletal remains as well as populationscaled gwSNP and genomic sequence analysis for east and northeast Asians will bring new insights into the complex demography in Japanese populations. Nakagome, Shigeki, et al. "Model-based verification of hypotheses on the origin of modern Japanese revisited by Bayesian inference based on genome-wide SNP data." Molecular biology and evolution 32.6 (2015): 1533-1543. |
|
|
|
Post by Admin on Oct 16, 2016 20:52:46 GMT
 Jōmon living in the Arctic-like northern islands of Japan subsisted on dolphins and whales, whereas those from Japan’s more tropical southern extreme engaged in more traditional fishing. Those in the middle—on the east coast of Japan’s largest island, Honshu—gathered shellfish but mainly lived off the forest. At first, this was a cold pine forest filled with large animals like elephants and bison. Then, as the world warmed early in the Holocene epoch, about 10,000 to 12,000 years ago, the pine gave way to a dense woodland of oak, maple, and birch inhabited by smaller prey like deer and wild boar.  Dogs, Perri reasoned, would have been highly valued in this Holocene forest, as they would be ideally suited to track, chase down, and hold these smaller prey animals. And indeed that’s what she found when she scanned the Japanese archaeological literature. Starting about 9000 years ago, the Honshu Jōmon began to bury their canine companions in shell middens—huge piles of seashells where they also typically interred their human dead. Like people, the dogs (which may have resembled Shiba Inus) were placed singly and appear to have been arranged in particular postures. “They looked like they curled up and went to sleep,” Perri says. Some had suffered what appeared to be hunting injuries—broken legs and teeth—and many of their bones had healed, suggesting people had taken care of them. Some were also found with grave goods, like shell bracelets and deer antlers. “They were treating their dogs the same way they treated their human hunters,” she says. Perri uncovered records of 110 burials in all, until about 2500 years ago, when the locals turned to agriculture. After this, canines appear in the archaeological record only as random piles of bones, often butchered, suggesting they were simply eaten and thrown away. The same was true all along in the Jōmon populations to the north and south, which did not need dogs for hunting.  The fact that the Japanese dogs were only revered in a time and place where they would have made ideal hunting companions strongly suggests that they did indeed play this role, Perri reports this week in Antiquity. She also points to a 2500-year-old bronze bell found on the east coast of Honshu that contains an engraving believed to depict an event from even further in the past: a boar surrounded by a hunter and his pack of dogs. Was the use of hunting dogs an adaptation to the post-glacial deciduous forest environment in the northern temperate zone? Dog burials in Jōmon Japan appear closely associated with a specific environment and with a related subsistence economy involving the hunting of forest ungulates such as sika deer and wild boar. Dogs were valued as important hunting technology, able to track and retrieve wounded animals in difficult, forested environments, or holding them until the hunter made the final kill. Greater numbers of dog burials during the later Jōmon phases may reflect a growing dependence on hunting dogs to extract ungulate prey from forests in an increasingly resource-strained seasonal environment. |
|
|
|
Post by Admin on Oct 20, 2016 21:04:17 GMT
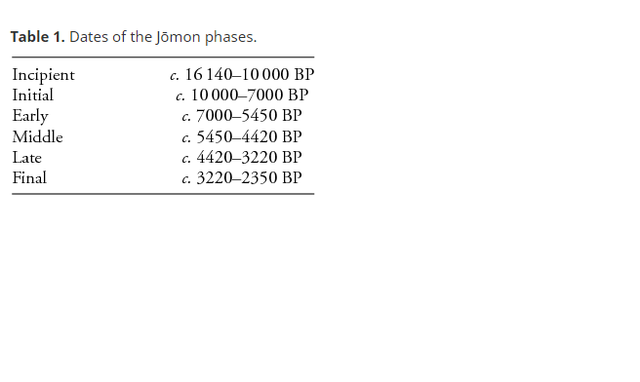 The Jōmon culture of Japan (c. 12 500–2350 BP; Table 1) is considered one of the best contexts for analysing complex prehistoric hunter-gatherer groups (Rowley-Conwy 2001). Although often discussed as a single culture that dominated the archipelago for over 10 000 years, the Jōmon actually comprised various subgroups with differing traits across regions (Bleed & Matsui 2010). These traits were specifically defined by the geography and climate of the various latitudinal zones from subarctic to subtropical, a result of the extreme north–south orientation of the islands. This variation in climatic conditions meant that subsistence systems practised by Jōmon hunter-gatherers in each area were highly varied, with different prey species, tool technology, hunting methods and environmental adaptations (Underhill & Habu 2006). Even within a small area in Japan, a mosaic of different environments is possible. Due to these variables, different degrees of complexity can be expected among the various Jōmon subcultures. This paper focuses on the Jōmon subculture that inhabited the eastern side of the main island, Honshu, hereafter referred to as the Pacific Honshu Jōmon (Figure 1). At the beginning of the Holocene, this region of Japan experienced a distinctive transition to environmental conditions that allowed Jōmon foragers to flourish in a forest-estuary ecotone consisting of abundant, nut-bearing deciduous trees, shellfish, coastal fish and forest ungulates such as sika deer (Cervus nippon) and wild boar (Sus scrofa leucomystax), which made up the majority of their diet. Also unique to this part of Jōmon Japan is the occurrence of the individual burials of domesticated dogs (Canis familiaris), only a few of which have previously been discussed outside of the Japanese literature. Researchers have long presumed that hunting dogs were kept by Jōmon foragers (e.g. Kraus 1953; Nishinakagawa et al. 1994), and in Japan “boar-hunting with dogs is seen as a quintessential Jōmon activity” (Knight 2003: 153). Yet this idea has not moved beyond the theoretical, even though archaeological dog remains have been systematically surveyed across Japan (see Kaneko 1978; Shigehara 1985; Niwa 1987; Kojima & Kikuchi 1999). Here, I discuss a possible relationship between the isolated clustering of Jōmon dog burials in just one region and the specialised adaptations to changing environments and prey in this locality. I document a strong association between the first appearance of Jōmon dog burials in eastern Honshu and a shift to primarily hunting terrestrial ungulates in the new Holocene deciduous forests of the region, signifying the probable use of dogs as a dense-forest hunting adaptation after the Pleistocene–Holocene transition.  Post-glacial shifts of flora and fauna in north-central Honshu prompted a reorganisation of subsistence strategies, requiring adaptations away from hunting the large terrestrial megafauna of the Pleistocene—e.g. Naumann's elephant (Palaeoloxodon naumanni) and Yabe's giant deer (Sinomegaceros yabei)—and towards a strategy of taking smaller, quicker ungulates in a densely forested environment (Inada 1986; Tsuji 1997). Changes in prey are reflected in the technological advances seen from the region, including a shift to smaller, triangular points used for the bow and arrow (Aikens & Higuchi 1982; Inada 1986), which were designed to induce heavy bleeding in ungulates (Friis-Hansen 1990; Churchill 1993). This change in blade technology began in the southern islands, moving north with the changing biota, suggesting strong connections between Jōmon environmental and cultural changes (Aikens & Akazawa 1996). In the temperate deciduous forests of north-central Honshu, there may have been variation between Jōmon on the Japan Sea and Pacific coasts, with an increased reliance on plant resources along the Japan Sea, although more dietary isotopic work is needed for this region. The Japan Sea coast, west of the central mountain range, is well known for its heavy, long-lasting snowfall, which sika deer and boar migrate to avoid (Tsujino et al. 2010). Minaki (1988) suggests that extensive chestnut cultivation occurred along this coast, with raised-floor longhouses—associated with the winter storage of nuts in high snowfall areas—found predominantly in this region (Kitagawa & Yasuda 2008). In contrast, a substantial primary dependence on sika deer and wild boar by Jōmon on the Pacific Honshu coast has long been established by researchers (Koike 1986; Hongo et al. 2007). In these two productive temperate forest environments, however, the importance of ungulate hunting on the Pacific coast, as compared to the Japan Sea coast, is probably closely related to the availability of prey during the key autumn and winter months.  Dogs as hunting technology A hunting partnership between dogs and humans has long been postulated in the archaeological literature, with some researchers suggesting that such a collaborative alliance was the basis for the initial domestication of dogs (e.g. Davis 1982; Clutton-Brock 1995). A partnership of this nature has often been proposed between Jōmon hunters and their dogs, given that terrestrial game hunting was an important part of the subsistence economy of some regional subgroups (Nishinakagawa et al. 1994; Kobayashi 2004). The shift in hunting strategies following the Pleistocene–Holocene transition probably included hunting dogs as a combined, dense-forest technological innovation along with the bow and arrow. The innate ability of a dog to sniff out, track, chase and hold prey can significantly enhance the success of human hunters in forested environments (e.g. Dwyer 1983; Ngima Mawoung 2006). Dogs are an important, and in some cases indispensable, hunting aid for many modern forager groups, as they probably were for foragers in prehistory. Their use is often a critical factor in the minimisation of subsistence risk and the maximising of hunting returns; they can prove an invaluable extension of the hunter and their toolkit (Mitchell 2008). The rapid spread of post-glacial temperate forests in north-central Japan increased the total ungulate biomass, which may have been a crucial variable in human behaviour, organisation and populations in the early Holocene (Mellars 1975; Rowley-Conwy 1986). These areas of high-value prey species were ideal hunting grounds for the Jōmon, yet the density of the temperate forests and swiftness of medium-sized ungulates would have required adapted hunting methods compared to the more open habitats and large herd animals of the previous glacial period. Clutton-Brock (1984) suggests that hunting dogs were heavily used in the early Holocene—in conjunction with microlith technology—to track and retrieve wounded game in difficult, forested environments. Wild boar are particularly sensitive to vegetation type, preferentially inhabiting areas with the densest cover (Melis et al. 2009; Saïd et al. 2012), making dogs particularly useful for boar hunting. The use of dogs as hunting tools is widespread in the ethnographic literature, especially in the hunting of deer and wild boar in forested environments (e.g. Ngima Mawoung 2006; Pannell & O'Connor 2010). Modern hunters emphasise the importance of hunting dogs in dense woodland, where human sensory and locomotor skills are diminished (e.g. Ellen 1999; Chitwood et al. 2011). Injured deer often run, leading hunters on long chases, and wild boar can be aggressive and quickly learn to evade capture. Hunting dogs mitigate these factors by tracking blood trails, forcing game into vulnerable positions (e.g. in water) and holding prey until the hunter can make the final kill (Rühe et al. 2006; Saïd et al. 2012). Specifically, the successful hunting of wild boar often requires highly skilled dogs, which are prized above all others, and without which many hunters attest boar hunting would be virtually impossible (Bulmer 1968; Dwyer 1983). The effectiveness of hunting dogs in the Pacific Coast Jōmon environment, along with the presence of many dog burials in this region, indicates that Jōmon hunters were probably using dogs as tools for the hunting of sika deer and wild boar, as hunters in Japan still do today. 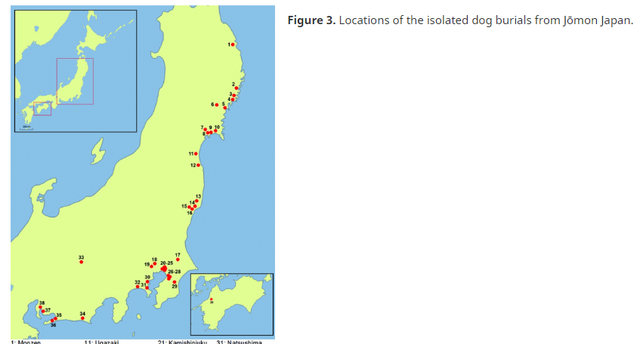 A comprehensive survey was undertaken of Jōmon dog burials in the archaeological literature (Japanese and Western language; details are available in the online supplementary material). Over 110 burials are identified from 39 archaeological sites (Figure 3). The dog burials discussed are all isolated burials: intentional, buried alone and with no obvious signs of butchery or human-induced death noted (cf. Perri in press). While 110 burials have been individually documented, some reports were ambiguous, noting only that dog burials were encountered. This implies the actual number of isolated burials is greater than 110. Importantly, isolated dog burials from Jōmon Japan are found almost exclusively in the eastern half of north-central Honshu, correlating with the deciduous forest-terrestrial ungulate economy of the Pacific Coast Jōmon. Burials begin in the Initial phase, with single burials at two sites, including the only example not located in north-central Honshu (Figure 4; Table 2). By the Early phase, burials occur farther north and in greater numbers. In the Middle phase, burials become more widespread across the Pacific coast of north-central Honshu, with more sites and more burials. The Middle phase also has the only reported inland dog burial(s), although the number of animals and details are not given. Large numbers of sites and burials continue during the Late and Final phases, with burials widespread across the entire Pacific coast of north-central Honshu. After the Final phase the practice of dog burials seems to terminate, as dog burials are unknown in the ensuing agricultural Yayoi period (beginning c. 2350 BP), further suggesting that dog burials are closely related to hunting activities during the Jōmon period. The association between Jōmon dog burials and the deciduous forest-estuary ecotone is strongly supported by the fact that 37 of the 39 dog burial sites are shell middens. Injuries, mostly healed broken bones, were evident on dog remains from seven sites. It is possible these are related to the hunting of ungulates, as has been suggested for other prehistoric dogs (Warren 2004) and modern wolves (Mech & Nelson 1990). The ages of the dogs range from newborn to over 12 years old. The burial of immature dogs may not normally be associated with those distinguished as capable hunters, yet the ethnographic record shows that puppies in hunter-gatherer groups are often valued for their potential as future hunting partners (e.g. Terashima 1983; Koster 2008), as Clutton-Brock (1995) has previously suggested for prehistoric puppies. Grave goods (an oyster shell bracelet; Horikoshi 1977) were noted from only one burial, although another dog burial was covered with stones (Otake 1983). Antiquity Volume 90 Issue 353 |
|
|
|
Post by Admin on Jan 23, 2017 20:35:15 GMT
 Two Cro-Magnons' mtDNA haplogroup (23,000-year-old Paglicci 52 and 24,720-year-old Paglicci 12) was identified as haplogroup N and mtDNA haplogroups N9b and M7a are called the Jomon haplotype. More specifically, the two Cro-Magnons carried macrohaplogroup N, containing haplogroups W, X, I, N1a, N1b, N1c, and N* (Caramelli et al. 2003). The Jomon and Cro-Magnons share common maternal ancestry, contributing to the morphological similarity between them. Moreover, Kennewick Man's mtDNA haplogroup was recently identified as haplogroup X2a by Rasmussen et al. (2015) and some Native Americans with this haplotype are genetically related to Cro-Magnons, which is present among the Algonquian people (25%), the Sioux (15%), the Nuu-chah-nulth (11%–13%), the Navajo (7%), and the Yakama (5%). 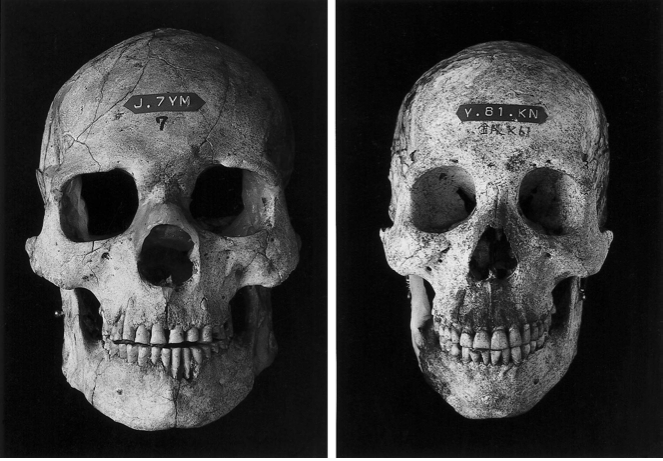 Jomon skulls The Out-of-Africa model (6), on the other hand, suggests that a.m.h. first arose in Africa some 150,000 years ago and then dispersed replacing archaic forms (Neandertals in Europe). This model does not imply any hypothesis regarding the structure of archaic humans populations, nor that modern humans migrating from Africa were a biologically distinct species. It only assumes that modern traits evolved recently in a single region, Africa, and the vast majority of the genomes of current human populations can be traced back to these African migrants. This hypothesis does not seem to differ radically from Relethford's (4) view of multiregional evolution, whereby the Neandertal's contribution to the modern European gene pool may have been small, yet nonzero. Analyses of morphological traits, Neandertal ancient DNA and modern DNA (e.g., refs. 7–9), appear to support a recent African origin of all humankind. However, it has been argued that patterns of genetic diversity are not incompatible with a multiregional model (5, 10–12). For instance, Nordborg (10) showed that the differences between Neandertal's and modern mitochondrial sequences are sufficient to rule out random mating between them, but not more complicated models of interbreeding. Those results reflect the existing uncertainties on the European demographic history of the last 30,000 years. Clearly, genetic typing of the earliest a.m.h. in Europe, sometimes referred to as Cro-Magnons or Cro-Magnoid from the site in France where they were first discovered, is a crucial step for solving this question (13–15), because that would allow a genetic comparison between individuals who lived at a much shorter (ideally, zero) time distance. The Out-of Africa model, in fact, predicts genetic discontinuity between Neandertals and early a.m.h. (the former being a separate lineage replaced by the latter) and genetic continuity along the a.m.h. lineages from the Upper Palaeolithic until the present. The multiregional models, on the contrary, predict at least some level of genetic continuity from the archaic Neandertal forms to the almost contemporary Cro-Magnon forms up to today's Europeans.  The D-loop sequence of Paglicci-25 shows no substitutions with respect to the Cambridge reference sequence (28). This sequence is fairly common (14%) in a database of 2,566 today's Europeans and Near Easterners from 35 different populations (updated from ref. 20). The average number of substitutions between Paglicci-25 and these 2,566 sequences is 2.34 (SD = 1.75, range 0–11), whereas the average number of substitutions between today's Europeans is 4.35 (SD = 2.32, range 0–18). The comparison of this 23,000-year-old a.m.h. with four Neandertals dated at ≈29,000, 40,000, 40,000, and 42,000 years ago (8, 16–18) produces very different results: 23, 28, 23, and 24 substitutions separate Paglicci-25 from the Neandertals, respectively. The second individual we typed, Paglicci-12, shows a single difference (a C → T transition at nt position 16,223) from the Cambridge reference sequence, and 3.17 differences on the average from today's Europeans (SD = 1.66, range 0–10). As in the case of Paglicci-25, Paglicci-12 is very different from the four Neandertal sequences: 22, 27, 22, and 23 substitutions, respectively. The consensus sequences of Paglicci-25 and Paglicci-12 are different from the sequences of all scientists who manipulated the bones during the laboratory analyses, thus also excluding the possibility of recent contamination. This result is graphically summarized in the multidimensional scaling analysis (MDS), where the plotted points correspond to the Paglicci, Neandertal, and representative today's sequences, and the genetic distances among them are proportional to the linear distances between the points in Fig. 1. Neandertals and a.m.h. form two distinct clouds of points, respectively, with the Paglicci sequences falling well within the range of variation of today's humans. We note that if the sequence from the 40,000 (30) years old sample Lake Mungo 3 (an anatomically modern Australian individual, ref. 31) is added to the MDS analysis, that pattern does not change, and two distinct clusters of data points remain evident. If genuine (25, 32–34), the sequence of Lake Mungo 3 is among the most divergent modern human mtDNAs, but yet it can be unequivocally attributed to the modern cluster of individuals in Fig. 1.  Fig. 1. MDS of HVRI sequences of 60 modern Europeans (filled squares), 20 modern non-Europeans (filled circles), 4 Neandertals (open diamonds), the Australian Lake Mungo 3 (open circle), and the two early a.m.h. typed in this study (open squares). European and non-European sequences in this figure were selected to represent the most divergent lineages observed in modern individuals. Note that the axes have different scales. The stress value for this analysis was 0.128. |
|
|
|
Post by Admin on Jan 24, 2017 20:24:12 GMT
 Fig. 2. Average genetic distance between ancient and modern samples (2,566 sequences of modern Europeans; y axis), as a function of the samples' age (x axis, in thousands of years). Vertical lines represent two standard deviations above and below the mean. Squares, a.m.h. Diamonds, Neandertals. The Paglicci samples typed in this study are indicated by open squares. The point at 0 years indicates the average pairwise difference between present-day samples. Specific mtDNA sites outside HVRI were also analyzed (by amplification, cloning, and sequencing of the surrounding region) to classify more precisely the ancient sequences within the phylogenetic network of present-time mtDNAs (35, 36). Paglicci-25 has the following motifs: +7,025 AluI, 00073A, 11719G, and 12308A. Therefore, this sequence belongs to either haplogroups HV or pre-HV, two haplogroups rare in general but with a comparatively high frequencies among today's Near-Easterners (35). Paglicci-12 shows the motifs 00073G, 10873C, 10238T, and AACC between nucleotide positions 10397 and 10400, which allows the classification of this sequence into the macrohaplogroupN,containing haplogroups W, X, I, N1a, N1b, N1c, and N*. Following the definition given in ref. 36, the presence of a single mutation in 16,223 within HRVI suggests a classification of Paglicci-12 into the haplogroup N*, which is observed today in several samples from the Near East and, at lower frequencies, in the Caucasus (35). It is difficult to say whether the apparent evolutionary relationship between Paglicci-25 and Paglicci-12 and those populations is more than a coincidence. Indeed, the haplogroups to which the Cro-Magnon type sequences appear to belong are rare among modern samples, and therefore their frequencies are poorly estimated. However, genetic affinities between the first anatomically modern Europeans and current populations of the Near East make sense in the light of the likely routes of Upper Paleolithic human expansions in Europe, as documented in the archaeological record (37). A pattern of genetic distances through time emerges clearly when four additional HVRI sequences from prehistoric anatomically modern Europeans dated between 5,500 and 14,000 years ago (38, 39) are considered (Fig. 2). Going back in time from the present to 25,000 years ago, prehistoric Europeans show an approximately constant number of differences in comparison with today's Europeans, very similar indeed to the number of differences between two randomly chosen modern sequences. Conversely, an abrupt increase of genetic distance from today's Europeans is observed for the Neandertals, even though the most recent of them is separated from the older Paglicci sample by only a few hundreds of generations. Therefore, these results suggest a pattern of genetic continuity in the modern humans' genealogy from the Upper Palaeolithic period to the present, but a clear discontinuity with respect to Neandertals of similar ages.  Neandertal mtDNA phylogeny from Dalen et al. 2012 Even the most stringent available criteria for validating ancient human DNA sequences do not allow one to prove that the sequences determined are authentic. Only if a sequence is radically different from modern ones, as is the case for Neandertals, can one be relatively sure that no contamination has affected the results. Therefore, a certain degree of prudence is necessary before drawing any conclusions from this study. Still, none of the biochemical tests we carried out suggests that different sequences (namely the endogenous one plus some contaminating sequences) were amplified from the 23,000- and 25,000-year-old specimens that we used. In addition, the amino acid racemization test strongly suggests that reasonably well preserved DNA should be present in those specimens. Because DNA from all four Cro-Magnon type bone fragments could be amplified and sequenced only by using primers specific for modern humans, and not for Neandertals, there is little doubt that the mtDNAs of early a.m.h. and of cronologically close Neandertals were, at least, very different. Under the multiregional model of human origins, Neandertals and modern humans are just one population observed at different times (1, 2). Therefore, when comparing sequences sampled along the single human–Neandertal genealogy, one should not observe any major discontinuity. More recent versions of the multiregional model (3–5) suggest that Neandertals and early a.m.h. were regional populations of the same evolving species connected by gene flow, and both archaic and modern forms contributed, possibly in different proportions, to the present day human gene pool. Under this updated multiregional model, the absence of Neandertal mtDNA lineages in living humans is regarded as a consequence of a random drift or a selection process of lineage extinction since the disappearance of Neandertals. In this case, unless the extinction of Neandertal lineages was almost instantaneous, the probability of finding such lineages in early a.m.h. should not be too low. All these expectations are inconsistent with the data and the analysis here presented. Two a.m.h. dated between 23,000 and 25,000 years ago appear to have HVRI sequences fully compatible with the variation observed both in contemporary and in ancient samples of a.m.h., and certainly they do not show any special relationships with the almost contemporary Neandertals. These results are at odds with the view whereby Neandertals were genetically related with the anatomically modern ancestors of current Europeans or contributed to the present day human gene pool. Although only six HVRI sequences of ancient a.m.h and four sequences of Neandertals are available to date, the sharp differentiation among them represents a problem for any model regarding the transition from archaic to modern humans as a process taking place within a single evolving human lineage. David Caramelli, 6593–6597, doi: 10.1073/pnas.1130343100 |
|
















